Dataset Presentation
The dataset used in this study consisted of consisting of 1,200 patient records. Collected in Excel files with structured labeled columns for different clinical and lifestyle data. This study used a subset of the publicly available cancer prediction dataset (1,500 out of 1,500 samples) shared under attribute 4.0 (4.0).32. All rows are individual patients, and all columns relay specific functions for cancer risk and diagnosis. The dataset includes a total of nine features in addition to the target variables used for prediction. A detailed description of all features and their types is given in Table 2.
Diagnostic variables are the targets for predictions for this study. It promotes patient classification according to the cancer diagnosis status. The dataset is distributed fairly between characteristics and target classes, facilitating an impartial training procedure for the ML model.
A frequency distribution chart called histograms created age, BMI, physical activity, and alcohol intake to visualize the distribution of continuous variables within the dataset. These factors represent important aspects of individual health and lifestyle that may be related to cancer risk. The age distribution of subplots is primarily uniform over the range of 20-80, with a slight increase in the frequency of older adults. BMI distribution of subplots Figure 2(b) is uniformly distributed between 15 and 40, meaning a heterogeneous patient population in terms of body composition. Physical activity figure 2(c) of the subplot shows variation across the population and is observed at all levels between 0 and 10 hours per week. The alcohol consumption depicted in the subplot is shown in Figure 2(d) with a uniform distribution ranging from 0 to 5 units, with no significant groupings. The histogram shows that the dataset exhibits authentic variability and is balanced and essential to construct a robust, generalizable ML model, as seen in Figure 2.
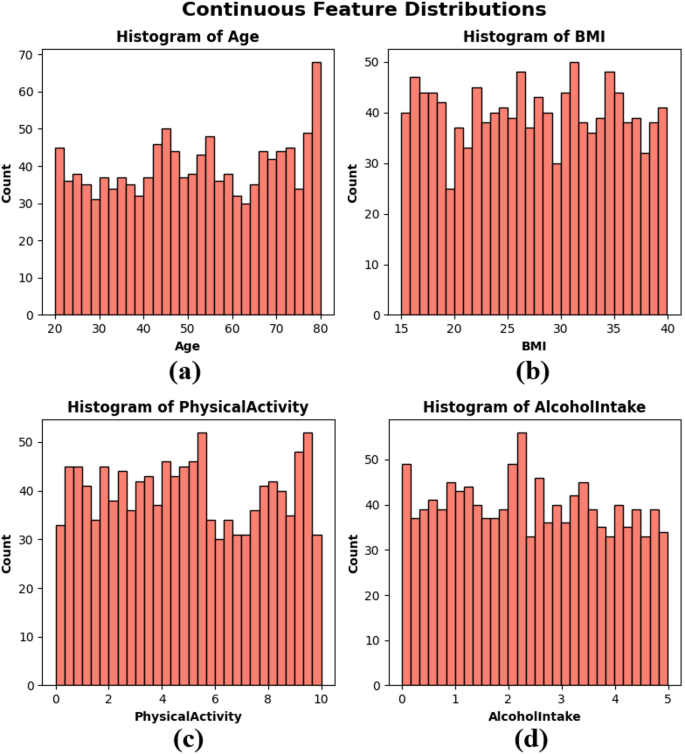
Continuous Function Histogram – (a) Year,b) bmi, (c) physical activity, and (d) alcohol intake.
Boxplots were created to show the distribution, central tendencies, and possible outliers of continuous variables within the dataset. Figure 3(a) shows the age distribution for the median age of about 50 years, the interquartile range (IQR) of about 35-65, and the absence of extreme outliers. BMI Functionality of the subplot Figure 3(b) shows a narrow, balanced distribution with a median value of around 27, with a narrow IQR showing similar body composition values between samples. Physical activity figure 3(c) in the subplot ranges from 0 to 10 hours per week with a median of nearly 5, reflecting a balanced distribution of patient physical activity levels. Finally, the subplot alcohol intake figure 3(d) also shows a wide distribution of 0-5 units per week, with a median slightly above 2 units. BoxPlots, as shown in Figure 3, shows that the dataset does not have large skewness or extreme outliers for continuous features, supporting the fit of the training model in ML.
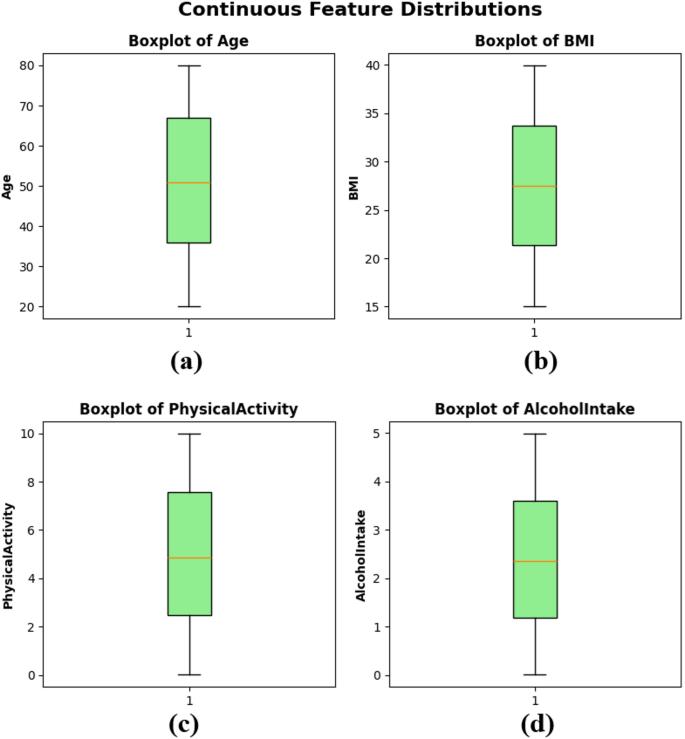
boxplots for continuous functionality – (a) Year,b) bmi, (c) physical activity, and (d) alcohol intake.
Count plots were created to assess the distribution of categorical and binary data, particularly regarding gender, smoking, cancer, and genetic environment. Figure 4 shows that the gender distribution of the subplots is roughly balanced between males (0) and females (1) and implies an impartial representation of the overall gender. Figure 4(b) shows that the majority of patients are nonsmokers, potentially indicating lifestyle trends in the population. In sidebar Figure 4(c), most patients do not show a personal history of cancer, but still a prominent proportion has been diagnosed previously. Genetic risk factors for subplots Figure 4(d) is categorized into low (0), medium (1), and high (2) levels. Most patients are classified as low-risk, while a small number is considered high-risk. These plots show the balanced properties and authentic variability of the dataset, confirming that the model is trained on a representative sample, as shown in Figure 4.
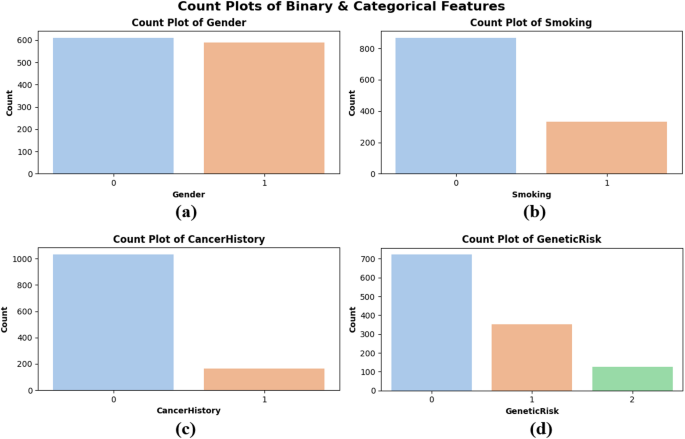
Count plots of binary and category features – (a) Gender, (b)smoking,(c) cancer history and (d) Genetic risk.
Matrix correlations provide statistical readings for each linear relationship between either the function of the dataset and the target (diagnostics). As can be seen in Figure 5, CancerHistory has the highest correlation with the target, with a correlation value of 0.41. This means a mild linear relationship. This means that patients who have had cancer in the past are more clinically relevant and are more likely to have cancer in the future. Other characteristics that are highly correlated with target variables include gender (0.28), genetic risk (0.27), and smoking (0.26). While most feature pairs exhibit weak or negligible correlations, such as BMI, gene inducibility, age and physical activity, this diversity indicates multidimensionality of the dataset and justifies the use of nonlinear models to capture complex interactions. The above correlation observations in Figure 5 are justified to further influence on predictive modeling, thus confirming the importance of selected features.
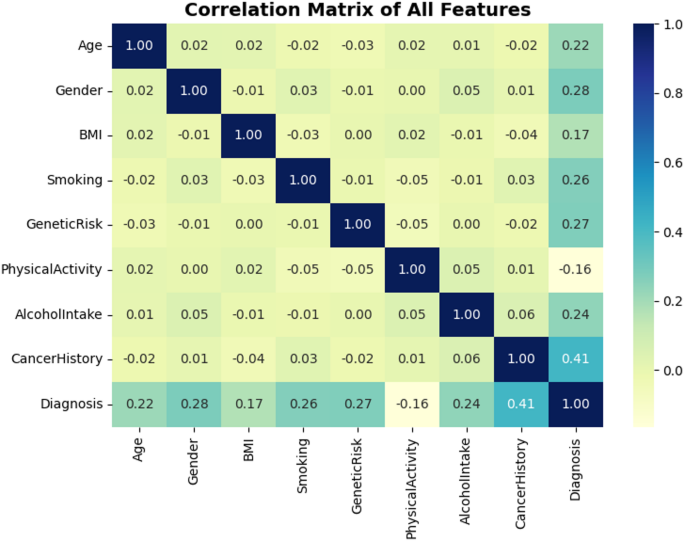
A correlation matrix for all functions including diagnosis.
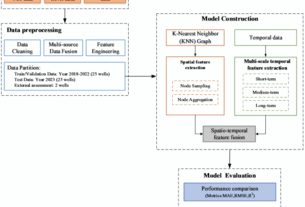
Workflow Overview
The methodology of this study follows a structured, data-driven approach to developing accurate cancer prediction systems using ML, as shown in Figure 6. The process begins with loading a dataset containing various patient characteristics such as age, gender, BMI, smoking status, genetic risk, physical activity, alcohol intake, and personal history of cancer. Once loaded, the dataset undergoes thorough investigation to understand its structure and integrity. Basic information such as shape, sample rows, and missing values is examined, followed by a statistical summary that provides insight into the distribution of each feature. To improve this understanding, EDA is done through visualizations such as histograms, box plots, and count plots to help identify patterns and anomalies. A correlation matrix is also generated to detect relationships between numerical variables.
Following the investigation, if the dataset is split into input functions and target labels, data preprocessing is performed. Continuous features are standardized using StandardScaler to ensure uniform scaling between models. The core of the methodology is in training logistic regression (LR), decision tree (DT), RF, gradient boost (GB), support vector machines (SVM), K-NEARest Neighbors (K-NN), CatBoost, Emptoce Boosting (XgBoost), and Light Gradient Boosting Machine (lightgifting cossgbmid), Catboost, extream boosting (k-nn), Support Vector Machines (SVMS), Support Vector Machines (SVMS), Support Vector Machines (SVMS), and Support Vector Machines (SVMS). This ensures a balanced and robust evaluation by evaluating the accuracy of each model across different data segments. The model with the highest average cross-validation accuracy is chosen as the best performance model.
To further verify the performance of the model, train test divisions are applied, and all models are evaluated with invisible test data. Key metrics such as accuracy, accuracy, recall, and F1 score are calculated for each model, and the confusion matrix is plotted to visualize classification performance. All evaluation results are saved as CSV files for reproducibility and reporting support. Finally, a GUI is developed using TKINTER, allowing users to enter patient information and receive instant predictions from the trained model. This end-to-end workflow ensures that the system is not only accurate and reliable, but also practically accessible and user-friendly.
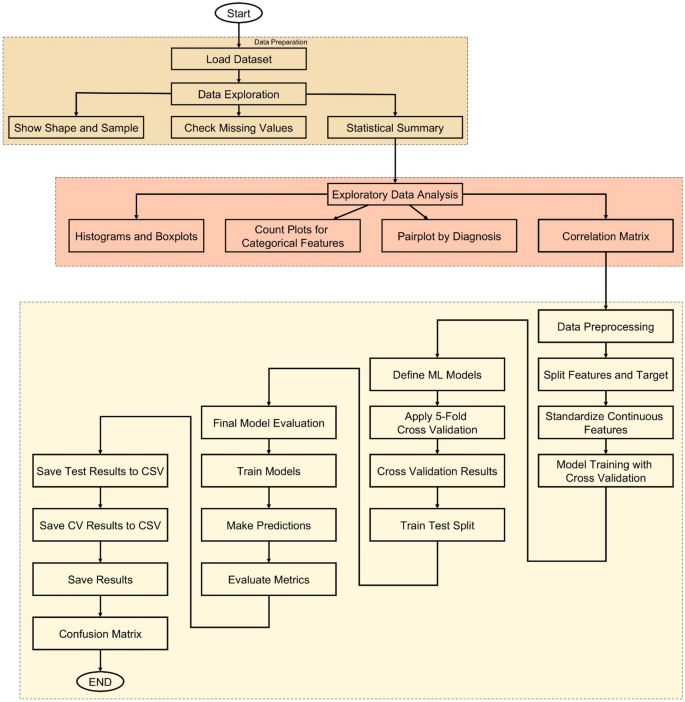
End-to-end workflow for cancer prediction systems that incorporate data exploration, model training, evaluation, and GUI deployment.
Rating Metrics
Several general classification metrics were employed to verify the effectiveness of the developed ML model. These measures help to have a complete overview of the efficiency of each model in isolating patients into patients with or without illness, a very important aspect of all health-related prediction tasks.
One of the indicators is the accuracy defined, in particular, as the percentage of correctly classified predictions (both negative and positive) across all predictions performed. This provides a general sense of model accuracy, as shown in the equation. (1). However, accuracy alone may not be sufficient for medical datasets, especially when FNS or FPS costs are high.
Accuracy and recall were also evaluated for deeper insights. Accuracy measures the percentage of true positive (TP) cases among all positive predictions. This is important to minimize false alarms and avoid unnecessary concerns and tracking procedures for healthy individuals. (2). On the other hand, also known as sensitivity, focuses on the ability of the model to correctly identify actual cancer cases, ensuring that as few positive cases as possible are not detected. This is calculated as explained in the equation. (3). Because there is usually a trade-off between accuracy and recall, this study used F1 scores to use a single performance score that balances both. This is useful when you take the accuracy and recall of the harmonic average and there is a slight class of imbalance, or when both FPS and FN are concerned. Mathematically, the F1 score is defined by an equation. (4).
In addition to these scalar metrics, a confusion matrix was created for each model, indicating the amount of TP, true negative (TNS), FPS, and FNS. This has encouraged a clear understanding of the performance of the model across different prediction types.
Each model was first evaluated using a 5x layered cross-validation approach, maintaining the class distribution within each fold. This provided a strong assessment of the model's generalization capabilities. The model was then evaluated with a clear 20% holdout test set, and identical metrics were calculated to measure actual predictive effects.
$$ \:surcocy = \frac {tp+tn} {tp+tn+fp+fn} \:$$
(1)
$$ \:precision = \frac {tp} {tp+fp} \:$$
(2)
$$ \:recall \:\left(sensitivity \right)= \frac {tp} {tp+fn} \:$$
(3)
$$ \:f1-score = \frac {precision \:\times \:\:recall} {precision \:+\:recall} \:$$
(4)