Drug discovery is like molecular Tetris. Chemists snap atoms together, tweak them until everything fits, and suddenly the molecule creates a promising new drug. Creating better molecules usually takes an enormous amount of time and money.
In a new study, researchers used machine learning to build a smarter predictive system that can speed up the process at a fraction of the cost.
“To understand new reactions, we sometimes use advanced physics-based computational chemistry tools, but these tools are too expensive to predict thousands of potential new molecules,” said Simone Galarati, co-first author of the study and a joint postdoctoral fellow at the University of Utah and the University of California, Los Angeles. “We wanted to train a statistical model that was as cheap as possible while still being ‘smart’ enough to make accurate predictions about untested responses.”
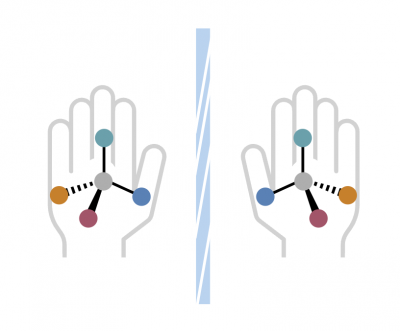
Molecules can exist as mirror images, a property known as “handedness.” Left and right hand form is very important. One may be curative, the other may be harmful. Chemists need to find the right set of tools: catalysts, ligands, and substrates to ensure they build the correct version.
The new system acts as a high-tech filter that can screen tens of thousands of chemical structures to predict how their parts will combine to make some molecules more “hands” than others. This workflow provides a cost-effective way to convert the components of a reaction into numerical data that computers can analyze and build a framework for machine learning predictions.
With surprisingly little input, the model reliably predicted how the component would behave, reducing the time, energy, and money spent on reaction testing in the lab.
“Most AI requires huge amounts of data to train the models. This is a problem in chemistry where obtaining high-quality, large datasets from experimental work is very expensive and very time-consuming,” said university chemist Matthew Sigman, co-author of the study. “The best part about this tool is that you can collect a small amount of data and build a pretty good model to accurately predict reactions that are known. You can also transfer the predictions to reactions that the model hasn’t seen yet.”
The study was published in accelerated preview in Nature on February 11, 2026.
high tech filter
The researchers built a workflow around asymmetric cross-coupling reactions, a powerful toolkit for drug development. This reaction uses a metal catalyst to join two carbon-based molecular fragments together to build a more complex compound. This reaction is called asymmetric because it is designed to favor one “hand” version of the molecule. Chemists often create both versions, but without guidance, experiments yield 50/50 results. In contrast, an asymmetric reaction yields, for example, 95% of the desired form and only 5% of the undesired mirror image.
Asymmetric cross-coupling reactions typically require at least three elements: a metal, a ligand, and a substrate. Metal catalysts do the heavy lifting of bonding carbon-based molecules together to build products. Ligands bind to metals, control which side of the molecule reacts, and influence the three-dimensional orientation of the products. Ligands are perhaps the most important element controlling the handedness of molecules.
To train the model, Galarati and his team identified four academic papers on asymmetric reactions, including previous work by co-authors Abigail Doyle and Sigman. All these papers used nickel-based catalysts with different ligands. These results were the only training data for the workflow. The team then asked the system to predict outcomes for hypothetical components that were not included in the training data. They added a series of increasingly difficult tasks that forced the algorithm to make predictions using material that was increasingly dissimilar to the original training data. The research team tested this prediction in Doyle’s lab. The study is an effort led by Erin Bucci, the study’s co-lead author and a doctoral student at UCLA.
“As a lab-based chemist, this tool is extremely valuable in saving time spent performing experiments,” Bucci said. “For example, instead of running 50 to 60 reactions, you can now run 5 to 10 reactions, potentially saving weeks or months. Each reaction component you test in the lab must be purchased or made from scratch. This tool significantly reduces the amount of money you would normally spend on materials.”
Although the authors tested the tool in the context of novel nickel-based reactions, the workflow can be applied across disciplines and can also improve our understanding of the chemistry itself.
“One of the nice things about the workflow is that it’s not a black box,” said Abigail Doyle, a UCLA chemist and co-author of the study. “Even if the predictions are wrong, we can still learn something about chemistry from the predictions. We apply our chemistry expertise to help us learn something we wouldn’t have learned without the tools.”
The pharmaceutical industry will soon benefit from such tools, Sigman added. Suppose a company needs to deliver a large quantity of a compound for clinical trials and wants to apply a reaction that is already in the literature. But it has never been done for a specific composite target.
“This is where this tool is very applicable,” he said. “Optimizing reactions and time costs is a value proposition when developing drugs. This streamlined process can make a difference when a molecule needs to move from Phase 1 to Phase 2.”
****
The study was published in the journal Nature under the title “Transferable enantioselectivity models from sparse data.” https://doi.org/10.1038/s41586-026-10239-7
This research was supported by the Swiss National Science Foundation (#222115), the U.S. National Science Foundation (CHE-2202693 and CHE-1048804), the National Institutes of Health (S10OD028644), and the University of Utah High Performance Computing Center.