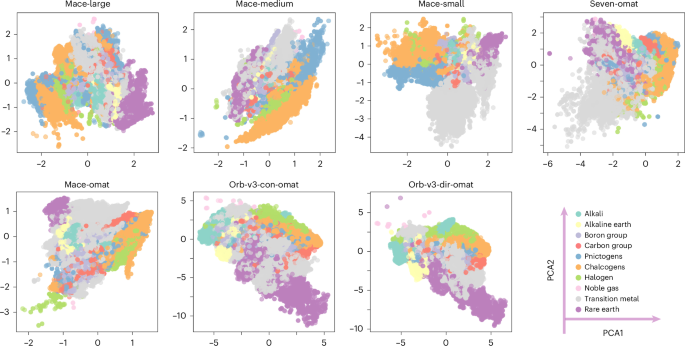
Aligning incompatible representations
We chose seven foundation MLIPs representing distinct architectures, datasets and approaches to equivariance and energy conservation. These include three MACE-MP-0 variants (large, medium and small)13 trained on the Materials Project Trajectory Dataset (MPtrj)14; two OMat24-based models15 (MACE-omat and Seven-omat); and two Orb-v3 models (Orb-v3-con-omat and Orb-v3-dir-omat), which differ in their treatment of force conservation. For each model, we extracted 282,847 atomic embeddings across 27,136 structures from the MP-20 dataset16.
We applied principal component analysis (PCA) to project these embeddings into a two-dimensional space, where the first two principal components (PCA1 and PCA2) capture the directions of greatest variance. While we also examined nonlinear visualization techniques such as uniform manifold approximation and projection (UMAP)17 (Supplementary Fig. 1), they distort global geometry and reconfigure relative distances. As shown in Fig. 1, the raw embeddings cannot be compared directly. Dimensionality varies by architecture; for instance, MACE embeddings use 128 dimensions, whereas Orb-v3 uses 256. More importantly, the representations depend strongly on the training set-up, including the specific architecture, loss function and random initialization. Even the MACE-MP-0 variants, which share the same training dataset (MPTrj) and objective, exhibit divergent latent spaces. This indicates that original representations act in arbitrary coordinates learned by each model architecture.

Two-dimensional PCA projections of atomic embeddings from seven foundation MLIPs reveal distinct variance directions and element clustering patterns. Although all models are trained to predict the same physical quantities for overlapping material sets, they learn embeddings in incompatible coordinate systems.
To enable meaningful comparison, a unified representation is needed. We identify four desiderata for its construction: (1) model agnosticism—the construction operates on any model’s latent space without access to architecture internals; (2) geometric faithfulness—relative distances and neighbourhood relationships among atomic environments are maximally preserved after projection; (3) sufficient diversity—the representation spans the chemical space broadly enough to resolve chemically distinct environments; and (4) robustness—the representation is robust to the choice of random seed and invariant to anchor permutation, ensuring that observed structure reflects intrinsic geometry rather than a particular realization. Therefore, each step of the following construction is designed to meet these criteria.
We first project the model-specific embeddings into a unified latent space. We achieve this using the relative representation strategy proposed by ref. 18, establishing a common coordinate frame independent of the original architecture—desideratum 1, model agnosticism. We use cosine similarity instead of distance functions (Euclidean L2 or Manhattan L1) as it provides scale invariance—critical when comparing embeddings with varying norms (for example, 3–5× differences across models; Table 1). This choice naturally bounds projections to [−1, 1], facilitating interpretation. The transformation framework is illustrated in Fig. 2a. Let \({{\bf{e}}}_{i}\in {{\mathbb{R}}}^{d}\) denote the original d-dimensional embedding of an atomic environment i. We select a set of K anchor vectors, {a1, a2, …, aK}, from the embedding manifold. The platonic transformation, T(ei), projects ei into the anchor-defined space based on its cosine similarity to these reference points:
$${{\bf{z}}}_{i}=T({{\bf{e}}}_{i})={\left[\cos ({{\bf{e}}}_{i},{{\bf{a}}}_{1}),\cos ({{\bf{e}}}_{i},{{\bf{a}}}_{2}),\ldots ,\cos ({{\bf{e}}}_{i},{{\bf{a}}}_{K})\right]}^{\top },$$
(1)
where the cosine similarity is defined as:
$$\cos ({{\bf{e}}}_{i},{{\bf{a}}}_{k})=\frac{{{\bf{e}}}_{i}^{\top }{{\bf{a}}}_{k}}{\parallel {{\bf{e}}}_{i}{\parallel }_{2}\,\parallel {{\bf{a}}}_{k}{\parallel }_{2}}.$$
(2)
The resulting vector \({{\bf{z}}}_{i}\in {{\mathbb{R}}}^{K}\) constitutes the embedding in the unified platonic space. By computing the cosine similarity between an input embedding and each anchor, we transform the absolute, model-dependent coordinates into a relative representation. The dimensionality of this new space is determined solely by the number of anchors, K, with each axis corresponding to the similarity to a specific anchor, rather than an arbitrary feature channel.

a, Schematic of the anchor-based transformation. Green, red and purple points represent a set of three anchor vectors (ak), and the yellow point indicates a sample vector \({{\bf{e}}}_{i}\in {{\mathbb{R}}}^{d}\). The transformation projects ei into the anchor-defined space via cosine similarity. b, Distribution of 100 DIRECT-sampled anchors (blue dots) overlaid on the PCA projection of the original embedding manifold (orange background).
To ensure that the anchor set captures the diversity of the embedding manifold, we compared random sampling with Dimensionality-Reduced Encoded Clusters with Sratified (DIRECT) sampling19. Originally developed to facilitate MLIP training, DIRECT sampling selects points that maximize coverage in the chemical latent space. We evaluated these strategies across multiple random seeds ∈ {0, 42, 12,345}. As detailed in Supplementary Table 1, DIRECT sampling yields substantially more diverse anchor sets, characterized by larger pairwise distances (>1.5) and lower Silhouette scores (<0.1) compared with random selection. Consequently, we use the DIRECT strategy throughout this work to ensure broad coverage and high representational diversity (Fig. 2b).
Convergence to a shared chemical geometry
The transformed representations obtained using anchor sets of varying sizes, K ∈ {3, 8, 20, 50, 100, 200, 400}, are presented in Fig. 3a. Even a minimal set of three anchors effectively aligns embeddings across disparate architectures, highlighting transition metals (grey) forming a distinct cluster, clearly separable from chalcogens (orange), halogens (green) and other main-group elements. As the anchor count increases, the unified representation stabilizes, with variance converging once the anchor set size K reaches 100. Here we show relative representations from Mace-medium, Mace-small and Orb-v3-con-omat, representing both equivariant and non-equivariant architectures. A more comprehensive trend of convergence across all models is provided in Extended Data Figs. 1 and 2.

a, Transformed representations as a function of anchor set size (K = 3 to K = 400). b, Two-dimensional PCA projections of converged representations using 100 randomly sampled anchors. c, Projections using 100 DIRECT-sampled anchors. Despite architectural diversity, all models transformed with DIRECT sampling show substantial alignment. Non-equivariant models (Orb-v3) exhibit systematic skewness. The dummy model (untrained, random weights) shows no chemical structure, confirming that alignment reflects learned physical knowledge. The colour map follows the labelling in Fig. 1.
These results indicate that anchors act as stable reference points defining a shared coordinate system, satisfying requirements 2 and 4—geometric faithfulness and robustness. Once the principal geometric relationships are pinned to these anchors, the remaining embeddings naturally align, reflecting the same underlying physical drivers regardless of the model origin. Furthermore, the evolution of the embedding topology with increasing K reveals a hierarchical organization within the chemical space. While a small number of anchors captures the coarse global structure, increasing the anchor density resolves finer structural details.
The quality of the unified representation also depends on anchor diversity—desideratum 3. As illustrated in Fig. 3b,c, representations constructed using DIRECT sampling yield a more unified manifold than those using random sampling. DIRECT-sampled anchors produce tighter clustering with mean pairwise distances ranging from 1.58 to 2.67, whereas random sampling leads to skewed, loosely distributed spaces (distances of 1.80–3.11; Supplementary Table 2). This effect is particularly pronounced for models trained on the OMat24 dataset (Orb-v3 variants). Their platonic embeddings show skewed distributions relative to the MACE-MP-0 models trained on MPTrj, particularly when anchors are chosen at random. This behaviour persists across anchor set sizes from K = 20 to K = 400 (Extended Data Figs. 1 and 2).
The role of anchor diversity parallels findings in image and language representation alignment20,21. Small, non-diverse anchor sets (≤20) provide only a coarse global alignment; as the anchor set grows (100–400), diversity becomes essential to constrain the mapping and recover a globally consistent alignment, where the fidelity of the unified space depends on the structural richness of the probes. To validate that this geometry arises from learned physics rather than mathematical artefacts, we applied the transformation to a ‘dummy’ MACE model initialized with random weights (Methods). As shown in the rightmost columns of Fig. 3b,c, this untrained model yields no discernible chemical structure. This contrast confirms that the platonic representation emerges from meaningful correlations learned from the data. We further demonstrate the universality of this framework by extending it to additional architectures, including NequIP-OAM-L22,23 and Mace-mpa-0, in Supplementary Fig. 3. All embeddings are extracted from the invariant component before energy readout, which captures the richest structural encoding after full message passing while avoiding distortion by the task-specific decoder (details in Supplementary Section 10).
Quantifying representational interoperability
The visual convergence observed above establishes qualitative alignment; to quantitatively assess this convergence and the interoperability of embeddings in the unified space, we compute three complementary metrics at different geometric scales. Procrustes analysis (ScoreProcrustes) measures global alignment: it finds the optimal rotation between two embedding distributions and reports their residual distance, quantifying whether two models organize chemical space in the same overall geometry24. Mutual k-nearest neighbours (ScoremKNN) measures local consistency: it computes the fraction of shared nearest neighbours between two models for the same atomic environments, quantifying whether two models agree on which environments are most similar. Normalized optimal transport (ScoreOT) measures distributional distance: it computes the minimal effort required to morph one latent distribution into another, capturing both global and local differences25.
As shown in Fig. 4b, foundation MLIPs show substantial global topological similarity, particularly among architectures sharing design principles (for example, MACE variants show ScoreProcrustes > 0.86). Similar universal convergence is independently confirmed by ref. 26. However, this macroscale alignment masks substantial local divergence. The local neighbourhood similarity (ScoremKNN) remains low (<0.38) across all pairs (Fig. 4a), notably dropping from the values observed in the original, unaligned spaces (Supplementary Fig. 4). This disparity indicates that while different models converge on the same global physical manifold, their local neighbourhood structure remains distinct. A recent work also revealed similar divergence27. Notably, non-equivariant models (Orb-v3) show near-zero ScoremKNN against equivariant models, confirming that the lack of symmetry constraints leads to a fundamentally different encoding of local atomic environments.

a, mKNN scores (local fidelity). b, Procrustes scores (global alignment). c, Normalized OT cost. d, The composite SuperScore. Colour bars indicate the magnitude of each metric. Higher mKNN and Procrustes scores and lower normalized OT cost indicate better alignment and contribute to a higher composite SuperScore. e,f, Element-level embeddings projected into the unified space (K = 100 anchors) reveal consistent periodic clustering across all seven models (Extended Data Fig. 3). g,h, Comparison of platonic representations from a generative diffusion model (Chemeleon, t = 1, the best step at reconstruction) trained without energy or force supervision (g) and a foundation MLIP (MACE-small) (h). Generative model without physical constraints shows less structured latent representations than foundation MLIP models trained with energy and force supervision. i, Density functional theory potential energy surface of Cmcm SnSe along a double-well vibrational mode with two soft modes (λ1, λ2) at imaginary frequencies; structures on either side of the well are related by symmetry. U, the double well potential energy; Q, normal model coordinates. j–l, Structure-level embedding traces along this vibrational path for Orb-direct (j), Orb-conservative (k) and the generative model (l). Non-equivariant Orb models produce smooth but geometrically distinct branches for the two symmetry-equivalent wells. The generative model produces a disordered trace with no smooth interpolation. The colour map in e–h follows Fig. 1.
The optimal transport (OT) cost map (Fig. 4c) partitions the seven models into three coherent clusters, revealing the interplay between architecture and data. High OT distances separate the OMat-trained equivariant models (Seven-omat, Mace-omat) from the MACE-MP-0 family, isolating the influence of dataset composition. The largest distances, however, separate the non-equivariant Orb-v3 models from all others, confirming that disparate physical constraints create distinct energy landscapes. We can define a combined metric:
$$\mathrm{SuperScore}=\frac{1}{2}({\mathrm{Score}}_{\mathrm{Procrustes}}+{\mathrm{Score}}_{\mathrm{mKNN}})\times {{\rm{e}}}^{-{\mathrm{Score}}_{\mathrm{OT}}}$$
that averages the global and local alignment scores, while applying a penalty based on the transport cost. Bounded between [0, 1], SuperScore offers a summary of representational compatibility.
As an illustration, the unified representation explicitly preserves chemical structure (Fig. 4e,f; all models in Extended Data Fig. 3). When projecting element-level embeddings that are mean-pooled over all atomic environments, all models produce a similar topology of the periodic table. Elements from the same group, such as halogens, chalcogens and pnictogens, cluster coherently, and periodic trends (for example, F–Cl–Br–I) form smooth trajectories. However, architectural biases persist: equivariant models (MACE, SevenNet) produce compact, spherical clusters, whereas non-equivariant models (Orb-v3) yield skewed distributions. This suggests that without explicit symmetry constraints, the model relies more heavily on statistical correlation than physical symmetry to organize chemical space.
Disentangling physics from data
Our results suggest that the platonic representation captures physically meaningful structure beyond what is captured by the training distribution alone (Fig. 4g–l). To isolate the role of physical supervision, we projected embeddings from Chemeleon28—a generative diffusion model trained on the same distribution of crystal structures and chemical compositions as the foundation MLIPs but without energy or force targets—into the unified platonic space. Despite identical data coverage, the generative model’s representation lacks the tight periodic chemical organization exhibited by all foundation MLIPs (Fig. 4g,h): chemically related element groups that form distinct, well-separated clusters in the MLIP representations are heavily mixed and unresolved. This contrast demonstrates that the periodic topology of the platonic representation requires physical supervision to emerge and cannot be attributed to training data distribution alone.
To further disentangle the contributions of optimization, architecture and physics, we traced structure-level embeddings along a double-well vibrational mode of Cmcm SnSe, whose phonon dispersion exhibits two soft modes (λ1, λ2) with imaginary frequencies29 (Fig. 4i). Structures sampled from opposite sides of the double well are related by symmetry and should map to the same point in a physically faithful representation. The non-equivariant Orb models produce smooth but geometrically distinct embedding trajectories for the two symmetry-equivalent wells (Fig. 4j,k), indicating that these models have learned the continuity of the potential energy surface but not its symmetry. In contrast, the generative model produces a disordered, non-smooth trace (Fig. 4l), reflecting a complete absence of potential energy surface topology in its representations. These observations define a diagnostic hierarchy: a model whose embedding trace is smooth but bifurcated has internalized potential energy surface continuity but not symmetry; a model whose trace is disordered has not learned the energy landscape at all. Both signals are accessible from the representation geometry alone.
However, a rigorous mathematical correspondence between manifold geometry and physical observables has not yet been established; we identify this as a key open problem at the intersection of machine learning and materials science.
Algebraic consistency and model stitching
A potential utility of a platonic representation is embedding arithmetic across different models. By mapping into a unified coordinate system, vector operations can be performed. We evaluate this through three case studies: a complex oxide (Na3MnCoNiO6), a symmetry-sensitive polymorphic pair of TiO2 and a solid-state reaction to synthesize BaTiO3. All embeddings are computed in the unified space (K = 100 anchors), with cross-model similarity (c-sim) measured relative to MACE-large and extended cases are presented in Supplementary Information.
We define the material-level embedding, zMater, as the centroid of its atomic constituents in the unified space:
$${{\bf{z}}}_{\mathrm{Mater}}=\frac{1}{{N}_{\mathrm{atoms}}}\mathop{\sum }\limits_{i=1}^{{N}_{\mathrm{atoms}}}{{\bf{z}}}_{i}.$$
(3)
As shown in Table 1, models exhibit agreement on zMater for Na3MnCoNiO6, with c-sim values consistently between 0.79 and 0.87 (excluding Orb-v3-con-omat). Notably, OMat24-trained equivariant models (Seven-omat, Mace-omat) produce vectors with substantially larger norms (l ≈ 4.5–5.4) compared with MACE-MP-0 models (l ≈ 1.1). This scaling factor (~3–5×) is consistent across materials (Supplementary Table 3), indicating that while architectures may vary in signal magnitude, they encode similar angular information relative to the transformed axes.
To assess sensitivity to structural degrees of freedom, we analysed two TiO2 polymorphs: one orthorhombic (Pbcn) and one tetragonal (P42/mnm) space group. While intra-model similarity (i-sim) between polymorphs is high (≥0.95), the cross-model agreement on the difference vector, zMorph = zMater1 − zMater2, is low (≤0.40). This suggests that while global material identity is preserved, current foundation models, and by extension their unified representations, smooth over the subtle local distortions that distinguish polymorphs, highlighting a resolution limit in current pooling strategies.
We define the reaction embedding as zReact = ∑zproducts − ∑zreactants. For the formation of BaTiO3 from its binaries, we observe consistency across models (about ≥0.7 c-sim, except the Orb-v3-con model). Embedding compatibility can support zero-shot model stitching. This allows us to algebraically substitute the product state representation of one model with that of another, treating them as compatible vectors within the shared geometry. We constructed a hybrid reaction embedding, zReact-stitch, by pairing reactant embeddings from MACE-large with product embeddings from other models. As detailed in Table 1, inter-model compatibility is high. MACE-MP-0 variants show strong agreement (>0.88). Surprisingly, Orb-v3-con-omat exhibits higher stitchability with MACE-large (0.82) than the other OMat24-trained models. This demonstrates that models trained on non-overlapping datasets (MPtrj versus OMat24) can be algebraically combined to yield geometrically reasonable embeddings, opening potential routes for modular reuse of pretrained foundation potentials.
Platonic representation as a diagnostic framework
Beyond enabling cross-model algebraic operations, the shared platonic geometry also defines a physically meaningful reference frame for detecting representational deviations. We demonstrate three diagnostic applications, progressing from individual model training to architectural fidelity to physical symmetries, and structural typicality of unseen configurations.
Tracking training dynamics
In the context of transfer learning (Fig. 5a,b), platonic representation enables the visualization of the trajectory of atomic embeddings, distinguishing catastrophic forgetting from stable adaptation. Naive fine-tuning on Cu–Cu dimers causes unseen Au–Au embeddings to collapse (teal cross, Fig. 5a), dragging Cu weights from the correct trajectory (purple star) towards the contaminated one (teal star). Multi-head fine-tuning preserves Au knowledge (teal cross, Fig. 5b) while maintaining consistent Cu trajectories (overlapping purple and teal stars).

a,b, Purple trajectories denote the fine-tune strategy (ft1) targeting only Cu during fine-tuning; teal trajectories represent the fine-tune strategy (ft2) targeting both Cu and Au. For both cases, only Cu–Cu dimer information is provided. a, Naive fine-tuning on additional Cu data causes embeddings for unseen Au atoms to collapse (teal cross (Au–Au) trajectories are artificial due to catastrophic forgetting). b, Multi-head fine-tuning keeps unseen Au embeddings stable near their pre-trained positions (the pair of teal cross (Au–Au) trajectories) while keeping the weights of fine-tuned Cu–Cu interactions (overlapping stared trajectories). c, The equivariant fraction of embeddings from different models based on two-NN analysis. d, Rotational sensitivity test on BaCeO3: equivariant models preserve embeddings under rotation; Orb-v3 models do not. e, OMAT non-equilibrium structures at 1,000 K projected into the platonic space. Left: atomic-level platonic embeddings. Right: material-level platonic embeddings (pink) compared with that of stable MP-20 structures (blue). f, Distribution of structures across manifold regions (dense interior, sparse interior, near exterior, far exterior) at atomic level (top) and material level (bottom).
Diagnosing architectural limitations
Beyond training stability, the proposed framework also highlights fundamental limitations in architectures that do not maintain strict equivariance in their learned representations. We adopt the two-nearest-neighbour (two-NN) approach 30 to quantify effective dimension reduction arising from symmetry equivalence. Considering a reference atom (O) as an example in Fig. 5d, the ratio d1/d2—where d1 and d2 denote the first and second nearest-neighbour distances in representation space—serves as a metric for how the model distinguishes (or fails to distinguish) symmetry-equivalent sites. For perfectly equivariant representations, symmetry-related atoms collapse onto the same point, leading to d1 = 0. By measuring the fraction of atoms with d1 = 0, we estimate each model’s capacity to preserve equivariant structure in its learned embeddings.
Statistics in Fig. 5c show that all equivariant MACE models maintain a consistent level of detected equivalent points across the four representative space groups studied (P63/mmc, Fm\(\bar{3}\)m, I4/mmm, Pm\(\bar{3}\)m), indicating robust preservation of equivariant representations. SevenNet, despite incorporating E(3)-equivariant neural network layers, does not detect d1 = 0 equivariance in its original embeddings, probably owing to architectural truncation at lmax; NequIP uses stricter numerical tolerance (1 × 10−8 to 1 × 10−10, float64) but still shows imperfect detection. Both recover equivariance close to MACE levels at a tolerance of 1 × 10−6 (Supplementary Table 6). In this regard, transformation into platonic space enables more robust detection of equivariant structure in models with numerical precision limitations, evidenced by the partial re-identification of equivalence clusters for SevenNet and NequIP models (~15–30%; Fig. 5c), suggesting that the platonic projection may reduce numerical noise.
In contrast, non-equivariant Orb models fail to identify any equivalent atoms in either their raw or platonic embeddings, deviating from physically expected symmetry relations regardless of the numerical accuracy applied, indicating that platonic transformation cannot restore equivariance that was fundamentally missing in the architecture’s learning phase. This failure directly manifests in downstream predictions: as illustrated in Fig. 5d, rotating the BaCeO3 structure causes the Orb-v3 embeddings to diverge rather than remain invariant, a clear instance of symmetry breaking. This violation propagates into the force field, producing qualitatively incorrect phonon dispersions (Supplementary Fig. 5).
Ground-truth-free measure of structural deviation
More importantly, the manifold hypothesis31 holds that physically realistic atomic configurations concentrate near a low-dimensional manifold in representation space. Based on this, the platonic projection provides a geometry in which distances from this manifold can be interpreted as structural deviation, that is, the unified platonic space enables a ground-truth-free measure of structural typicality based on manifold distance (details in Supplementary Section 11), which is computed by (1) project an unlabelled query structure into the platonic space using the pre-computed anchor set, and (2) compute its distance and density to the reference manifold.
By partitioning the representation space into density-distance-based regions (dense/sparse interior, near/far exterior) relative to the reference manifold of known stable materials, we quantify how typical a query structure is without requiring any property labels. Relative to the near-equilibrium MP-20 anchors and the training sets of all foundation MLIPs studied, we projected configuration sampled from the non-equilibrium OMat dataset (rattled, 300 K and 1,000 K) into the platonic space (Extended Data Fig. 4c,d). Their projections remain interpretable and physically organized for both atomic-level and structural-level embeddings: 0 K relaxed structures occupy predominantly the dense interior, 300 K structures shift towards the sparse interior, and 1,000 K (Fig. 5e,f) structures shift further towards the near- and far-exterior regions. This same label-free measure extends naturally to quantifying the typicality of structures from generative models, which correlates strongly with their ‘novelty’ metric. In this regard, structures from GNoME32, MatterGen (MP-20)33 and Chemeleon are collected and their embeddings predominantly occupy the dense interior (82.8%, 78.5% and 79.6% respectively; Extended Data Fig. 4a,b), consistent with their proximity to the equilibrium training distribution. In contrast, as shown statistically in Fig. 5f, MatterGen structures conditioned on magnetic materials shift towards the sparse interior (~55% dense, ~45% sparse), reflecting configurational novelty detectable through geometry alone. We note that a rigorous mathematical proof of the correspondence between manifold geometry and physical observables does not yet exist; these results demonstrate the potential of platonic manifold distance functioning as a prospective detection measure for structural novelty. This geometric plausibility measure could provide a computationally efficient pre-screen for large-scale generative material searches, complementing real-space structural matching.