Preoperative laboratory findings and tumor biomarker levels
In this study, a total of 297 participants were recruited comprising five groups: healthy control (HC; n = 50), cholangiocarcinoma (CCA; n = 138), gallbladder cancer (GBC; n = 16), hepatocellular carcinoma (HCC; n = 65), and pancreatic ductal adenocarcinoma (PDAC; n = 28). Prior to clinical and peptidome analyses, the samples were divided into a training set (n = 198) and a testing set (n = 99).
The experimental results revealed that liver function parameters (ALT, AST, ALP, bilirubin) and tumor biomarkers (CEA, CA19-9) were markedly elevated in patients with hepatobiliary and pancreatic (HPB) cancers compared to healthy individuals. Specifically, in the training set, median levels of AST and ALT in CCA patients were 32 and 29 U/L, respectively, compared to 21 and 18 U/L in healthy controls. PDAC patients showed the highest median levels of ALT (72 U/L) and AST (66 U/L), with some values reaching over 300 U/L, indicating severe hepatic involvement. Similarly, elevated total bilirubin levels were observed particularly in PDAC patients (median 4.0 mg/dL; range 0.5–31.0 mg/dL), in contrast to healthy individuals (median 0.5 mg/dL; range 0.1–6.4 mg/dL). Albumin levels tended to be slightly lower in cancer groups, reflecting impaired liver synthetic function, although the differences were not statistically significant.
In terms of tumor markers, both CEA and CA19-9 showed wide variation and tended to be higher in cancer groups; however, no statistically significant differences in these markers were found among the various cancer types. For example, median CA19-9 levels were 31.13 U/mL in CCA, 27.51 U/mL in GBC, 23.56 U/mL in HCC, and 25.5 U/mL in PDAC within the training cohort. CEA levels showed a similar pattern of elevation, particularly in CCA and PDAC cases, but again without statistical significance between cancer types.
The testing set exhibited consistent trends with the training set. Elevated liver enzymes and bilirubin levels persisted among HPB cancer patients, especially in PDAC, which had the highest AST (median 94 U/L) and ALT (median 93 U/L) levels. However, comparisons among different cancer types again did not reveal statistically significant differences in these parameters. Collectively, these results suggest that while liver function tests and tumor biomarkers can distinguish cancer patients from healthy individuals, they are insufficient to differentiate between specific cancer types in HPB malignancies (Table 1).
Peptide mass fingerprints for hepato-pancreato-biliary cancer diagnosis
A total of 1,100 peptide features were detected in serum of the training set by MALDI-TOF MS that showed markedly different patterns of peptide mass fingerprints (PMFs) between healthy and HPB cancers (Fig. 1A). The PMF spectra were transformed into expression z-scores and visualized as a heatmap to represent peptide mass fingerprint expression in healthy individuals and cancer patients. A total of 1,100 peptides within the m/z range of 1000–4000 were analyzed. In the heatmap, red indicates upregulated peptide expression, while blue indicates downregulated expression at each corresponding m/z position. The results revealed a clear distinction in PMF expression patterns between the healthy group and HPB cancers (Fig. 1B).
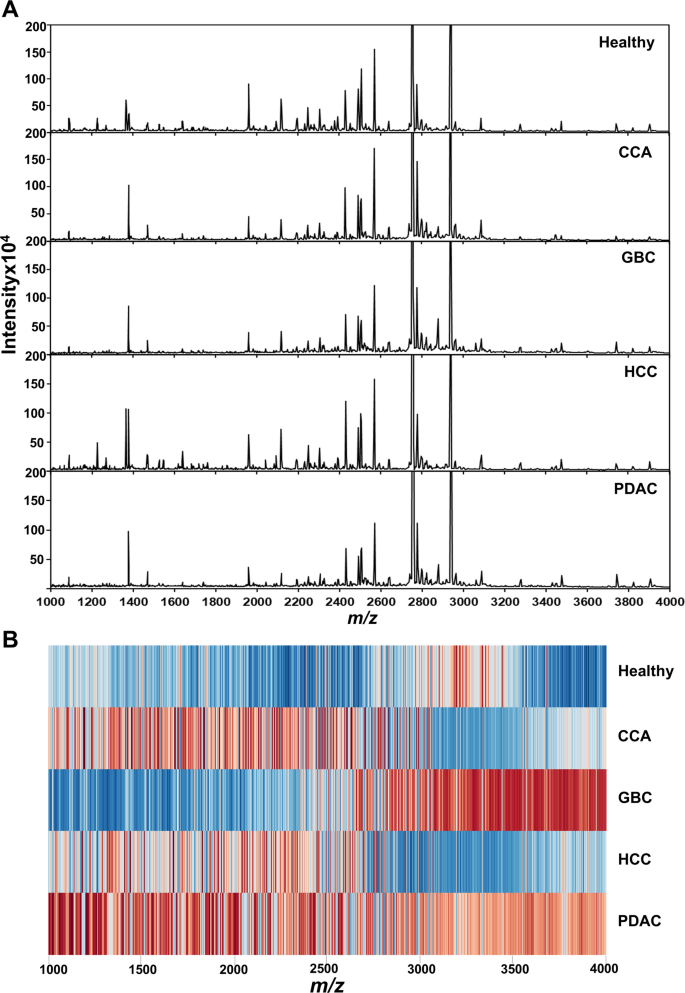
Peptide mass fingerprints (PMFs) of healthy controls and hepato-pancreato-biliary (HPB) cancer patients analyzed by MALDI-TOF MS. (A) Representative MALDI-TOF MS spectra showing markedly different peptide mass fingerprints between healthy individuals and HPB cancer patients within the m/z range of 1000–4000. A total of 1,100 peptide features were detected in serum samples from the training set. (B) Heatmap visualization of PMF expression after transformation into z-scores. Red indicates upregulated peptide expression, while blue indicates downregulated expression. The heatmap reveals a clear distinction in peptide expression patterns between the healthy and cancer groups.
Selection of key peptide mass fingerprints for hepato-pancreato-biliary cancer classification
Global PMF analysis was performed to identify peptide features capable of distinguishing healthy individuals from those with HPB cancers. In the training set, a total of 1,100 peptide peaks derived from MALDI-TOF MS were analyzed using MetaboAnalyst 6.0. Feature selection was carried out exclusively on the training set to prevent information leakage and to maintain the integrity of downstream analysis. Peptides with a variable importance in projection (VIP) score ≥ 1 from partial least squares discriminant analysis (PLS-DA) and statistical significance (p < 0.05) from one-way ANOVA were retained. This resulted in 71 peptides considered as informative features for distinguishing between groups.
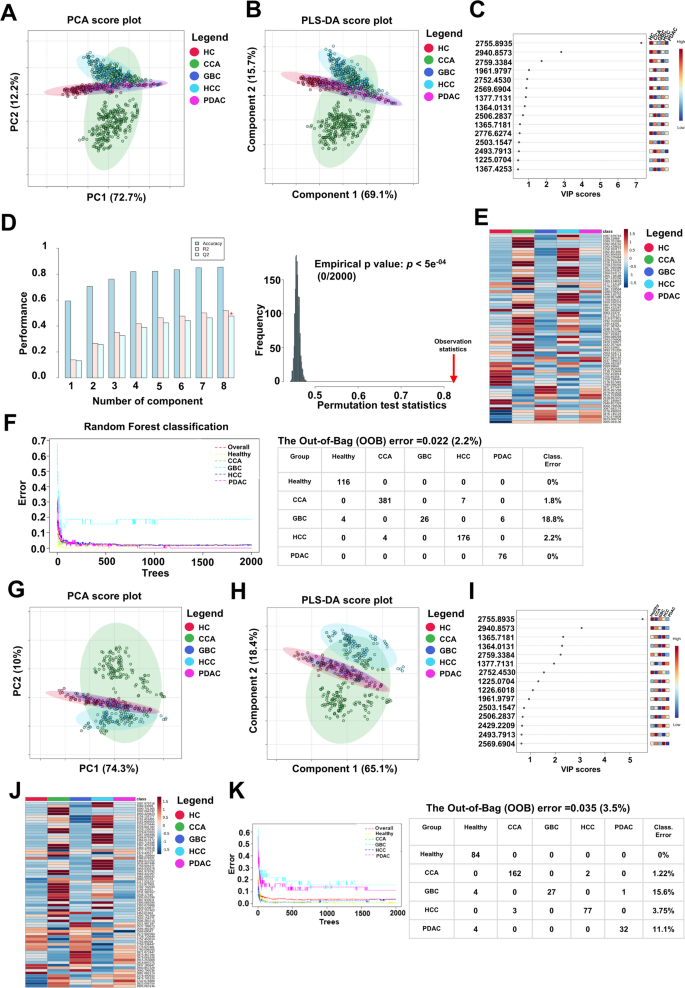
In the training set, principal component analysis (PCA) revealed that the first two components accounted for 84.9% of the total variance (PC1 = 72.7%, PC2 = 12.2%). As illustrated in Fig. 2A, partial separation was observed between the CCA and HCC groups, whereas the healthy control, GBC, and PDAC groups tended to cluster together, suggesting shared peptide expression profiles among these latter groups. Similarly, the PLS-DA, a supervised classification method, demonstrated a comparable clustering pattern. The first two latent variables explained 84.8% of the variation (Fig. 2B), and the group distribution was consistent with the PCA results. Although PLS-DA typically improves group separation due to its supervised nature, in this case, the separation between groups was similar to that observed in PCA, indicating that intrinsic differences among groups were already evident without model supervision. These findings were further supported by the analysis of the global PMF dataset containing 1,100 peptides, which exhibited consistent distribution patterns in both the PCA and PLS-DA score plots (Supplementary Fig. S1A and B). The top 15 peptide features with the highest VIP scores were identified as key contributors to this separation (Fig. 2C). Cross-validation of the PLS-DA model demonstrated strong robustness, with increasing R² and Q² values as additional components were included. The optimal model achieved R² = 0.564 and Q² = 0.502, indicating good explanatory and predictive performance. Furthermore, permutation testing with 2,000 iterations confirmed the absence of overfitting (p < 5 × 10⁻⁴; Fig. 2D).
Multivariate analysis and classification performance based on 71 selected peptide features in the training and testing sets. (A) PCA score plot showing partial separation of CCA and HCC groups, with clustering of healthy control, GBC, and PDAC groups in the training set. (B) PLS-DA score plot demonstrating a comparable group distribution to PCA, based on the 71 selected peptides. (C) Top 15 peptide features ranked by VIP scores from the PLS-DA model. (D) PLS-DA cross-validation results showing optimal model performance (R2 = 0.564, Q2 = 0.502), with permutation testing (N = 2,000) confirming the absence of overfitting in the training set. (E) Heatmap of average peptide expression levels across groups, with peptides ordered by increasing mass. (F) Random Forest model classification results, achieving an out-of-bag (OOB) error rate of 2.2%. (G,H) PCA and PLS-DA score plots in the independent testing set, showing clustering patterns consistent with the training set. (I) Top 15 VIP-ranked peptides in the testing set, 13 of which overlapped with those identified in the training set. (J) Heatmap of average peptide expression levels across groups in the testing set, showing consistent expression trends. (K) RF classification performance in the testing set, yielding an OOB error rate of 3.5%.
The heatmap illustrating the expression patterns of 71 peptides across each group, ordered by increasing mass, based on average peptide expression per group, revealed distinct differential expression profiles (Fig. 2E). Individual peptide expression profiles for each participant were also presented in Supplementary Fig. S2A, highlighting consistent variation both within and between groups.
To further evaluate classification performance, a Random Forest (RF) model was constructed using the 71 selected peptides. This model achieved an out-of-bag (OOB) error rate of only 2.2%, substantially lower than the 5.56% error rate observed when using all 1,100 peptides (Supplementary Fig. S1C). Subgroup classification error rates were similarly low: 0% for healthy controls and PDAC, 1.8% for CCA, and 2.2% for HCC. The only notable misclassification occurred in the GBC group, with an error rate of 18.8% (Fig. 2F).
To assess the robustness and generalizability of the 71 selected peptide features, the same analytical workflow was applied to the independent testing set. Both PCA and PLS-DA score plots (Fig. 2G–H) demonstrated clustering patterns that closely mirrored those observed in the training set. Partial separation was maintained between the CCA and HCC groups, whereas the healthy control, GBC, and PDAC groups continued to cluster more closely, suggesting similar expression trends. These observations support the stability of the peptide-based classification across independent datasets. Among the top 15 features ranked by VIP scores, 13 peptides overlapped with those identified in the training set, further indicating strong reproducibility of discriminative markers (Fig. 2I). Cross-validation and permutation testing in the testing set confirmed the reliability of the PLS-DA model, with no signs of overfitting (Supplementary Fig. S3).
The heatmap of the 71 peptides, based on average expression and ordered by increasing mass, also revealed group-specific expression profiles (Fig. 2J), which was consistent with those seen in the training set. Individual-level peptide expression profiles were presented in Supplementary Fig. S2B, highlighting consistent intra- and intergroup variation.
Furthermore, the RF model demonstrated sustained classification performance in the testing set, achieving an OOB error rate of 3.5% (Fig. 2K). Collectively, these findings reinforce the discriminative power and reproducibility of the 71 selected peptides and underscore their potential utility as peptide mass fingerprint-based biomarkers for distinguishing HPB cancer subtypes from healthy individuals.
Investigation of classification performance in PMFs for hepato-pancreato-biliary cancers diagnosis using support vector machine and random forest models
To evaluate the diagnostic performance of the 71 candidate PMFs, binary classification models—Support Vector Machine (SVM) and Random Forest (RF)—were employed to distinguish healthy individuals from patients with HPB cancers, including CCA, GBC, HCC, and PDAC. These models were constructed using the web-based MetaboAnalyst platform with default settings. The performance evaluation was based on commonly calculated from confusing matrix, including accuracy, precision, recall, F1-score, and the area under the receiver operating characteristic curve (AUC-ROC). In addition, Matthews Correlation Coefficient (MCC) was also calculated to provide a more balanced assessment of classification performance. MCC is particularly valuable in binary classification problems involving imbalanced class distributions, as it considers all four categories of the confusing matrix (true positives, true negatives, false positives, and false negatives) and provides a more balanced measure than accuracy alone. An MCC value of + 1 indicates perfect prediction, 0 indicates random prediction, and − 1 indicates total disagreement between prediction and observation.
In the training set (n = 198), the SVM model demonstrated excellent classification performance, achieving 98.74% accuracy for distinguishing healthy individuals (n = 29) from all cancer cases (n = 169), with a precision of 99.70%, recall of 98.82%, F1-score of 0.99, TNR of 98.28%, MCC of 0.95, and ROC of 0.999. Individual comparisons with each cancer type revealed perfect classification for CCA, GBC, and HCC (MCC = 1.00), and nearly perfect classification for PDAC (MCC = 0.99) (Table 2; Fig. 3A). In the testing set (n = 99), SVM performance remained robust. The model yielded 98.55% accuracy for healthy vs. all cancer cases, with an MCC of 0.97, and ROC of 0.999. Comparisons between healthy individuals and each cancer type also showed high MCC values: CCA (0.99), GBC (0.94), HCC (1.00), and PDAC (0.94) (Table 2; Fig. 3A).
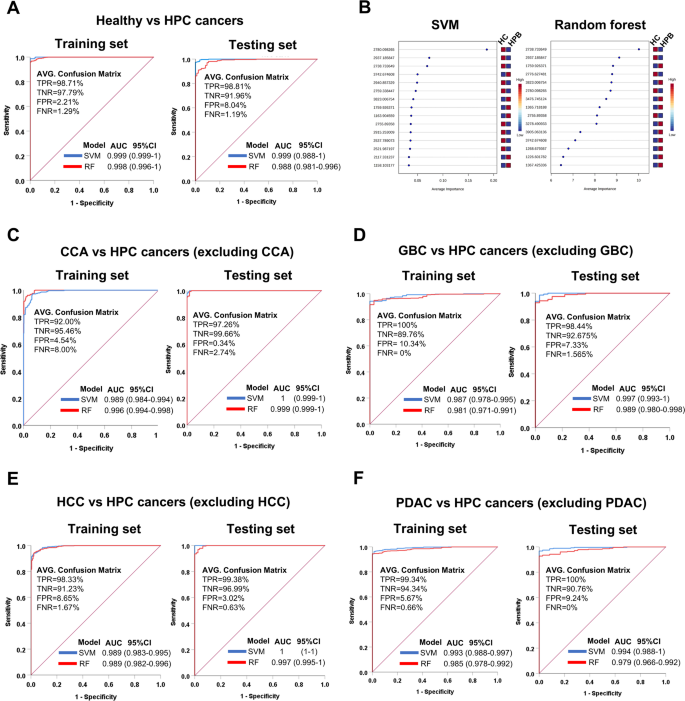
Receiver operating characteristic (ROC) curves and feature importance of SVM and Random Forest models based on 71 candidate peptides. (A) ROC curves comparing healthy controls and HPB cancers. (B) Top 15 peptides ranked by mean importance scores from SVM and random forest. (C–F) ROC curves for pairwise comparisons between CCA vs. HPB cancers (C), GBC vs. HPB cancers (D), HCC vs. HPB cancers (E), and PDAC vs. HPB cancers (F). Blue curves represent SVM models, while red curves indicate Random Forest models. ROC performance is reported as AUC with 95% confidence intervals. Classification metrics include TPR (true positive rate), TNR (true negative rate), FPR (false positive rate), and FNR (false negative rate), calculated from the corresponding confusion matrices.
The RF model also performed well. In the training set, the healthy vs. all cancer classification achieved 97.10% accuracy, with precision of 99.85%, recall of 96.75%, MCC of 0.90, and ROC of 0.998. Comparisons with individual cancer types showed perfect classification in all cases (MCC = 0.99–.00) (Table 3; Fig. 3A). In the testing set, RF showed slightly lower performance for the healthy vs. all cancer group (accuracy: 89.14%, MCC: 0.76, ROC: 0.988). MCC values for individual comparisons were as follows: CCA (1.00), GBC (0.81), HCC (1.00), and PDAC (0.87) (Table 3; Fig. 3A).
In addition, to identify the most important peptides for classification, a mean importance measure was calculated for both SVM and RF models. In RF, peptide importance was derived from the mean decrease in accuracy across trees, while SVM used recursive feature elimination (RFE) with cross-validation to rank features by their contribution. The mean importance score reflects the average impact of each peptide across all model iterations. The Top 15 peptides with the highest importance scores were consistent across both models, indicating that despite their different learning methods, both algorithms identified a similar set of key features (Fig. 3B). This consistency points to a robust, model-independent signature, reinforcing the biological and statistical relevance of these peptides in distinguishing the sample groups. The convergence of important features across models strengthens confidence in their predictive value, suggesting that the observed classification performance is driven by strong, reproducible signals rather than being model-specific.
In addition, we utilized 71 PMFs to develop SVM and RF models for the classification of HPB cancers using a one-vs-all (OvA) classification strategy. This approach was adopted to support potential clinical application, as HPB cancers often present with overlapping anatomical locations, making differential diagnosis challenging. Therefore, the OvA strategy was applied to enhance the discriminative power of the models in this context.
In the training set, the SVM model demonstrated excellent performance in differentiating each HPB cancer subtype from the remaining cancer types. The classification accuracy was highest for HCC vs. other HPB cancers (accuracy = 94.97%, MCC = 0.88, ROC = 0.989), followed by PDAC (accuracy = 94.67%, MCC = 0.80, ROC = 0.993), CCA (accuracy = 92.60%, MCC = 0.85, ROC = 0.989), and GBC (accuracy = 90.98%, MCC = 0.56, ROC = 0.987) (Table 4; Fig. 3C–F). Notably, precision and specificity (TNR) values reached 100% for GBC and PDAC, although recall was relatively lower, particularly for GBC (34.41%).
In the testing set, the SVM model maintained high discriminatory power across all cancer subtypes. The model achieved near-perfect classification performance for HCC (accuracy = 99.68%, MCC = 0.99, ROC = 1) and CCA (accuracy = 99.04%, MCC = 0.98, ROC = 1), followed by strong performance for GBC (accuracy = 94.87%, MCC = 0.78, ROC = 0.997) and PDAC (accuracy = 91.67%, MCC = 0.73, ROC = 0.994) (Table 4; Fig. 3C-F). These results underscore the model’s robustness and reliability, particularly in distinguishing CCA and HCC from other HPB cancer types.
Similarly, the RF model also yielded strong classification performance across most comparisons. In the training set, classification accuracy was highest for PDAC (95.12%, MCC = 0.81, ROC = 0.985), followed by CCA (94.38%, MCC = 0.89, ROC = 0.996), HCC (91.42%, MCC = 0.82, ROC = 0.989), and GBC (90.24%, MCC = 0.54, ROC = 0.981) (Table 5; Fig. 3C-F). Precision remained high across all subtypes (≥ 98.68%), but recall was markedly lower for GBC (32.65%), similar to the SVM model.
In the testing set, the RF model performed particularly well for CCA (accuracy = 97.76%, MCC = 0.96, ROC = 0.999) and HCC (accuracy = 95.51%, MCC = 0.89, ROC = 0.997), and to a slightly lesser extent for GBC (accuracy = 91.67%, MCC = 0.71, ROC = 0.989) and PDAC (accuracy = 91.99%, MCC = 0.73, ROC = 0.979) (Table 5; Fig. 3C–F). These findings confirm the consistent diagnostic potential of PMFs in differentiating between HPB cancer subtypes, with both SVM and RF models demonstrating high reliability, though SVM generally showed slightly superior performance, particularly in handling class imbalance and recall rates for certain subtypes.
To reduce model complexity, the top five discriminative peptides from each group were selected and used to construct classification models using SVM and RF algorithms under an OvA framework. As shown in Supplementary Fig. S4–S6, reducing the number of peptides to five per class resulted in a noticeable decrease in classification performance in both the training and testing sets when compared to the full model using 71 peptides. This decline was particularly reflected by the lower MCC values observed across all comparisons, as detailed in Supplementary Table S1.
In addition, to evaluate the relative diagnostic performance of peptide-based models, we constructed additional SVM and random forest models using clinical biomarkers included in the STARD checklist (ALT, AST, ALP, total bilirubin, CEA, and CA19-9). Specifically, we developed models based on (1) clinical biomarkers alone, which served as the baseline, and (2) a combination of clinical biomarkers and the 71 peptide mass features (PMFs). The baseline models trained using only the clinical biomarkers (Supplementary Table S2–3) demonstrated inferior performance compared to the models using the 71 PMFs alone, as evidenced by lower overall metrics. This finding indicates that peptide-based features possess superior discriminatory power in our dataset. Notably, adding clinical biomarkers to the 71 PMFs (Supplementary Table S4–5) did not significantly improve model performance, suggesting that the selected peptides alone are sufficient and may already capture the diagnostic information provided by conventional biomarkers.
These results suggest that PMFs provide high discriminatory power in differentiating healthy individuals from HPB cancer patients. The high MCC values across most comparisons, particularly in the SVM model, confirm the reliability and robustness of these models, even in the presence of class imbalance. Both SVM and RF classification models demonstrated strong diagnostic potential for identifying HPB cancers, with SVM performing slightly better in most scenarios, including the ability to handle class imbalance more effectively. These findings emphasize the value of PMFs as promising biomarkers for cancer diagnosis, offering both high sensitivity and specificity for clinical applications in HPB cancer screening and detection. In addition to distinguishing between healthy individuals and patients with HPB cancers, PMFs also exhibited strong classification performance in differentiating among individual HPB cancer subtypes, further supporting their utility for both general diagnosis and precise cancer subtype classification.




