Physics-informed machine learning digital twin framework for prostate cancer
We present a computational framework for prostate cancer in digital twins, combining a physics-based model for tumor growth with a machine-learning model that regulates its growth dynamics (Fig. 1). This patient-specific digital twin of the prostate is generated from the T2-weighted image (T2WI) sequences on magnetic resonance imaging (MRI), which include Diffusion Weighted (DW) and Dynamic Contrast Enhanced (DCE) image sequences, as well as the tumor segmented by expert radiologists from HULAFE, all common procedures in the evaluation and diagnosis of prostate cancer (Fig. 1a). Using these MRI sequences, a 3D voxelized geometry is generated, containing the patient’s data such as the cellularity c(x, t), the spatial distribution of vascularization considered through ktrans(x), which provides crucial information about tissue perfusion and permeability, and the tumor binary mask Tmask(x, t) (see Imaging and postprocessingMRI sequences in Methods for further details).
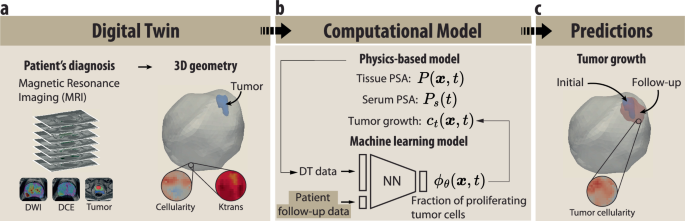
a The digital twin geometry is reconstructed from the T2-weighted image sequences on magnetic resonance imaging, including the spatial distributions of cellularity from the apparent diffusion coefficient maps in the diffusion-weighted images, ktrans from dynamic contrast-enhanced sequences, and the tumor mask. b The computational model consists of two main parts. First, a physics-based model simulates the evolution of tissue PSA P(x, t), serum PSA Ps(t), and tumor growth ct(x, t). Second, a machine learning model based on a neural network (NN) determines the fraction of proliferating tumor cells ϕθ(x, t) in the tumor growth equation based on the data from the digital twin and the patient follow-up serum PSA blood test. c The outcome of the model is the patient’s tumor growth from diagnosis to the follow-up date.
We propose a physics-informed machine learning model to simulate the growth of prostate cancer (Fig. 1b). The purpose of the physics-based model is to represent the biological processes involved in prostate cancer within the digital twin framework to predict the growth of the diagnosed tumor. In this model, we simulate the tissue PSA per unit volume of prostatic tissue P(x, t) as a consequence of the PSA leakage from cancer cells. Additionally, we incorporate the exchange of PSA between the tissue and the bloodstream. To achieve this, we consider the capillaries located in the tissue, represented through ktrans(x). This exchange depends on the concentration difference between the tissue PSA distribution (P(x, t)) and the serum PSA (Ps(t)), as well as the permeability of the capillaries. We also consider the natural decay of the tissue PSA. The serum PSA (Ps(t)) is obtained by integrating the flux between the bloodstream and the tissue, also considering its natural decay. Finally, we model the evolution of the concentration of tumor cells ct(x, t), which represents tumor growth. This tumor cell concentration is responsible for the production of tissue PSA, and therefore, for the variations in the simulated serum PSA concentration. Hence, the dynamics of tumor growth cause the variations in serum PSA (see Physics-based model for prostate cancer in Methods for details of the physics-based model).
In this physics-based model, we integrate a machine-learning model to determine the dynamics of tumor growth (ct(x, t)) based on the patient’s serum PSA tests from their follow-ups. This machine learning model approximates, through a fully connected neural network (NN), the fraction of proliferating tumor cells ϕθ(x, t), which is incorporated into the equation describing tumor growth (see next section for further details). To accomplish this, the deep-learning model receives data from the physics-based model and the patient follow-up data based on serum PSA blood tests. Consequently, it regulates the tumor growth dynamics in the physics-based model so the simulated PSA accurately reproduces the patient’s serum PSA values. Finally, we predict the tumor’s growth from the time of diagnosis to the follow-up date which is responsible for the observed serum PSA variations.
Machine learning model for tumor growth dynamics
We propose a deep-learning algorithm to determine tumor growth by regulating the fraction of proliferating tumor cells (ϕθ(x, t)) in Eq. (3) (Fig. 2). To approximate this function that controls tumor growth, we employ data from the digital twin and the patient. First, for each voxel of the digital twin, we extract the following data: cellularity (c(x, t)), normalized tissue PSA (\(P({\boldsymbol{x}},t)/\max (P({\boldsymbol{x}},t))\)), normalized ktrans (\({k}_{{\rm{trans}}}({\boldsymbol{x}})/\max ({k}_{{\rm{trans}}}({\boldsymbol{x}}))\)), the tumor mask (Tmask(x, t)), and the fraction of proliferating tumor cells (ϕθ(x, t)). Second, we use as patient data the serum PSA measured in the follow-up blood test (\(({P}_{{s}_{t}})\)), which we aim to replicate. This is compared to the current simulated PSA at the time the NN is called (Ps), and normalized as \(({P}_{{s}_{t}}-{P}_{s}
Source link




