Study design and participants
This study was granted ethical approval by the institutional review board of The University of Texas MD Anderson Cancer Center (MDACC), Mayo Clinic, and Dana-Farber Cancer Institute (DFCI). All procedures were conducted in accordance with ethical standards of the 1964 Declaration of Helsinki. Informed consent was waived due to the retrospective nature of this study.
The objective of this retrospective study was to use machine learning to build a treatment recommendation system that enabled individualized treatment plans for systemic therapy (i.e., immunotherapy with or without chemotherapy). Patients from MDACC, Mayo, and SU2C were utilized for model discovery and tuning of hyperparameters, which was subsequently subjected to external validation using patients from the DFCI cohort. These cohorts are briefly described below.
MDACC cohort—We queried GEMINI, at the University of Texas MD Anderson Cancer Center, funded by the Lung Cancer Moon Shot program, to find patients treated with ICIs between 2020 to 2022. Patients were included if they met the following criteria: (1) pathologically confirmed NSCLC, including adenocarcinoma, squamous cell carcinoma, and others; (2) stage IV disease at the time of commencing ICIs; and (3) received at least 2 cycles of ICI alone with or without chemotherapy.
Mayo cohort—The same inclusion criteria were applied to a cohort of patients treated with ICIs at Mayo Clinic. This cohort constituted patients who were seen at Mayo Clinic in Florida between 2019 and 20208.
DFCI cohort—The same inclusion criteria were applied an external cohort of patients treated with ICIs at Dana-Farber. This cohort constituted consecutive patients who were seen at Dana-Farber Cancer Institute, consented to correlative research protocols 02-180, 20-000 and 17-000 and received at least one dose of PD-(L)1 blockade with or without chemotherapy between 2013 to 2023. Details of this data are published elsewhere26,27.
SU2C cohort–This was a public consortium of 393 patients with advanced NSCLC across nine cancer centers treated with PD-1/PD-L1 agents. This cohort was predominantly adenocarcinoma patients, and details of this data are published elsewhere28.
Genomic profiling
We collected somatic sequencing results from genomic profiling or external vendors from pathology reports and clinical notes. Samples with in-house somatic sequencing data were included for extended genomic analyses. Mutational profiling was conducted on formalin-fixed paraffin-embedded tumor tissue or blood samples as previously described29,30. For the MD Anderson data, tissue molecular profiling used NGS-based analysis to detect mutations in 134 or 146 genes. Sequencing or circulating tumor DNA (ctDNA) was performed using the MD Anderson liquid biopsy panel (70 genes) or the Guardant360 panel (74 genes). For computational power, samples from any time point were included, and a mutational event was considered present if it was detected in tissue and/or in the blood. The sample closest to the date of ICI initiation was selected for patients with multiple samples across different time points. For consistency in analysis across sequencing panels, we limited the genomics set to the 70 genes contained in the ctDNA panel. All reported non-synonymous mutational events were included; copy number alterations with log2 copy ratio<1 (deletion) or >5 (amplification) were included in the analysis. Tumor mutational burden was excluded due to smaller sequencing panel size. Further details are described elsewhere8. Similar genomic event filtering was used in the Mayo, SU2C, and DFCI datasets. In the end, 28 overlapping genes across different cohorts were kept for downstream modeling analysis.
Individualizing treatment plans
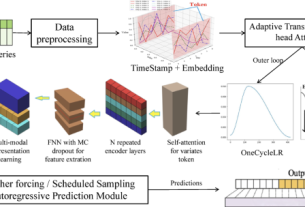
Figure 1 summarized the overall pipeline to characterize heterogeneity in treatment response and enhance customized treatment strategies regarding the addition of chemotherapy to baseline immunotherapy (or lack thereof). We integrated 28 genomic features with clinical risk factors including gender, age, tobacco exposure, pathology, PD-L1 expression, and line of therapy as the foundation of the framework. The computational system encompassed several key steps. First, the pooled data were pre-processed and quality checked. Details of genomic profiling were done per prior reports8,26,27,28. To address retrospective biases and confounding, propensity score matching was performed to identify paired samples with similar characteristics (Supplementary Note 1). Subsequently, an optimal subset was identified through recursive feature elimination for model construction. The individualized benefit of adding chemotherapy was estimated through repeated cross-validation (n = 30) using five different weighting methods (Supplementary Note 3). When the stand-alone scoring’s predictive performance did not achieved the desired performance during validation, we have introduced different strategies to integrate them (H-STEP, S-STEP, and A-STEP, Supplementary Note 4) to mitigate the potential biases of any single model by leveraging the diversity of different approaches to capture wider patterns in the data. Similarly, the integrated system subjected to performance validation in both the discovery and external validation cohort to ensure reliability and generalizability. Finally, we investigated the model’s decision-making through the lens of clinical translation and model interpretation.
Step 1: Modeling treatment effect
In this work, we made standard assumptions shown to be sufficient for causal dependencies on the interventions (details in Supplementary Note 2). Without loss of generality and according to Rubin’s potential outcome framework31, the expected outcome \(E\) can be expressed given the desired outcome \(\left(O\right),\) treatment \((T)\), and initial covariates \(C\) as \(E\left[O | T,C\right]={{\varnothing }}\left(C\right)+T\times \triangle \left(C\right)/2\), where ∅(C) is a function that represents the main prognostic effect and can be calculated as \(E\left[O | T=1,C\right]+E\left[O | T=0,C\right]\) and \(\triangle \left(C\right)\) indicates the treatment effect which can be expressed as \(E\left[O | T=1,C\right]-E\left[O | T=0,C\right]\). To construct a model for \(E\left(O | T,C\right)\) involves formulating functions for both \({{\varnothing }}\left(C\right)\) and \(\triangle \left(C\right)\) based on the covariates. Variables used in constructing \({{\varnothing }}\left(C\right)\) are referred to as prognostic variables, while those used in constructing \(\triangle \left(C\right)\) are called treatment moderators24,32. Conventional methods for creating optimal ITRs involve \({{\varnothing }}\left(C\right)\) and \(\triangle \left(C\right)\) simultaneously predicting outcomes and estimate ITEs using these model-based estimates. However, this approach requires accurate specification of both \({{\varnothing }}\left(C\right)\) and \(\triangle \left(C\right)\), even though only the latter is ultimately used to guide treatment selection. As a result, \({{\varnothing }}\left(C\right)\) becomes a nuisance parameter, and its specification may impact the estimation of the treatment function \(\triangle \left(C\right)\), which is particularly problematic because there are often many prognostic variables but only a few treatment moderators that influence treatment recommendations10,33. Nevertheless, if our aim is to rank ITEs or develop ITRs, only the treatment function \(\triangle \left(C\right)\)’s ranks or signs would be relevant. Therefore, it is desirable to have a robust estimate of ITEs that does not require the estimation of \({{\varnothing }}\left(C\right)\)9,10.
The endpoint in this study was progression at 3 months (denoted as \({O}^{(1)}\)) or no progression at 3 months (denoted as O(0)). The two treatments for comparison were ICI-Chemo, denoted as T=1, or ICI-Mono, denoted as T=0. Additionally, we assumed that only a single treatment outcome, either (O(1) or O(0), can be observed per patient. We also assumed that T is not dependent on (O(1)\(,\)O(0))31,34. To assign treatment, we assumed that the probability of receiving treatment, given certain covariates \(C\), is represented by the propensity score π(\(C\)), calculated as \(\pi \left({C}_{i}\right)=\Pr \left(T=1 | C\right)\). In randomized trials, this score is typically known and not influenced by \(C\), but it requires estimation in observational studies. The data we observe is represented by \(n\) independent and identically distributed copies of \(\left(O,T,C\right),\) denoted as \(\{({O}_{i},{T}_{i},\,{C}_{i}),\,i=1,\,.\,.\,.,\,n\}\).
Step 2: Individual benefit score estimation through weighting method
The objective was to utilize the weighting subgroup identification method9,10 to develop a personalized benefit scoring system \(f(C)\) based on the covariates \(C\), in which customized treatment would be suggested for individual patients using \(f(C)\). To clarify, we defined a benefit score as any function \(f(C)\) that met two criteria: (i) it reflects the extent to which patients benefit from a certain treatment and (ii) it has a meaningful and known cut-point value \(c\), such that for a given level of covariates \(C\), \(f\left(C\right) < c\) indicates that ICI-Chemo is more effective than ICI-Mono, and \(f\left(C\right)\ge c\) indicates the opposite. Clearly, \(\triangle \left(C\right)\) can serve as a benefit score, as it represents the expected benefit a patient will derive from ICI-Chemo in terms of their outcome. Thus, estimating \(\triangle \left(C\right)\) or its sign can enable optimal recommendation of different treatments for different patient subgroups. By definition, \(\triangle \left(C\right)\) can also be used to rank patients based on the magnitude of treatment effect9,10.
Another measure of interest is ITR, a function that maps patient covariates to the treatment decision, \(d\left(C\right):C\to T\). Optimal ITRs aim to maximize the average outcomes across the population by making treatment decisions for patients in a way that maximizes the value function \(V\left(d\right)={E}^{d}\left(O\right)=\int {Od}{P}^{d}\) where \({P}^{d}\) is the distribution of \(\left(O,T,C\right)\), given \(T=d\left(C\right).\) As a result, \(\triangle \left(C\right)\) can be used to develop optimal ITRs. Specifically, the sign of \(\left\{f\left(C\right)\right\}\) is considered for optimal ITRs9,10. A convex loss function \(M\left(y;v\right)\) can be used to estimate benefit scores, for example the squared error loss, \(M\left({y;v}\right)={(y-v)}^{2}\). Chen et al.10 originally proposed that \(M\left({y;v}\right)\) should meet two conditions. First, \({M}_{v}\left({y;v}\right)=\,\partial M\left({y;v}\right)/\partial v\) is increasing in \(v\) for every fixed \(y\). Second, \({M}_{v}\left({y;}0\right)\) is monotone in \(y\). These conditions are adequate for Fisher consistent subgroup identification, but they are not mandatory9. Our formulation of different weighting methods to estimate benefit scores and subsequently identify subgroup effects is presented in Supplementary Note 3.
Step 3: Integrating the individual sub-scoring systems
Using the benefit scores from five distinct scoring systems, we further built an ensemble model using three fusion mechanisms. The impetus was to capture common and potential complimentary signals presented across different stand-alone systems while reducing the individual system’s bias and variance by aggregating across multiple model predictions. The assembling steps are detailed in Supplementary Note 4.
Step 4: Model training and validation
We adopted a repeated 5-fold cross-validation strategy (n = 30) to develop and fine tune sub-scoring models’ hyperparameters including the feature importance based on discovery cohort. Features were ranked based on their frequency of being selected from 150 runs and were considered informative if selected at least 50% of the time. We then locked the optimized hyperparameters and refitted the individual models on the whole discovery set, which were further tested on an external validation cohort. All models were developed using open-source Python version 3.9.8 and R version 3.6.1. All models were trained on the NVIDIA DGX A100 station.
Clinical validation
To assess the predictive capability of sub-models and integrated models in estimating the benefit of adding chemotherapy to immunotherapy for a patient, an individualized recommendation was made by contrasting two treatment arms, in terms of treatment effect conditional on the subgroup. The personalized benefit score was utilized to recommend whether chemotherapy was beneficial for a patient. Using this recommendation, patients were stratified into two subgroups, (i) those who were treated according to the model recommendation, or “Follow-recommendation’’, and (2) those who were treated against the recommendation, or “Anti-recommendation”. Subsequently, we investigated clinical significance by conducting a comprehensive analysis by comparing the “Follow” versus “Anti” subgroups in terms of effect measures particularly the weighted risk reduction (WRR), Kaplan–Meier plots, log-rank tests, and hazard ratios (HRs). In brief, WRR represents the percentage of patients potentially saved from early disease progression by adhering to model’s recommendations (details in Supplementary Table 3).
Model interpretation
Finally, we interpretated the association between individual features with a model’s recommendations (ICI-Chemo vs ICI-Mono) using SHAP (SHapley Additive exPlanations) analysis35 to quantify the impact of each feature on both direction and magnitude of the treatment effects.
Statistical analysis
The primary endpoint was defined as early disease progression at 3 months. Disease progression was determined by the recorded assessment of the attending physician based on imaging reports of tumor growth or new disease sites, pathologic conformation, and/or through clinical assessment. We also collected PFS data, where patients who were alive without progression were censored at their last image assessment. The interaction p-value between the recommended and treatment groups was calculated via ANOVA. Kaplan-Meier analysis and log-rank tests were used to evaluate statistical significance of patient stratification. The Wilcoxon signed-rank test or χ² test were used to test the differences in distribution of continuous or categorical variables, respectively, as stratified in different groups. To adjust for multiple statistical testing, the Benjamini-Hochberg method was used to control the false discovery rate (FDR). All statistical tests were two-sided, with a p value less than 0.05 or FDR less than 0.1 considered to be statistically significant. All statistical analyses were done with R version 3.6.1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.