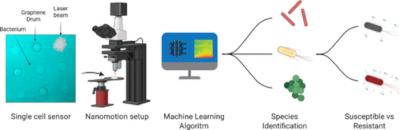
Researchers from Delft University of Technology, its spin-off company SoundCell, and Reinier Haga MDC have shown that graphene “nanodrums” combined with machine learning can identify bacteria and determine their antibiotic susceptibility within hours of nanomotion of single cells. This approach combines bacterial identification and antimicrobial susceptibility testing (AST) into one label-free measurement at the single-cell level.

Each nanodrum consists of a bilayer graphene membrane less than 1 nanometer thick, suspended above an 8-micrometer-wide cavity that can house a single bacterium. When living cells attach to the drum, their inherent motion causes nanoscale vibrations in the graphene, which are read optically as a time-dependent signal. This configuration avoids ensemble averaging and captures the mechanical behavior of individual bacteria.
Instead of analyzing noisy time-domain traces, the nanomotion signal is converted into a time-frequency spectrogram that preserves both spectral and temporal information. These spectrograms are used as input to machine learning models (Convolutional Neural Networks (CNNs) and Support Vector Machines (SVMs)) to learn discriminative features directly without manual feature engineering. A total of 456 measurements were used for species identification and 347 susceptibility tests, and performance was evaluated using accuracy, sensitivity, specificity, and ROC-based metrics.
Using this framework, the researchers were able to distinguish between E. coli, Staphylococcus aureus, and Klebsiella pneumoniae, as well as distinguish between resistant and susceptible strains. Species identification reached an accuracy of up to 88%, and susceptibility testing of meropenem-resistant and susceptible E. coli achieved an accuracy of up to 98.6%. This corresponds to an accuracy of 98% for resistant/susceptible classification. CNN showed the best performance for susceptibility profiling, whereas SVM provided more stable generalization across species for identification.
Because the model operates with single-cell nanomotion, this method achieves both ID and AST in 1–2 h without additional culturing after loading the graphene drum with bacteria. With further developments such as parallel cartridge-based sensor arrays and training datasets enriched with resistant bacteria, this graphene-ML platform could evolve into a rapid label-free diagnostic and powerful clinical tool for investigating antimicrobial resistance and cell biophysics at the single-cell level.