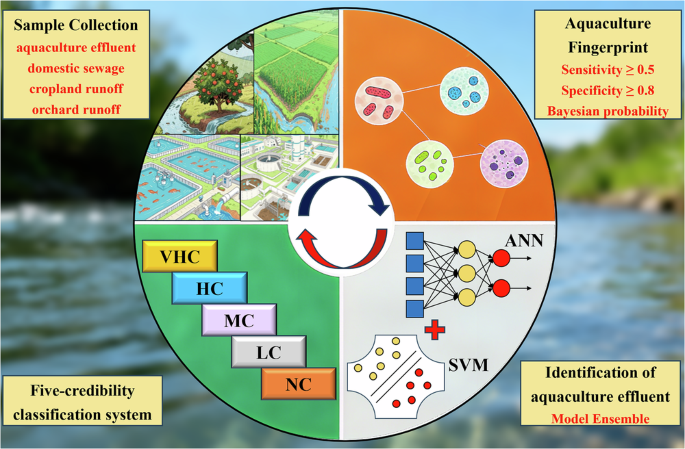
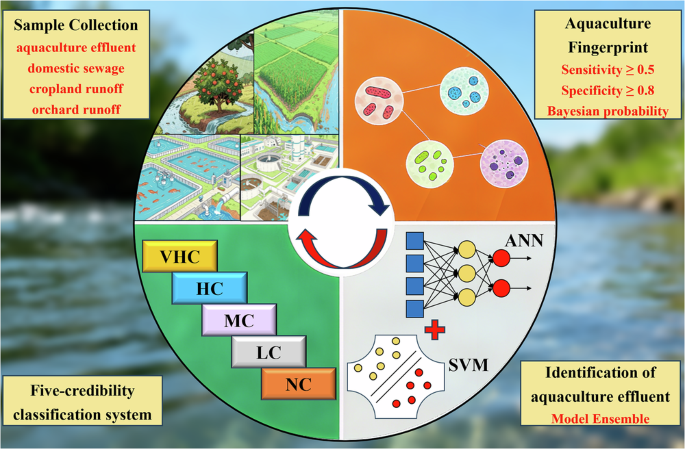
Aquaculture wastewater threatens the health of aquatic ecosystems, but tracing its source remains difficult due to the lack of specific microbial fingerprints and a preference for results. Here, we developed a microbial fingerprint-based machine learning approach for hierarchically tracking aquaculture wastewater sources. We screened four microbial taxa through high-throughput sequencing of 386 source samples (aquaculture wastewater, domestic sewage wastewater, agricultural land and orchard runoff).g_ML602J-51, g_Silisimonas, g_Lewinella and f_Furaceae) shows high sensitivity (0.512 to 0.649) and specificity (0.804 to 0.974) as a fingerprint for aquaculture wastewater. Artificial Neural Network and Support Vector Machine-Radial were the best models using fingerprint relative abundance and presence data, with accuracies of 0.8335 ± 0.0090 and 0.8221 ± 0.0047, respectively. The ensemble of models improved accuracy to 0.8706 ± 0.0175, outperforming individual classifiers by 4.44% to 5.90% and fingerprint matching by 17.29%. The predicted uncertainty was stratified into five confidence tiers to track primary aquaculture wastewater sources. Application to three coastal regions of China demonstrated the generalizability of the approach.