image:
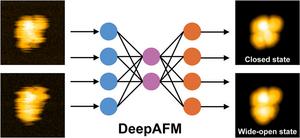
DeepAFM is a deep learning-based method for analyzing high-speed atomic force microscopy images of proteins. Eliminate noise to identify protein shapes and enable accurate detection of transitions between closed and wide open states during protein function.
view more
Credit: Associate Professor Ryuji Mori, Tokyo University of Science Image source link: https://pubs.acs.org/doi/10.1021/acs.jcim.6c00142
In 2018, an artificial intelligence (AI) program called AlphaFold made significant progress by winning first place in the Structural Prediction Critical Assessment, a competition to predict the three-dimensional structure of proteins. It scored nearly 90 out of 100 for moderately difficult targets, marking a turning point in the use of AI to understand protein structure and highlight its potential applications. Although it has been a major advance in predicting the structure of proteins, proteins in biological systems are not fixed. They constantly move, change shape, and interact with other molecules. AI is now playing a role in supporting this.
Traditionally, determining the different shapes that proteins take has required fitting known three-dimensional structures to two-dimensional high-speed atomic force microscopy (HS-AFM) images, which allows us to capture proteins in action at the single-molecule level. However, HS-AFM images are noisy and can be distorted by the line-by-line scanning process, where each part of the image is recorded at slightly different times. This time delay and background noise make it difficult to determine the exact shape of the protein at any time.
Associate Professor Ryuji Mori from the Department of Chemistry, Faculty of Science, Tokyo University of Science (TUS) explains the problems with current approaches. “Due to the presence of noise in the images, these methods can lead to overfitting, and the model may capture noise-induced artifacts or false details rather than the true structural features of the protein.”
To address this challenge, his team developed a deep learning-based method called DeepAFM. This technique aims to reduce noise in HS-AFM images and accurately estimate the different shapes that proteins adopt as they move and function.
The team included Katsuki Sato of the University of Tokyo, who completed his master’s program in 2025, and Dr. Takayuki Uchihashi and Dr. Yui Kanaoka of Nagoya University. and Dr. Tomoya Tsukasaki of Nara Institute of Science and Technology. The study was published online. Journal of Chemical Information and Modeling Published on April 4, 2026 and published in Volume 66, Issue 8 on April 27, 2026.
In this approach, the team creates a dataset of synthetic HS-AFM images representing different protein shapes from molecular dynamics simulations. Each image is labeled according to the 3D structure of the corresponding protein. These simulated images include both ideal versions without noise and more realistic versions that incorporate experimental effects such as background noise, scan distortion, and Brownian motion.
The researchers trained DeepAFM on a protein called SecA that can switch between a closed state and a wide open state. They used molecular dynamics simulations to generate a wide range of possible protein shapes, which they used to create millions of synthetic HS-AFM images. This dataset was then used to train a deep learning model that can remove noise from HS-AFM images and identify the underlying protein shape.
When we tested this method, it produced a denoised image that closely matched the ground truth, with an error on the order of 0.1 nm. In addition to improving image quality, AI was able to accurately classify the conformational state of the protein. Across 800,000 test images, the model correctly identified the correct state among 19 possible conformations with 93.4% accuracy. This accuracy was further improved when small tolerances were taken into account. Importantly, when applied to experimental HS-AFM images, the AI estimated protein conformational states in agreement with independent experimental measurements, demonstrating its utility.
“DeepAFM provides a new deep learning-assisted strategy for analyzing noisy HS-AFM data, facilitating the study of protein dynamics.” says the associate professor. Mr. Mori.
The researchers also demonstrated that the method can be extended to other protein systems using transfer learning, which applies knowledge from one system to another. This suggests that DeepAFM could become a widely useful tool for studying a wide range of biomolecules.
The initiative, developed by the RIKEN Center for Computational Science in collaboration with Fujitsu and NVIDIA, is part of a broader effort to advance AI-driven research in preparation for next-generation computing platforms such as Fugaku NEXT, which is scheduled to be operational around 2030.
***
reference
DOI: 10.1021/acs.jcim.6c00142
About Tokyo University of Science
Tokyo University of Science (TUS) is Japan’s largest private scientific research university with four campuses in central Tokyo, its suburbs, and Hokkaido, and is a well-known and well-regarded university. Founded in 1881, the university has continued to contribute to the advancement of science in Japan by instilling a love of science in researchers, engineers, and educators.
Our university engages in a wide range of research from basic science to applied science, with the mission of “creating science and technology for the harmonious development of nature, humans, and society.” TUS takes a multidisciplinary approach to research and has conducted intensive research in some of today’s most important areas. TUS is a meritocracy, where the best science is recognized and fostered. It is a private university in Japan that has produced Nobel Prize winners, and is the only private university in Asia that has produced Nobel Prize winners in the natural sciences.
Website: https://www.tus.ac.jp/ja/mediarelations/
About Associate Professor Ryuji Mori, Tokyo University of Science
Ryuji Mori is an associate professor at the Department of Chemistry, Faculty of Science, Tokyo University of Science. He holds a Doctor of Science degree from Nagoya University and has over 15 years of experience in molecular simulation of biomolecules. His research focuses on theoretical and computational chemistry, integrating molecular dynamics and artificial intelligence to study the structure and dynamics of biomolecules. He has authored 42 peer-reviewed publications and numerous presentations. His research spans life sciences, drug discovery, and environmental chemistry, with a focus on understanding molecular function at the atomic level through advanced computational approaches.
Funding information
This research was supported by the JSPS KAKENHI (grant number JP25K02226 Mr. Tomoya Tsukasaki, grant number JP24K01309 Mr. Takayuki Uchihashi, grant number JP21H05157, JP24K03035, JP25H01329 Mr. Ryuji Mori), the Japan Scientific Research Grant K Program (JPMJKP23H2) This project was supported by Tomoya Tsukazaki). Japan Science and Technology Agency (JST), Joint Usage and Research System Development Promotion Project: Program for Excellence in Collaborative Research with Universities (CURE) (Takayuki Uchihashi Grant Number JPMXP1323015482), Ministry of Education, Culture, Sports, Science and Technology’s “Supercomputer Fugaku Research Promotion Project” (Development and Application of Inference Based on Large-Scale Simulations of Biomolecules JPMXP1020230119 to Takaharu Mori), HPCI Project (hp230209, hp240215, hp240277, hp250233, and hp250280 Takaharu Mori), Computational Science Research Center (Okazaki City) (Project: 25-IMS-C151 Takaharu Mori).
journal
Journal of Chemical Information and Modeling
Research method
Computational simulation/modeling
Research theme
not applicable
Article title
Estimation of protein conformational state from high-speed AFM images using molecular dynamics and deep learning
Article publication date
April 27, 2026
Conflict of interest statement
The authors declare that they have no competing financial interests.