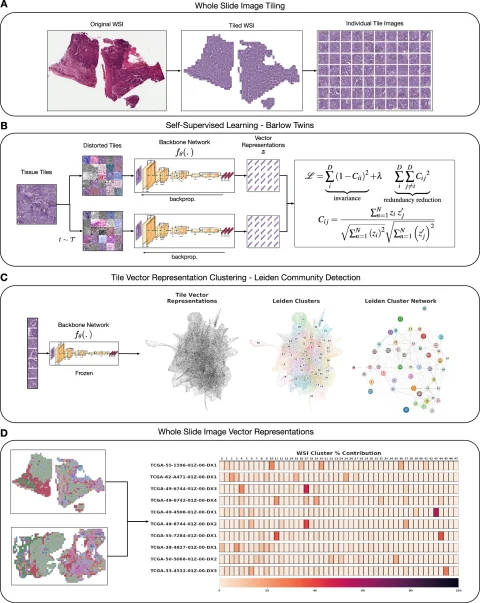
Histomorphological phenotypic learning (HPL) framework architecture overview: The entire slide image (WSI) is processed for tile extraction and normalization of staining. b Self-monitored training of the backbone network Fθ creates a tile vector representation. The C tiles are projected onto the Z vector representation using the frozen backbone network Fθ. Continuingly, phenotypic clusters of histology morphology (HPCs) are defined using Leiden Community detection for nearest neighbor graphs of Z-tile vector representations. D WSI or patient (one or more WSIs per patient) is defined taking into account compositional vectors with dimensions equal to the number of HPCs and the proportion of HPCs to total tissue area. HPL creates WSI and patient composition vector representations that can be easily used in interpretable models such as logistic and COX regressions that associate tissue phenotypes with clinical annotations.
Image sources: Claudio Quiros A, Coudray N, Yeaton A et al. , Nature Communications 2024 (CC by 4.0)
From there, H&E images can be converted into maps of histomorphological clusters (HPCs) that provide interpretable quantitative histopathology, containing classical forms of cancer, and predicting recurrence through its maps that provide a detailed morphological picture. “It provides us with a truly interpretable quantitative tool to classify images and search for tissues, and we can see the morphological content of that tissue very quickly,” he continued. “More than that, it can be used using compositions to ask if there are clusters that have a significant impact on patient outcomes. The HPC composition is highly prognostic in the validation cohort.
Powerful Tools – There are some remaining challenges
Experts described it as a “powerful tool” that surpasses AI grading that he oversaw pathologists' supervised. 'Self-learning methods like HPL learn the meaningful pathology present in these images without supervision. It provides quantitative, interpretable morphological dictionary, automate identification, outperforms supervised methods and human experts of key tasks such as subtyping and prognosis, and discovers “new” features and morphology,” Le Quessnet said.
He further highlighted how self-learning AI can be deployed to other imaging modalities such as multiple immunofluorescence (IF) and applied to the immune microenvironment. “We found that using images that go through the same process with much smaller tiles allows us to use greater power than human grading across the slide image to really accurately predict the outcome,” he added.




