Herein, the development of two separate but related models to evaluate static images of denuded MII oocytes is described. The first model performs multiclass segmentation of the oocyte into ooplasm, ZP, and PVS, and the resulting masks are utilized as inputs for the second model—a classifier model (referred to as mask model) that extracts features from the masks and generates predictions of whether an oocyte develops into a blastocyst (Fig. 6). We summarize how the performance of the mask model is assessed and methods of determining the significance of the features it uses to make predictions. Finally, the mask model is compared to an ensemble model incorporating an additional DL model.
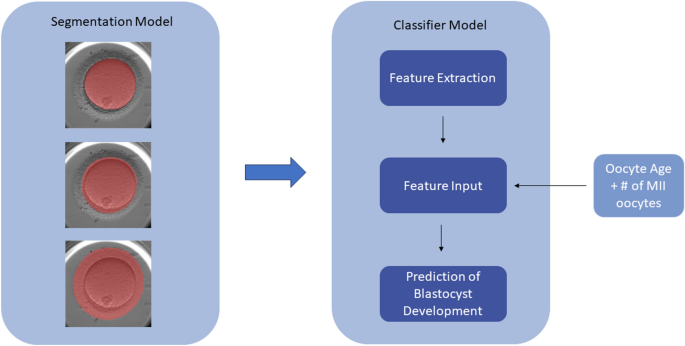
The proposed workflow utilizes two models—the first creates masks for each image of the oocyte, which is then used along with clinical variables as inputs into a classifier model (mask model) to generate a prediction of blastocyst development.
Ethical approval
Research methods were all performed in accordance with relevant guidelines and regulations, and in accordance with the Declaration of Helsinki. Ethics and IRB approvals were obtained for the clinics involved from Veritas (2022-2547-13359-1), FutureLife Scientific Advisory Board, and Indira IVF Hospital Institutional Ethics Committee (IIHPL-UDR/RCT/004_2022), as necessary. Under the guidelines of the Human Fertilization and Embryology Authority (HFEA), studies on oocytes in the UK do not require a research license. Each participating centre reviewed the study protocol and provided approval. All subjects contributing their oocyte images to the study provided written informed consent to data sharing for research purposes prior to undergoing ICSI. Datasets were anonymized to protect patient privacy before being shared.
Development of segmentation model
Images of 7792 denuded MII oocytes were utilized for the development of our segmentation model. 3258 images were obtained from five globally dispersed fertility clinics (located in Canada, UK, Spain, Czechia, and India). An additional 3138 images were obtained from an open-source dataset licensed under a Creative Commons Attribution 4.0 International36. The open-source data contained oocytes from various species—sea urchin, mouse and humans, however, only images of human oocytes, consisting of a combination of metaphase I (MI) and metaphase II (MII) maturation stages, were utilized. The resulting dataset included microscope and time-lapse (EmbryoScope and Geri) images of MI and MII human oocytes immediately pre- and post- ICSI. The task of manually assigning ground truth labels (ooplasm, PVS, and ZP) for 4654 images using the Computer Vision Annotation Tool (CVAT) was divided between three senior embryologists. This step was not necessary for the open-source dataset, which already possessed annotations for the ooplasm. Annotations of ZP and PVS for the open-source images were automatically generated by Oocytor, a Fiji Plugin tool capable of segmenting oocyte contours from multiple species, and then manually reviewed36. Of the 3138 open-source images, 380 samples with incorrect labels were removed during error analysis, leaving 2758 to be used. A final combined dataset of 7412 images of denuded oocytes underwent a split of approximately 60:20:20 to generate the model training, validation, and test sets—consisting of 4453, 1476, and 1483 images respectively.
A model with Fully Convolutional Branch TransFormer (FCBFormer) architecture was developed with PyTorch to perform multi-class segmentation of the oocyte37,38. We initially built a model of U-Net architecture, first to segment each region of the oocyte individually (background vs. region of interest), then in a second iteration to segment all four classes (background, ooplasm, PVS, ZP). The shift from binary segmentation to multi-class segmentation was justified by poorer performance on the PVS, a finding consistent with another study24. U-Net models are commonly applied to the task of medical image segmentation, due to their ability to extract both high-level contextual information (e.g. size, shape) and finer details that allow them to generate accurate segmentations39. The incorporation of skip layers in the U-Net architecture allows for the propagation and therefore the preservation of information from earlier layers to later layers in the network, which facilitates effective learning even with smaller datasets. However, the convolutional operator in CNNs such as U-Nets acts locally, making it challenging for these models to capture long-range dependencies. In contrast, transformer models efficiently make use of long-range dependencies to learn global context40. Thus, the FCBFormer architecture was recently proposed to exploit the merits of both fully convolutional neural networks and transformer models.
The FCBFormer model described here consists of a fully convolutional branch and transformer branch. The architecture of the transformer branch is Pyramid Vision Transformer v2—pre-trained on ImageNet—which is effective at dense prediction tasks (predicting at pixel-level) through generating multiscale feature maps and has the additional advantage over other transformer models in its ability to represent local features41,42,43. Both the input image and the segmented image generated by the model are resized to 352 by 352, the dimensions the FCBFormer architecture was developed on38.
Several augmentations were implemented prior to training using Torchvision44. Colour jitter with a brightness factor and a contrast factor uniformly sampled from 0.8 to 1.2 with a 0.7 probability was applied to vary the brightness and contrast of samples, respectively. Random JPEG compression with quality uniformly sampled from 35 to 100%, with a 0.5 probability was also applied, to vary the amount of detail present in each sample. Image sharpness was randomly adjusted by a factor of 2 with a probability of 0.5. Images also underwent random posterization with a 0.5 probability by reducing two bits for each color channel. To improve image contrast, the histogram equalization technique was applied randomly with a probability of 0.5. The order of the augmentation steps was random, and images were normalized to an interval of − 1 to 1.
The model was trained on 4453 images for 40 epochs using a batch size of 24 and Lion optimizer with an initial learning rate of 0.000145. The Lion optimizer is memory-efficient and outperforms other optimizers such as Adam. Focal Loss was selected as the loss function, as it addresses class imbalance in the training set and facilitates the model’s ability to efficiently learn challenging or under-represented classes–relevant in the case of the oocyte, as the PVS is smaller than the other regions46.
To assess the accuracy of the FCBFormer model’s segmentation on the test set, intersection over union (IoU) scores—which determine the percentage of overlap between the ground truth and the model’s predictions—were calculated for each image and averaged to give a mean score for the ooplasm, ZP, and PVS. IoU scores were calculated separately for each mask to permit the monitoring of performance on each region of the oocyte, using the following formula:
$$IoU\left( {Y_{label} ,Y_{pred} } \right) = \frac{TP}{{TP + FP + FN}},$$
(1)
where \({Y}_{label}\) is the ground truth, which was manually assigned or checked by embryologists, and \({Y}_{pred}\) is the predicted masks. \(TP\) is the true positive, which represents the intersection between the ground truth and model prediction; \(FP\) is the false positive which represents the area where the ground truth is negative, but the prediction is positive; and \(FN\) is the false negatives, representing the area where the ground truth is positive, but the prediction is negative.
Development of mask model
51,831 static images of denuded MII oocytes with known blastocyst development outcomes, collected from seven clinics across six geographical locations (Canada, USA, Spain, Czechia, India, and UK), were used to develop a binary classification model (herein, referred to as the mask model) utilizing extracted features in addition to patient characteristics (i.e. oocyte age and number of MII oocytes) as inputs, with the output being a prediction of blastocyst positive or blastocyst negative. 2058 images of this dataset were utilized to develop the segmentation model described above. Images were captured either immediately pre- or post-ICSI, from various microscope setups (e.g. Leica, Nikon, Olympus, and Zeiss microscopes; with a designated Basler acA3088-57 camera) and time-lapse (e.g. Embryoscope, Geri) systems. All images were two-dimensional, single-plane, taken within a magnification range of ×200– ×400. Greater details on the imaging systems were unavailable due to the retrospective nature of the data collection. 6793 patients of mean age 37.1 ± 4.3 undergoing 8089 ICSI cycles contributed to the dataset. Conventional IVF cycles were not included in the model, as oocytes are not denuded during conventional IVF prior to insemination. Oocyte age (mean 36.0 ± 4.5 years) was used as a clinical feature in place of patient age to account for donor oocytes. Oocyte cohorts consisted of a mean 6.9 ± 4.6 oocytes. The dataset underwent a split of approximately 60:20:20, with 29,262, 10,812, and 11,757 images allocated between the training, validation, and test sets respectively. The dataset underwent splitting at the patient level rather than at the oocyte level to prevent data leakage. 20,927 oocytes had a ground-truth outcome of blastocyst positive (40.4% of the dataset), while 30,904 were labelled as blastocyst negative (59.6%). A blastocyst was defined as a day 5–7 (post-fertilization) embryo with a minimum Gardner grade of 1CC. Low-quality blastocysts were not filtered out. Dataset details are found in Supplementary Table S6.
The oocyte features to investigate were selected in accordance with morphometric descriptors commonly measured in two-dimensional cell shape analysis12. An ellipse was fitted to each mask to calculate the major (\({L}_{maj})\) and minor (\({L}_{min})\) axis, while perimeter \(\left({L}_{p}\right)\), area \(\left(A\right)\), and area of the convex hull, \({A}_{hull}\) , were calculated from the masks themselves–these measurements were obtained using inbuilt OpenCV functions. These features were used in downstream calculations to define the aspect ratio, roundness, circularity, and solidity of each oocyte region, as summarized in Table 1. Ratios between two different masks (e.g. ooplasm vs ZP) for major axis, minor axis, perimeter, and area–here, termed relative features—were calculated and averaged for the entire cohort of oocytes. For each oocyte, cohort relative features were calculated by subtracting the cohort average from the value of the relative feature—thus, negative values for a given cohort relative feature indicate that the value of the relative feature for the oocyte under consideration is lower than the average of the cohort. Area of the convex hull was only used to calculate solidity and was not compared between oocyte regions. Altogether, 47 features were extracted to use as inputs for the mask model, encompassing relative features, average cohort features, cohort relative features, and mask-specific features. Relative features (e.g. ooplasm vs ZP area ratio) were used as inputs over absolute features (e.g. ooplasm area), due to their greater generalizability, which allowed the model to achieve a more balanced performance between clinics. Extracted features were also statistically compared between the group of blastocyst-positive and blastocyst-negative oocytes using the Welch’s t-test.
We experimented with two approaches to addressing the task of oocyte classification. The first approach was to use the Python-based automated machine learning toolkit, Auto-sklearn, which leverages Bayesian optimization, removes the need for the user to select algorithms and perform hyperparameter tuning, and constructs ensembles47,48. The Auto-sklearn model was constructed from an ensemble of extra trees classifiers—a type of machine learning model consisting of decision trees, where each decision tree randomly learns from the training set—and Gradient Boosting classifiers—where a strong model is created by combining several weak models, with each weak model correcting the mistakes of previous models. The second approach was training and evaluating a model with the light gradient boosting machine (LightGBM) framework. The LightGBM framework applies the technique of gradient boosting to quickly and efficiently train a collection of decision trees and is designed to be capable of handling large datasets and using less memory. To evaluate these two approaches, the performance metrics of area under the curve (AUC), sensitivity, and specificity were calculated.
To address the high dimensionality and potential interdependencies of the 47 features included in the mask model, a PCA was conducted prior to model training for comparison to a model trained without PCA. PCA attempts to reduce data dimensionality while maintaining the most relevant information, which can potentially improve the resulting model performance. Model performances were compared by AUC with a paired DeLong test49.
Feature importance with respect to blastocyst formation was assessed using the SHAP method. MASV across all samples in the dataset were used as a measure of the input features’ importance relative to one another, where a higher value is indicative of greater importance to the model’s prediction. SHAP values also facilitate the understanding of how each feature impacts a specific prediction. We additionally assessed the impact of removing the following features on model AUC: features related to each mask, clinical features, and cohort features. Changes in AUC were assessed for significance with the paired DeLong test49.
Ensemble model
To assess if the problem of oocyte evaluation benefits from incorporating DL or if a single ML model is sufficient, an ensemble model consisting of the described LightGBM model and a DL model of ConvFormer architecture was constructed50. The ConvFormer architecture combines the strength of convolutional neural networks in capturing local details and the strength of transformer models in learning long-range dependencies to effectively classify images. It is pre-trained on ImageNet41.
The ConvFormer model was trained and validated on 48,363 and 10,812 denuded MII oocyte images, respectively, with associated outcomes from seven clinics. This dataset comprises of the same images utilized for the mask model development described above. Images underwent data augmentation using the AugMix method, which eliminate the need for hyperparameter tuning51. Pixels outside the ooplasm, PVS and ZP were set to zero to avoid the model paying attention to irrelevant parts of the image, such as residual cumulus cells or background noise. Images were cropped by an algorithm trained with Faster-RCNN and cropped images were resized to 224 by 22452. Binary cross entropy loss and an SGD optimizer with momentum of 0.9 and weight decay of 7.9e-5 were used to train the model, and the learning rate was adjusted using a cosine annealing schedule with warm restarts53. The model was trained for 40 epochs with a batch size of 128. To balance the positive and negative classes of the training dataset, samples of the minor class were oversampled to the same size of the major class for each clinic.
AUC, sensitivity, and specificity were all calculated for the ensemble model. Additionally, AUC of the DL model with and without the LightGBM was compared for statistically significant differences with the paired DeLong test.
Subgroup analysis
Subgroup analyses by clinic and age group were performed to assess clinical relevance of the mask model and ensemble model for different patient populations (i.e. by geographic location and age). The age groups chosen for analysis were < 30 (n = 1607), 30–34 (n = 3515), 35–37 (n = 2646), 38–39 (n = 1594), and ≥ 40 (n = 2395) years, based on the age-related statistical decline of live birth in ICSI cycles31,54.
External validation
A dataset of 9789 MII oocytes with known laboratory outcomes was obtained from a single Spanish clinic, independent of model development. 140 oocytes were removed from the analysis due to unknown blastocyst development outcomes (i.e., were only cultured until the cleavage stage prior to embryo freezing or transfer; in the case of transferred embryos, had unknown or negative implantation outcomes, as a positive implantation outcome would indicate in vivo blastocyst development). The impact of oocyte age on quality is especially pronounced after age 40, however this represents a significant percentage (17%) of the dataset. Therefore, we excluded oocytes > 43 years old, as the chances of live birth have been observed to fall below 5% in this age group54. The final dataset consisted of 9346 oocytes (mean age 31.0 ± 7.8 years), corresponding to 909 patients. 4691 of the oocytes developed into blastocysts (50.2%), while 4655 (49.8%) failed to develop into blastocysts. Performance of the mask model and ensemble model were evaluated on this dataset for AUC, sensitivity, and specificity, and AUC values were compared with the paired DeLong test.




