David Ussery, professor of biomedical and information science at UAMS, and his student Aakash Bhattacharyya, discuss using machine learning methods to predict the pathogenicity of bacterial infections based on genome sequencing
The bacterial genus Salmonella is a common source of food-borne development, infecting more than 1 million people each year. In most cases, the illness is short and recovery occurs within a few days. However, Salmonella infections result in more than 25,000 hospitalizations, resulting in approximately 400 deaths in the United States each year. To promptly direct appropriate treatment, it is necessary to quickly determine whether a salmonella infection may be serious.
Salmonella was first described by the Lignieries in 1901 as “Le Microbe du Hog-Cholera de Salmon” (1,2), and was named after Daniel Elmer Salmon, an American veterinary surgeon. There are over 2500 types of salmonella (“serotypes”). Historically, each serotype was named as a species (1) However, in 1987 it was reduced to one species (Salmonella enterica). (2) Last revised in 2005 (3) Added another species (S. boneri).
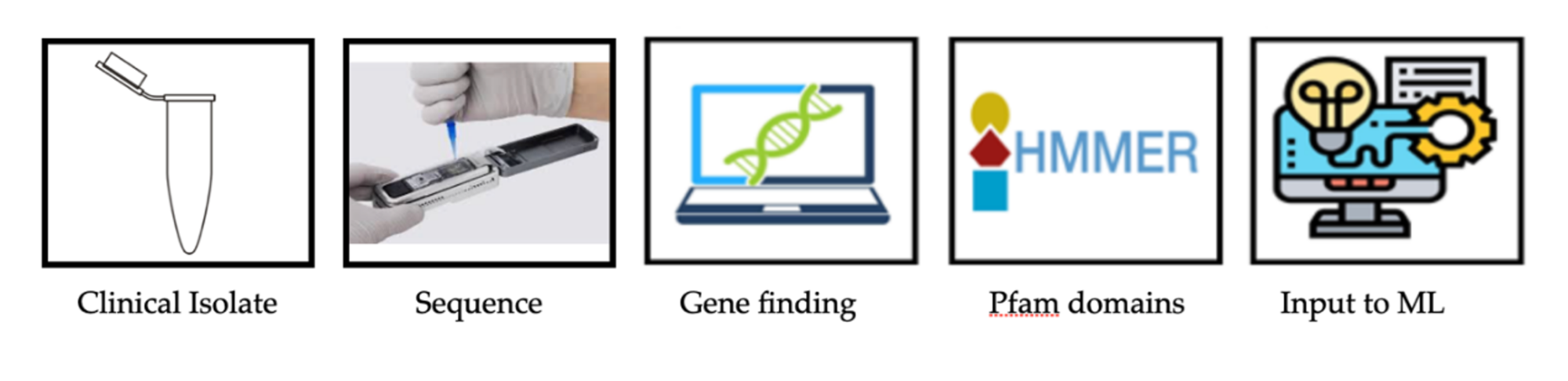
It is now possible to predict the severity of Salmonella infection based on the genomic sequence of clinical isolates using high-throughput calculation methods. As a rule, as shown in the diagram above, it is possible to transfer from the sample to the genomic sequence and predict severity levels within a few hours.
Disruptive changes in AI and sequencing technology
The first single molecule or “third generation” sequencing machine was introduced in 2014, dramatically reducing the cost and time required to obtain bacterial genomic sequences. This technology allows for much longer reads. For bacterial genomes, approximately 20,000 nucleotides (NT) per read are routinely obtained, corresponding to approximately 20 genes. Careful sample preparation also allows for approximately 1 million nt reads to be obtained from human chromosomal DNA. Oxford Nanopore Flow Cells arrange single molecules by measuring current changes as a single strand of DNA passes through small pores about 1 nm (10 atoms) wide. This change in current is converted into sequences with a variety of machine learning methods, such as artificial neural networks trained with known sequences. (4,5) Includes modified bases, (6) 5MC etc. This disruptive technology allows for quick and inexpensive sequencing, allowing newer versions of flow cells (R10.4.1) to read DNA fragments. (7) Long reads can be used to fully assemble and assemble the bacterial genome and plasmids with sequences in a few hours to improve the quality (and amount) of the sequenced genome. As mentioned in my previous profile article, (8) As of December 2024, approximately 1.2 million Salmonella genome sequences were compared. Over the past five months, another 200,000 Salmonella genomes have been available (as of May 2025), and the number continues to grow rapidly.
Computational analysis of proteomes
This raises the question of how to rapidly analyze literally millions of genome sequences, particularly aimed at helping doctors determine whether certain strains of salmonella isolated from patients can cause severe, fatal diseases. Genomic sequences can be stored as four letters (GATC) strings representing the four bases of DNA, but digital computers use numbers and simply compare characters is not enough. In living salmonella cells, the genome is transcribed into RNA, most of which encode proteins. “Information” is in the protein sequence, but again, can be thought of as a mere string of 20 characters representing each amino acid. How is it calculated with a set of characters? The basis of this dates back to the 1960s by the creation of protein sequences and structural atlas. (8,9) It was eventually part of the Uniprot database. (10)
Profile HMM to abstract proteome
We have previously explained how to use profile HMM to find functional domains such as PFAM domains. (11) Within all the proteins in the genome, they are used to quickly extract a set of specific proteins (such as Sigma factors) from thousands of genomes in thousands of seconds. These PFAM domains can be used to search for enrichment of genomes known to cause pathogenicity, and this information serves as an input for machine learning methods.
Finally, we identified a set of PFAM domains that could be used as biomarkers to accurately predict case severity in 93% of the test set. The method explained here is just one of many examples. At the time of writing (May 2025), there are over 200 articles on PubMed when searching for “Salmonella Machine Learning.” The number of publications could increase rapidly as labs around the world continue to apply machine learning methods to study the salmonella genome and pathogenesis.
The projects described here are accepted for publication – new references: Bhattacharyya A, Panday S, Ussery D. Protein family domain analysis and rapid assessment of clinical severity of Salmonella cases via machine learning. Academia Molecular Biology and Genomics 2025;Volume. https://doi.org/10.20935/xxx
reference
- John-Brooks, E. St. , (1934). “Genus Salmonella Lignieres, 1900,” Journal of Hygiene (London). 34 (3): 333-350. doi:
- Le Minor, L. , & Popoff, My, (1987). “Salmonella entéricasp. no. nom. Rev. designation, as a type and only species of the genus Salmonella,” International Journal of Systematic Bacteriology, 37:465-468. doi:
- Tindall B.J., Grimont Pad, Garrity G.M., Euzeby JP, (2005). “Nomenclature and Taxonomy of the Genus Salmonella,” International Journal of Systematic Bacteriology, 55:521-524. doi:
- Bao Y, Wadden J, Erb-Downward JR, Ranjan P, Zhou W, McDonald TL, Mills RE, Boyle AP, Dickson RP, Blaauw D, Welch JD, (2021). “Squigglenet: Real-time, Direct Classification of Nanopore Signals”, Genome Biology, 22(1):298. doi:
- Hall MB, Wick RR, Judd LM, Nguyen AN, Steinig EJ, Xie O, Davies M, Seemann T, Stinear TP, Coin L, (2024). “Benchmarks reveal the advantages of deep learning variant callers over bacterial nanopore sequencing data,” Elife, 2024;13. doi:
- Takaguchi S, Takeuchi N, Shensin V, Jains G, Genot AJ, Nivara J, Ronderes Y, Kawano R (2025). “Utilizing DNA computing and nanopore decoding for practical applications: from informatics to microRNA targeting diagnosis,” Chemistry Society Review, 54(1): 8-32. doi:
- Kim Bai, Gerato HR, Church SH, Suvorov A, Anderson SS, Barmina O, Beskid SG, Comere AA, Crown KN, Diamond SE, Doras S, Fujichika T, Hemka, Furusek J, Kankare M, Kato T, Magnacca KN, Martinla M, Simoni S, SteenWinkel TE, Syed ZA, Takahashi A, Wei KH, Yokoyama T, Eisen MB, Kopp A, Matute D, Obbard DJ, O'Grady PM, Price DK, Toda MJ, Werner T, Petrov DA, (2024). “Single fly genome assembly fills the major phylogenetic gaps across the Drosophila family trees of life,” PLOS Biology, 22(7): E3002697. epub20240718. doi:
- Open Access Government, January 2025, pages 48-49
- Strasser BJ. Collecting, (2010). “Comparison and Computing Sequence: Creating a Protein Sequence and Structure Atlas by Margaret O. Deihoff, 1954-1965”, Journal of the History of Biology, 43(4): 623-660. doi:
- Palmblad M, Hoopmann MR, Dorfer V, (2025). “Special Software Issues to Celebrate Margaret Deihoff's 100th Birthday,” Journal of Proteome Research, 24(3): 977-978. doi:
- Uniprot Consortium (2025). “Uniprot: Universal Protein Knowledge Base for 2025”, Nucleic Acid Research, 53 (D1): D609-D17. doi:
- Cook H, Ussery DW, (2013). “Sigma factors in the Thousand E. coli genomes,” Environmental Microbiology, 15 (12): 3121-3129. epub20130829. doi: