image:
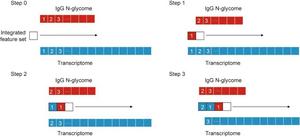
A typical forward feature selection (FFS) workflow for the AlphaSnake algorithm.
view more
Credit: Yao Xia, Syed Mohammed Shamsul Islam, et al.
The research team developed a new biological aging clock, called gtAge, by integrating the immunoglobulin G (IgG) N-glycome and blood transcriptome using a multi-omics integration method with deep reinforcement learning called AlphaSnake, according to a study published in . engineering. This study provides a new approach to estimate biological age by combining complementary molecular signatures that change with age.
This study points out that both the IgG N-glycome and transcriptome serve as potential biochemical markers of chronological and biological age, but few attempts have been made to construct aging clocks using integrated multi-omics data. To address this, researchers established gtAge based on data from 302 people with a mean age of (56.98 ± 5.23) years in the Busselton Healthy Aging Study cohort. They employed minimum angle regression within a bootstrap-based feature selection framework to identify predictive features from each omics layer, and then designed AlphaSnake, a deep-Q network-based agent, to dynamically select the most informative features across the two omics sets during forward feature selection.
Performance evaluation using 10-fold cross-validation shows that AlphaSnake-derived gtAge achieved coefficient of determination R²/0.853, which outperforms traditional concatenation-based integration methods. R0.820 of ². The gtAge model explained up to 85.3% of the variance in chronological age and outperformed age prediction from IgG N-glycome alone. This is called gAge. R² is 0.290, referred to as tAge from the transcriptome only. R0.812 of ². The final model after feature refinement contained 144 features, including 137 genes and 7 glycan traits, and removing glycan features significantly degraded the predictive performance, confirming the contribution of the IgG glycosylation signature.
Further analyzes considered associations between delta age, the difference between predicted and chronological age, and age-related phenotypes. Both delta gtAge and delta tAge are negatively correlated with high-density lipoprotein; p Delta gAge was positively correlated with total cholesterol, triglycerides, fasting plasma glucose, low-density lipoprotein, and glycated hemoglobin, although the values were 0.02 and 0.022, respectively. p The values are 0.006, 0.002, 0.014, 0.006, and 0.039, respectively. These associations suggest that gtAge, tAge, and gAge carry complementary information about biological aging beyond chronological age.
The researchers also conducted feature importance analysis using SHAP values and pathway enrichment analysis. Genes in the tAge model were enriched in immune and inflammatory pathways such as chemokine activity and natural killer cell-mediated immunity, whereas the integrated gtAge model contained important immunoregulatory genes that support the role of immunosenescence and chronic inflammation in aging. The results of this study demonstrate that multi-omics integration captures a more comprehensive molecular landscape of aging, and AlphaSnake provides a flexible framework for feature selection of high-dimensional, heterogeneous omics data beyond aging research. This study is based on a midlife cohort and requires future validation in a larger and more diverse population to increase generalizability.
The paper, “Deep reinforcement learning-driven multi-omics integration to build gtAge: A new aging clock from IgG N-glycome and blood transcriptome,” is authored by Yao Xia, Syed Mohammed Shamsul Islam, Xingang Li, Abdul Baten, Xuerui Tan, and Wei Wang. Full text of open access article: https://doi.org/10.1016/j.eng.2025.08.016. For more information, engineeringvisit the website https://www.sciencedirect.com/journal/engineering.
Article title
Deep reinforcement learning-driven multi-omics integration to build gtAge: a new aging clock from IgG N-glycome and blood transcriptome
Article publication date
February 17, 2026
Disclaimer: AAAS and EurekAlert! We are not responsible for the accuracy of news releases posted on EurekAlert! Use of Information by Contributing Institutions or via the EurekAlert System.