Data collection
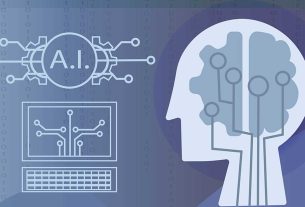
The target variable in this study is the logarithmic measure of aqueous solubility (logS) of compounds present in moles per liter (mol/L), ranging from − 5.82 (thioridazine) to 0.54 (ethambutol). This dataset, derived from the comprehensive study by Huuskonen et al32, encompasses experimental solubility values for 211 drugs and related compounds. Its selection was based on its extensive coverage and consistent experimental data, which is a prime requirement for robust training and validation of ML models. The dataset includes diverse drug classes, with their respective contributions illustrated in Fig. 1.

This pie chart illustrates the percentage distribution of drugs and derivatives across various classes: barbiturates (25%), steroids (16%), pteridines (14%), NSAIDs (8%), pyrimidines (8%), xanthine (7%), RTIs (6%), phenothiazine (4%), and “others” (12%).
In addition to MD-derived properties, the compounds’ octanol/water partition coefficient (logP) was incorporated as a critical feature extracted from scholarly literature, which is detailed in Table S4 of the supplementary materials. The inclusion of logP facilitates a comparative analysis between MD-derived properties and a well-established experimental descriptor, widely recognized for its significant influence on aqueous solubility, thereby enabling an evaluation of the efficacy of MD-derived properties. From 211 drugs in the Huuskonen et al.32 dataset, 12 Reverse-Transcriptase Inhibitors (RTIs) were excluded since the literature did not contain trustworthy logP values available. Including compounds with missing or untrustworthy logP values would be prejudicial to data integrity and adversely affect model performance. ML methods’ performance is highly dependent on having completeand accurate features, indicating the imperative of this exclusion.
MD simulations setup
Molecular dynamics simulations were conducted in the isothermal-isobaric (NPT) ensemble using the GROMACS 5.1.1 software package 33. The GROMOS 54a7 force field was employed to model the molecules’ neutral conformation, generating topology and initial coordinate files34,35. Simulations were performed within a cubic simulation box of dimensions (4 × 4 × 4) nm3, with periodic boundary conditions applied in all three spatial dimensions to ensure continuity of system properties. A variable number of water molecules modeled using the Extended Simple Point Charge (SPC/E) model were incorporated to hydrate the system.
The steepest descent energy minimization technique was applied to eliminate unfavorable atomic interactions and stabilize water molecules within the systems36. Drug molecule positions were constrained during the preparatory phase, and simulations were conducted for 10 ns to achieve equilibrium. Subsequently, simulations proceeded for an additional 20 ns in the NPT ensemble without positional constraints. Thermal conditions were maintained at 300 K using the V-rescale thermostat with a time constant of 0.1 ps37, while pressure was regulated at 1 bar, using the Parrinello-Rahman barostat with a compressibility factor of (4.5 × 10–5) bar -1 and a time constant of 2 ps38. The leap-frog algorithm was employed to integrate Newton’s equations of motion39. Long-range electrostatic interactions were computed using the Particle Mesh Ewald (PME) method40, with a cutoff radius of 1 nm set for both Coulombic and van der Waals interactions. Given the large number of compounds analyzed, results of the MD simulations for a representative compound are presented in the supplementary materials (Figs. 4S–8S).
Feature extraction
The analysis of molecular interactions during the dissolution of a compound in a solvent is essential for elucidating solubility characteristics. Based on this principle, molecular descriptors contain three energetic terms: lattice energy associated with crystal packing, the energy required to form a cavity in the solvent, and solvation energy, which collectively facilitate the modeling of aqueous solubility41. In this study, properties derived from molecular dynamics simulation were extracted to assess their impact on examining the aqueous solubility of drugs. To this end, key features were categorized into four aspects.
-
(1)
Structural properties Solvent Accessible Surface Area (SASA), Solvent Orientation around solute (Sorient), and Average number of solvents in Solvation Shell (AvgShell)
-
(2)
Intermolecular properties Average Number of Hydrogen Bonds (Hbond), Coulombic and Lennard–Jones (LJ) Energies
-
(3)
Dynamic property Root Mean Square Deviation (RMSD)
-
(4)
Thermodynamic property Estimated Solvation Free energies (DGSolv)
These features were derived from all drug molecules in the dataset and used as input features for machine learning models.
Additionally, logP was included as an intermolecular property to quantify the hydrophobicity/hydrophilicity of a substance in water. This parameter provides a comprehensive measure of solute polarity, influencing its solubility behavior.
SASA was calculated from the average geometrical structure of the output trajectory using the Connolly algorithm, quantifying the molecular surface area exposed to the solvent. A higher SASA typically correlates with enhanced solubility due to increased molecular exposure to solvent interactions. Additionally, SASA is a valuable metric, alongside other molecular properties, for accurately assessing lipophilicity.
The solvation free energy was estimated by evaluating the solvation energy contribution of each atom per unit area of the solvent-accessible surface. The solvent orientation around the solute, defined as the cosine of the angle between the solvent dipole and the drug’s center of mass as a reference vector. Furthermore, the AvgShell metric, representing the average number of solvent molecules in the first solvation shell (0 – 0.5 nm), is a critical indicator of aqueous solubility. A higher AvgShell value suggests improved solvation and greater solubility. RMSD quantities a drug molecule’s structural stability and flexibility throughout the simulation. Lower RMSD values indicate a stable molecular conformation associated with well-defined and consistent solvent interactions.
The average number of hydrogen bonds is closely related to a molecule’s solubility, hydrophilicity, and hydrophobicity. Hydrophilic molecules, typically bearing polar groups, form hydrogen bonds that enable strong interaction with water molecules. The average number of hydrogen bonds, alongside the Coulombic and Lennard–Jones energies, quantifies the strength of intermolecular interactions.
It is worth mentioning that MD solubility simulations often rely on simplifying assumptions to make the problem computationally tractable while focusing on explicit aspects of molecular behavior. A prevalent assumption is the infinite dilution condition, wherein solute concentration is sufficiently low that solute–solute interactions are negligible compared to solute–solvent interactions. This condition enables a focused analysis of a single solute molecule’s interactions with the solvent, minimizing the complexity of solute aggregation or clustering. Consequently, Coulombic and Lennard–Jones energies were computed for the system’s solute–solvent and solvent–solvent interactions. This study aims to evaluate the significance of these properties through ML algorithms to ascertain their influence on aqueous solubility and improve its prediction.
Machine learning algorithms
A well-structured machine learning project involves several key components: data preprocessing, model training, hyperparameter tuning, and model evaluation. Data cleaning and partitioning into training and test sets are performed during preprocessing to ensure data quality and enable robust model validation. Subsequently, models are trained to select the appropriate algorithm, fitting it to the training data, thereby ensuring generalization to unseen data. Hyperparameter tuning is conducted to optimize the hyperparameters of selected models, enhancing their predictive capability. Finally, model performance is quantified using appropriate metrics to assess accuracy and reliability. This organized framework establishes a rigorous foundation for developing robust and reliable machine learning models.
Data preprocessing
The phase commences with identifying and eliminating outliers to ensure the dependability and precision of statistical models. Four methods were employed for outlier detection: Interquartile Range (IQR), Z Score, Isolation Forest, and Local Outlier Factor (LOF). Following outlier elimination, the dataset was partitioned into training and test sets with an 80/20 split ratio implemented via the Stratified splitting technique from the Verstack package (version 3.9.2). This method ensured the proportional distribution of the continuous target variable, logS, in both datasets. To minimize biases due to varying feature scales, the StandardScaler method was applied to scale each feature to a mean of zero and a standard deviation of one.
Models training
Ensemble methods have emerged as a prominent approach for enhancing predictive performance, leveraging the concept of the wisdom of crowds. This approach assumes that aggregating predictions from diverse models yields superior results compared to relying solely on a single model’s prediction. Ensemble methods categorized by their training strategies are broadly classified into two main types: Parallel and Sequential ensembles.
In Parallel ensemble methods, base learners are trained independently in parallel, categorized into sub-grouped: Homogeneous and Heterogeneous Parallel ensembles, distinguished by the type of base learners and training algorithms employed. Homogeneous ensembles utilize identical base learners, whereas heterogeneous ensembles incorporate diverse learners. Sequential ensemble methods take advantage of dependencies among base learners during the training process to enhance predictive performance. Each subsequent base learner is trained to mitigate errors committed by previously trained learners, thereby improving overall model accuracy. Sequential ensemble methods can be divided into two primary approaches: Adaptive Boosting and Gradient Boosting Ensemble. Adaptive Boosting trains consecutive base learners by dynamically adjusting their weights to focus on error correction from past iterations. However, Gradient Boosting further enhances the combination of gradient descent optimization with adaptive boosting to minimize the prediction error and yield more precise results.
This study investigates the application of sequential ensemble methods to predicting aqueous solubility by employing two homogeneous parallel ensemble algorithms, Random Forest (RF) and Extremely Randomized Trees (ExtraTrees), alongside two sequential gradient boosting algorithms: Gradient Boosting Regression (GBRT) and XGBoost. Bagging or Bootstrap aggregation represents the foundational homogeneous parallel ensemble technique. It utilizes a consistent learning algorithm and dataset, training base estimators on multiple bootstrap replicates of the dataset. Ensemble diversity is achieved through bootstrap sampling, also known as sampling with replacement, which introduces variability among base learners. Predictions from ensemble models are aggregated to produce a final output via two primary methods: majority voting or model averaging.
The Random Forest (RF) algorithm, pioneered by Leo Breiman and Adele Cutler42, represents a cornerstone of ensemble learning, renowned for its robustness and reliability across various applications, including drug discovery43,44. RF constructs a collection of randomized decision trees as base learners through bagging and integrates their outputs via a collective voting process. Unlike conventional decision tree algorithms, which evaluate all features to determine the optimal split point for each decision node, Randomized Decision Trees (RDTs) employ a modified tree learning approach. RDTs introduce a random subset selection of features at each step, effectively lowering the inherent overfitting risk in isolated decision trees and yielding precise and consistent predictions. This approach reduces bias, while variance can be controlled by optimizing the hyperparameters of weak learners. Extremely randomized trees (ERT)45 is a bagging-like ensemble method that constructs an ensemble of extremely randomized decision trees. ERT differs from other tree-based ensemble methods in two key aspects: it employs random selection of features and thresholds, increasing tree variability, and uses the entire dataset to train each decision tree rather than bootstrap replicas. This combination of Randomization and ensemble averaging reduces variance more robustly than other methods. Compared to parallel ensembles, sequential ensembles intend to combine multiple weak learners to form a single robust learner. A critical component of machine learning frameworks, the loss function facilitates the identification of the best-fitting model within the training set. It can treat all training set equally or prioritize specific instances by assigning them greater weights. Gradient boosting46 a sequential ensemble method that aims trains a series of base estimators, typically decision trees by utilizing the gradient descent concept to minimize the loss value at each iteration. This process involves learning effectively from the residual errors of preceding models, repeating for a predetermined number of iterations or until predictive accuracy improvements diminish. The final prediction aggregates the contributions of all decision trees, adjusted by a learning rate known as shrinkage so that it until predictive accuracy improvements diminish. Newton boosting, another boosting approach, combines elements of AdaBoost and Gradient Boosting. Adaboost highlights misclassified examples by assigning them more wights, ensuring that subsequent base estimators prioritize these instances during the learning process. Conversely, Gradient boosting identifies misclassified examples through the gradient of the loss function, termed residuals. Newton boosting combines these strategies by employing weighted gradients to detect misclassified examples. eXtreme Gradient Boosting (XGBoost)47, an open-source framework for tree-based Newton boosting, enhances this approach through regularized loss functions, efficient tree construction, and parallelizable implementation, making it highly effective for predictive modeling tasks.
Hyperparameter tuning
The performance and complexity of machine learning models depend highly on selecting hyperparameters in different model architectures. Identifying optimal hyperparameters is often challenging and necessitates systematically exploring of diverse configurations. A conventional approach to hyperparameter optimization is grid search, which performs an exhaustive search across a predefined subset of the hyperparameter space for a given estimator. This method systematically examines every possible combination of parameters and selects the most effective set based on a performance score. However, as each hyperparameter adds a new dimension to the search space, grid search can become computationally intensive, resembling brute-force approach with significant time requirements. Cross-validation (CV)48 method is employed to avoid overfitting, typically through the GridSearchCV class in the Scikit-learn49,50. CV is a resampling technique characterized by a single parameter, k, that determines the number of groups or folds, in which dataset is divided. The training data is randomly partitioned into k unique, equally sized, independent folds. The model is trained on k-1 folds and assessed on the remaining fold, with the process repeated k times to ensure each fold serves as the validation set once. Finally, the optimum hyperparameter set is determined by averaging the k performance scores, providing a robust estimate of model performance.
Evaluation metrics
The performance of the regression models was commonly assessed using two metrics: Root Mean Square Error (RMSE) and the coefficient of determination, (R2) (Eq. 1–2).
$$RMSE\left(Y,\widehat{Y}\right)=\sqrt{\frac{1}{n}\sum_{i=1}^{n}{\left({Y}_{i}-{\widehat{Y}}_{i}\right)}^{2}}$$
(1)
$${R}^{2}\left(Y,\widehat{Y}\right)=1-\frac{\sum_{i=1}^{n}{\left({Y}_{i}-{\widehat{Y}}_{i}\right)}^{2}}{\sum_{i=1}^{n}{\left({Y}_{i}-\overline{Y }\right)}^{2}}$$
(2)
In Eq. 1–2n is the number of samples,\({Y}_{i}\) is the ith real value and \({\widehat{Y}}_{i}\) is the ith predicted value.
Feature selection
Irrelevant and redundant features can adversely affect ML model performance, compromising accuracy and computational resources. Therefore, feature selection (FS) plays a crucial role in identifying a compact subset of informative and robust features in the machine learning pipeline. This process not only reduces overfitting arising from better performance of models on training data rather than unseen data, but also improves model stability and declines its sensitivity to minor feature variations. FS methods are broadly classified into Filter, Wrapper, and Embedded techniques. Comprehension of the characteristics of different FS algorithms is beneficial for choosing the right one based on the dataset and modeling objectives. These characteristics encompass a range of factors, such as the type of analysis (univariate/multivariate), the learning paradigm (supervised/unsupervised/semi-supervised), and feature space search strategy (e.g., random, sequential, exponential, or greedy). Multivariate FS methods account for interdependencies and correlations among features, whereas univariate methods evaluate features independently, disregarding their relationships. Notably, not all FS methods belong to the univariate category51. Supervised Learning Techniques exert effort to use a combination of features with labeled target datasets to assess feature relevance, and all FS categories incorporate supervised learning within their evaluation metrics. Moreover, feature space exploration is accomplished sequentially, iteratively assessing feature contributions to the target variable. Filter-based FS methods examine feature subsets through statistical measures (i.e., correlation, consistency, information gain, or distance) without applying learning models. A fundamental aspect of filter methods is features’ ranking, which assigns each feature a relevance score. Features are retained or discarded based on a predefined threshold, informed by various ranking techniques. Spearman’s rank correlation coefficient, a statistical measure, is used to quantify a monotonic relationship between variables and targets, considering whether the relationship between two variables is linear, unlike Pearson’s correlation, which is limited to linear relationships.
Spearman’s rank is calculated per (Eq. 3) where ρ is the Spearman’s rank correlation coefficient, di is the difference between the ranks of the i-th pair of variables, and n is the number of observations. It ranges from -1 to 1, enabling straightforward interpretation of variable relationships. Values between 0 and 1 indicate a positive correlation, while values between -1 and 0 indicate an inverse correlation.
$$\rho =\frac{6\Sigma {d}_{i}^{2}}{n\left({n}^{2}-1\right)}$$
(3)
Wrapper-based FS techniques adopt a black-box approach, utilizing the performance output of classifiers, typically accuracy, as the evaluation criterion. These methods identify relevant features by assessing the improvement in model performance for selected feature subsets, as determined by machine learning algorithms. This study employs three wrapper-based approaches, namely Sequential Forward Selection (SFS)52, Sequential Backward Selection (SBS), and Recursive Feature Elimination with Cross Validation (RFECV). These methods are implemented using a range of classifiers, including: RF, XGBoost, Gradient Boosting, Extra Trees, Light Gradient Boosting, CatBoost, and AdaBoost. SFS procedure begins from an empty feature set and incrementally constructs a feature subset by evaluating the performance metrics derived from ML algorithms, continuing until a predefined number of remaining features. Conversely, SBS commences with the complete feature set and iteratively eliminates features based on poor performance scores, as determined by evaluation methods. RFE follows a logic similar to SBS, relying on feature ranking. It involves an iterative process of fitting a model using all features, and a relevance score is computed for each feature. Low-ranking features are subsequently eliminated, and the model is refitted with the updated feature subset until the optimal subset is identified. An important measure for selecting similarity between features for both forward–backward selection algorithms is the Symmetrical Uncertainty between features X and Y, as defined by (Eq. 4)51. Here, H(X) and H(Y) are the entropies of features X and Y, respectively, and H(X∣Y) is the conditional entropy of X given Y. This formulation, incorporating joint entropy H(X,Y) implicitly through the mutual information, guides the feature selection process to prioritize highly informative features.
$$\text{SU}=2\left[\frac{\text{H}(\text{X})-\text{H}\left(\left.\text{X}\right|\text{Y}\right)}{\text{H}\left(\text{X}\right)+\text{H}(\text{Y})}\right]$$
(4)
Compared to Filter-based techniques, which are straightforward to implement and require reasonable computational resources, wrapper-based techniques suggest superior learning capabilities and accuracy. Through filter methods tend to be less accurate, wrapper methods are complex and costly. Embedded techniques seek to merge the strengths of both approaches, integrating search strategies with subset assessment through ML models. This study employs two embedded techniques: Importance-based selection and Shapley Additive exPlanation (SHAP)-value-based selection53. Importance-based selection, increasingly utilized with Random Forest (RF), leverages RF’s inherent capacity to handle complex, high-dimensional datasets and capture feature interactions. Feature importance in RF is performed by measuring the impurity reduction attributable to each feature, with greater impurity reductions indicating higher feature significance. Features are then ranked by importance, and the most relevant are selected.
In contrast, SHAP, introduced by Lundberg and Lee54, draws on game theory principles. This process involves two stages: first, training ML models using the entire feature set, and second, computing SHAP values for each feature. These values are ranked in descending order to identify and select the most impactful features55.