Overview of HAPIR
HAPIR is a machine learning model used for predicting response to ICIs. The HAPIR workflow mainly consists of four parts: (1) Extracting refined Hallmark gene sets; (2) Constructing ICIs response prediction model; (3) Evaluating performance of the model; (4) Predicting potential targets and drugs. Briefly, we extracted Hallmark gene sets enriched in DE genes between responders and non-responders, and refined these gene sets by retaining only the DE genes. Then, we calculated the activity levels of the refined Hallmark gene sets using AUCell, and applied logistic regression to train a model for predicting responses to ICI therapy. Next, we performed model evaluation by internal and external validation. Finally, we identified potential targets and predicted drugs to increase sensitivity to ICI therapy (Fig. 1).
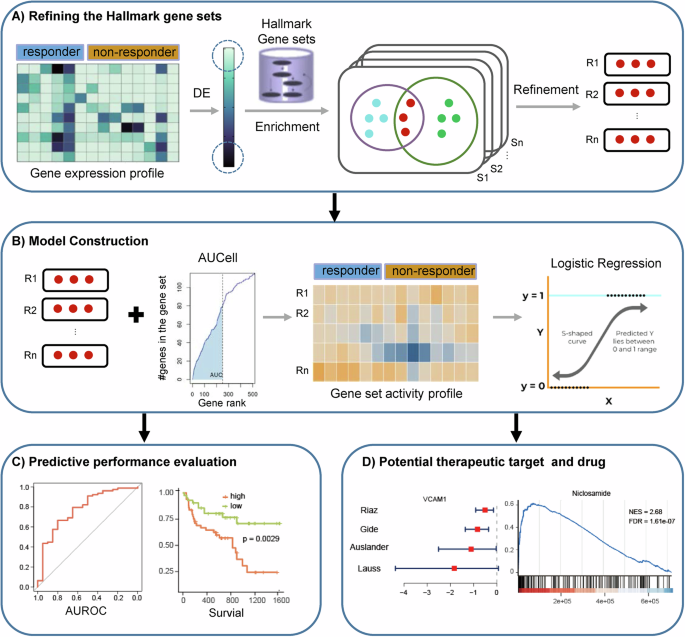
A Extracting refined Hallmark gene sets. B Constructing ICI response prediction model. C Evaluating performance of the model. D Predicting potential target and drug.
Refined Hallmark gene sets
To identify the feature genes related to ICIs immunotherapy response, we focused on Riaz et al. cohort, which contains 20 ICIs responders and 78 non-responders of melanoma. The top 200 significantly (P < 0.05) up- and down-regulated genes were obtained between responders and non-responders (Fig. 2A and Supplementary Data 1). Several genes have been reported to contribute to ICIs response. For example, Exhausted T cell with high expression of CXCL13 were correlated with favorable responses to ICIs in triple-negative breast cancer3. Granzyme K (GZMK) is a crucial mediator released by immune cells to eliminate tumor cells, and plays a significant role in inflammation19. These genes were significantly enriched in seven Hallmark gene sets (FDR < 0.05, Fig. 2B), including immune response related function (Allograft rejection, coagulation and complement), KRAS_signaling up, xenobiotic metabolism, IL2 STAT5 signaling, and epithelial mesenchymal transition (EMT). These functional gene sets have been previously implicated in immune therapy response and tumor immune evasion20,21,22,23,24,25. For example, KRAS-mutant signaling upregulates PD-L1 expression in tumor cells by enhancing the stability of PD-L1 mRNA24, directly linking oncogenic pathways to immune evasion. IL-2 signaling orchestrates the differentiation and functional regulation of both effector and regulatory CD4+ T cell subsets25, influencing tumor immunogenicity and ICI efficacy. To demonstrate the stability of enriched Hallmark gene sets, we analyzed the Jaccard Index, which quantifies the similarity between two sets by calculating the ratio of their intersection to their union, across multiple datasets. Compared to DE genes, the enriched Hallmark gene sets demonstrated a more stable pattern across the six datasets (Fig. 2C, D), highlighting their robust effectiveness as features for differentiating resistant from sensitive samples. Furthermore, a total of 77 DE genes were included in the seven Hallmark gene sets (Fig. 2E). These genes were retained to create the refined Hallmark gene sets (Supplementary Data 2) (Details in “Refining the Hallmark gene sets” of “Methods” section).
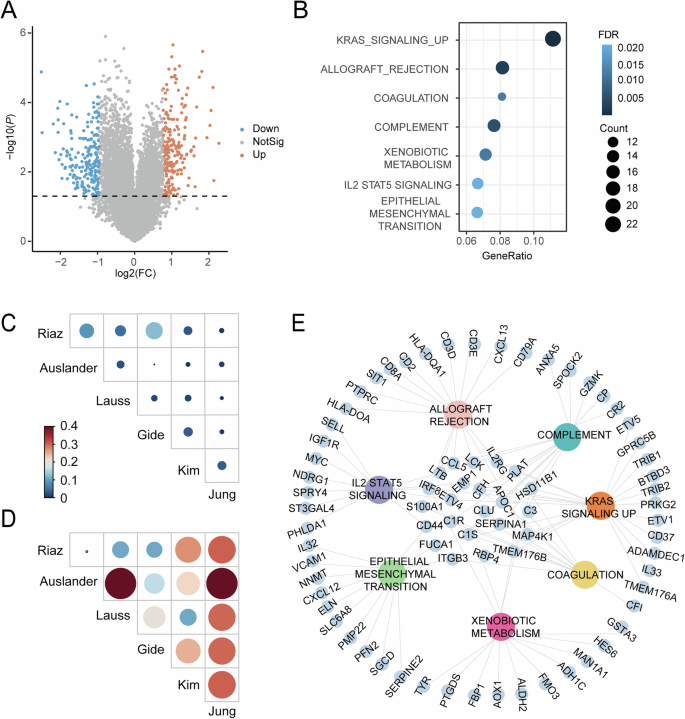
A Volcano plot of DE genes between responders and non-responders of melanoma. The top 200 significantly (P < 0.05) up- and down-regulated genes are colored by orange and blue. B Bubble plot for Hallmark gene sets enrichment analysis. C, D Heatmap of Jaccard Index for DE genes (C) and enriched gene sets (D) across six cohorts. E Network visualization for 77 DE genes across the seven refined Hallmark gene sets.
Construction and within-study cross-validation of HAPIR
Next, we calculated the activity levels of the refined Hallmark gene sets, and then constructed a machine learning model to predict immunotherapy response (details of model parameters in the “Construction and evaluation of HAPIR” of “Methods” section). To evaluate the predictive performance of HAPIR in training dataset, ten-fold cross-validation was conducted. Notably, we observed an improvement in prediction performance compared to the expression-based models of ICIs targets (PD-1 and PD-L1) (Fig. 3A). The AUROC value was 0.778 (95% CI [0.657–0.889]) for HAPIR (Supplementary Data 3), outperforming PD-1 (0.678 95% CI [0.536–0.809]) and PD-L1 (0.54 95% CI [0.403–0.685]).
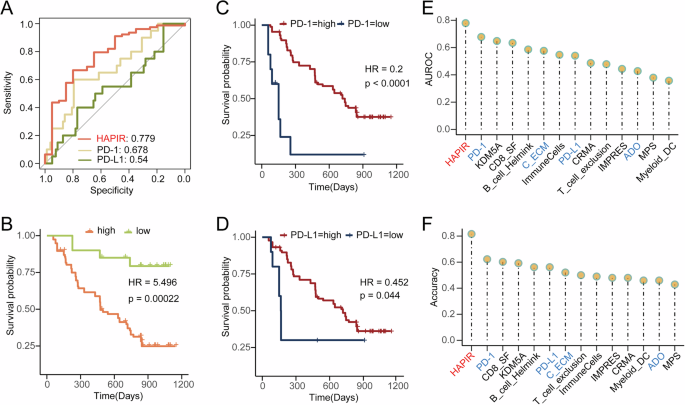
A ROC curves of HAPIR, PD-1 and PD-L1. B Kaplan–Meier survival curves generated for HAPIR. Patients were divided into high and low-predicted resistance probabilities groups. C, D Kaplan–Meier survival curves generated for the expression of PD-1 (C) and PD-L1 (D). E, F AUROC (E) and accuracy (F) comparison of HAPIR and 13 ICIs response-related biomarkers. Blue color text represents pan-cancer biomarkers and black color text represents melanoma-specific biomarkers.
We further compared the survival differences between patient groups with low- and high-predicted resistance probabilities. Patients with a low-predicted resistance probabilities exhibited significantly prolonged survival (Fig. 3B, log-rank test P = 0.0002). This survival benefit was comparable to that observed in patients with higher PD-1 expression (Fig. 3C, log-rank test P < 0.0001), and outperformed patients with high PD-L1 expression (Fig. 3D, log-rank test P = 0.04). These results indicate that HAPIR provides superior predictive performance compared to established ICIs targets.
Next, we compared the predictive performance of HAPIR with other previously identified ICIs response-related biomarkers, including melanoma-specific markers like KDM5A and CD8_SF, as well as pan-cancer biomarkers such as C_ECM, and ADO (details in Supplementary Data 4). HAPIR achieved the highest AUROC and accuracy when compared to both melanoma-specific and pan-cancer biomarkers (Fig. 3E, F). These results demonstrated that HAPIR outperforms both ICIs target expressions and other ICI response-related biomarkers in predicting immune therapy response and patient survival, making it a promising candidate for improving immunotherapy outcomes.
Validation of HAPIR based on validation and test sets
We further validated performance of HAPIR using validation and test sets. In three melanoma datasets, the AUROC exceeded 0.8 for the Lauss et al. and Auslander et al. cohorts, and it was 0.66 for the Gide et al. cohort (Fig. 4A). To assess the broader applicability and effectiveness of the HAPIR beyond melanoma, we expanded our validation to include NSCLC and STAD, both of which are suitable for immunotherapy. In these datasets, the AUROC reached 0.81 for Jung et al. cohort, and 0.62 for Kim et al. cohort (Fig. 4A). Additionally, HAPIR further exhibited a significant association with survival time in two datasets where survival data was available. In Gide et al. cohort, we observed significant differences in survival time between patient groups with low- and high-predicted resistance probabilities (Fig. 4B, log-rank test P = 0.0029), comparable to the survival association observed with PD-1 (Fig. S1A, log-rank test P < 0.0001) and PD-L1 expression (Fig. S1B, log-rank test P < 0.0001). Similarly, in Lauss et al. cohort, high-predicted resistance probabilities were associated with poorer survival (Fig. 4C, log-rank test P = 0.022), consistent with the survival outcomes for patients with lower PD-L1 expression (Fig. S1C, log-rank test P = 0.034). However, PD-1 expression was not significantly associated with survival time in this cohort (Fig. S1D, log-rank test P = 0.19). Then, we compared the predictive performance of HAPIR with the other biomarkers across these five datasets. In general, HAPIR exhibited significantly (P < 0.05) higher AUROC values than the other biomarkers, although the differences were not significant compared to PD-1 and PD-L1 (Fig. 4D). These results highlight HAPIR’s strongly robust predictive performance across multiple datasets, underscoring its potential as a valuable immunotherapy-related biomarker.
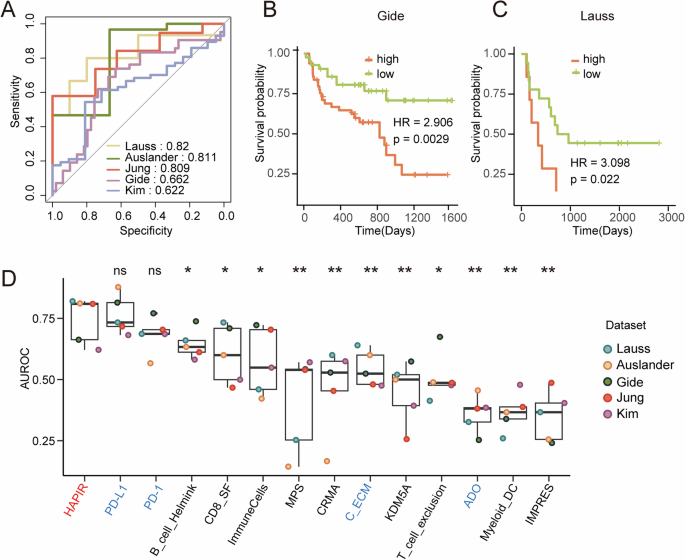
A ROC curves of HAPIR. B, C Kaplan–Meier survival curves generated for HAPIR in Gide et al. (B) and Lauss et al. (C) cohort. Patients were divided into high and low-predicted resistance probabilities groups. D Boxplot of AUROC values of HAPIR and 13 curated ICIs response-related biomarkers. Blue color text represents pan-cancer biomarkers, and black color text represents melanoma-specific biomarkers. *P < 0.05; **P < 0.01; ns: non-significant.
Comparative analysis of the refined Hallmark gene sets
To demonstrate the superiority of the refined Hallmark gene sets, we performed comparison analysis of HAPIR with models based on genes and other functional gene sets. Specifically, for gene-based models, we utilized the 77 genes included in HAPIR and the top 200 up- and down-regulated genes in combination with seven machine learning models to predict immune therapy response. HAPIR demonstrated the highest average AUROC value across the six datasets (Fig. 5A), indicating that the use of gene set scores provides a superior predictive capability compared to individual gene expressions.
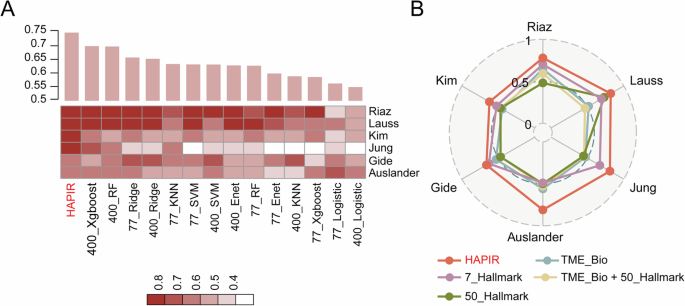
A Heatmap of AUROC comparisons between HAPIR and other methods based on genes. Bar plot showed the average AUROC across the six datasets. B Radar plot of AUROC comparisons between HAPIR and other methods of gene sets. 7_Hallmark: seven original Hallmark gene sets; 50_Hallmark: the full set of 50 Hallmark gene sets; TME_Bio: tumor microenvironment (TME)-related gene sets; TME_Bio + 50_Hallmark: combination of TME-related gene sets and 50 Hallmark gene sets.
For gene set comparison, we firstly compared the performance of the refined Hallmark gene sets with seven original Hallmark gene sets (7_Hallmark, including 1118 genes), and the full set of 50 Hallmark gene sets (50_Hallmark). For a simple comparison, we used “AUCell” and “logistic regression”, the same parameter as HAPIR to predict ICIs response for gene sets. As a result, HAPIR showed highest AUROC values across all six datasets (Fig. 5B). This highlights the effectiveness of gene set refinement in improving model performance. In addition, we also collected TME-related gene sets (TME_Bio, Supplementary Data 5) and performed a comparison. Consistently, HAPIR achieved the highest AUROC compared to models based on these TME-related gene sets, as well as those combining them with the 50 Hallmark gene sets (TME_Bio + 50_Hallmark, Fig. 5B). In summary, these findings suggested that HAPIR, with its refined gene set, provides superior predictive performance compared to those models based on genes, Hallmark gene sets, and TME-related gene sets, reinforcing its utility in immune therapy response prediction.
HAPIR predictions recapitulate the immune microenvironment in multiple datasets
Since HAPIR demonstrated accurate and robust performance across diverse datasets, we further investigated whether HAPIR predictions could accurately reflect the immune microenvironment associated with immunotherapy responses. We assessed the correlation between HAPIR predictions and immune contextures in melanoma. The levels of immune infiltration for each patient were estimated using the Quantiseq algorithm. The predicted resistance probabilities using HAPIR showed a significantly negative correlation with CD8+ T-cell proportions in Riaz et al. dataset (Fig. 6A), suggesting that patients with high-predicted resistance probabilities have low immune infiltration levels of CD8+ T-cells. This finding aligns with the well-established role of CD8+ T-cells as cytotoxic effector cells that could recognize and eliminate the tumors26.
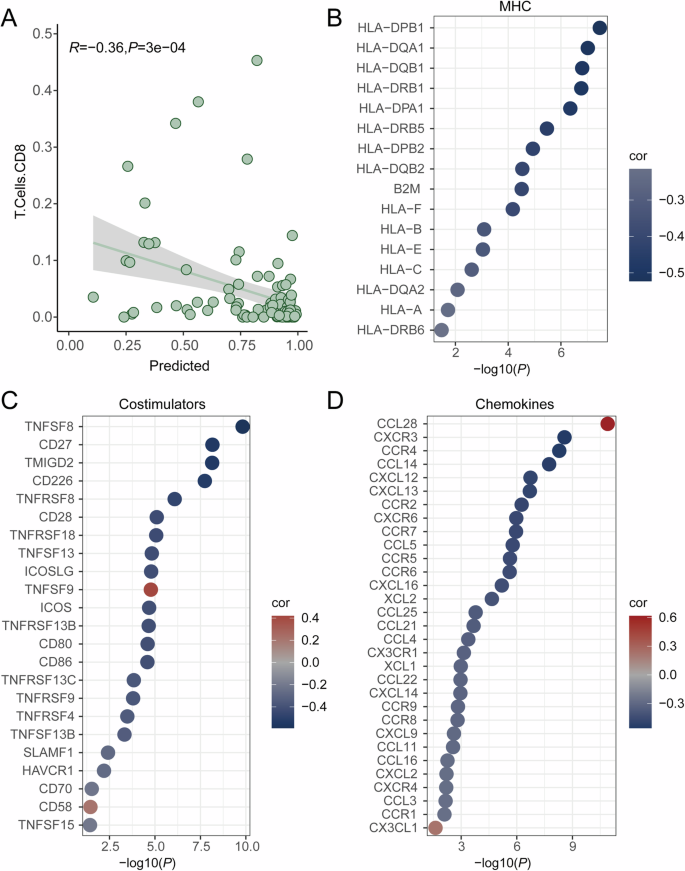
A Correlation between HAPIR predictions and the proportion of CD8+ T cell. B–D Bubble plot of the correlation between HAPIR predictions and the gene expression of MHC (B) and co-stimulators (C) and chemokines (D).
Additionally, we explored the relationship between predicted drug response and the expression levels of immune-related genes, including major histocompatibility complex (MHC) molecules, co-stimulators and chemokines (details in “Methods”). These three molecules form the classic three-signal model essential for effective T cell priming, differentiation and activation through antigen presentation, playing a pivotal role in generating anti-tumor immune responses27. Among the MHC molecules that were significantly correlated, almost all exhibited a negative correlation with the predicted resistance probabilities (Fig. 6B), as did co-stimulators (Fig. 6C) and chemokines (Fig. 6D). These results demonstrated that HAPIR could effectively capture the key features of the tumor environment, including CD8+ T-cell proportions and antigen presentation, in melanoma. We further evaluated these relationships in three independent testing datasets including Lauss et al., Auslander et al. and Gide et al. cohorts. Consistent with the Riaz et al. cohort, we observed negative correlations between the predicted resistance probabilities and both CD8+ T-cell infiltration and immune-related gene expression (Figs. S2–4). To validate these conclusions further, we examined the skin cutaneous melanoma (SKCM) dataset from The Cancer Genome Atlas (TCGA). Immune infiltration scores for TCGA-SKCM were obtained from Thorsson et al.28. We found the consistent results across multiple datasets (Fig. S5), reinforcing the robustness of HAPIR predictions. In summary, HAPIR predictions could effectively recapitulate the immune microenvironment, highlighting its potential utility in enhancing tumor treatment.
Exploration of potential therapeutic targets from HAPIR
The immune response data of knockout genes in CRISPR cohorts have been utilized to identify potential therapeutic targets in synergy with ICIs8. To explore potential therapeutic targets among HAPIR genes, the Normalized z score of 77 genes in 17 CRISPR Screen datasets were obtained from TIDE29 (http://tide.dfci.harvard.edu/prioritization/). Genes with negative values in at least half (nine) datasets were defined as immune-resistant genes, which may enhance anti-tumor immunity after knockout. Conversely, genes with positive values were classified as immune-sensitive, which may suppress anti-tumor immunity after knockout. We found 21 immune-resistant genes, such as ANXA5, CD8A and ITGB3, and 17 immune-sensitive genes, such as VCAM1, SGCA and so on (Fig. 7A). Notably, several genes have been implicated in the cancer treatment or patient survival. For example, the deletion of ANXA5 enhanced the killing of head and neck squamous cell carcinoma cells by activated CD8+ T cells30. High baseline serum levels of VCAM1 are associated with prolonged survival in NSCLC patients receiving anti-PD-1 therapy with nivolumab31. Moreover, we found that higher expression of VCAM1 was associated with a significantly improved survival in three melanoma datasets (Fig. 7B–D) and a better immune response in four melanoma datasets (Fig. 7E). These results suggested that VCAM1 could serve as a potential therapeutic target for overcoming resistance to ICIs.
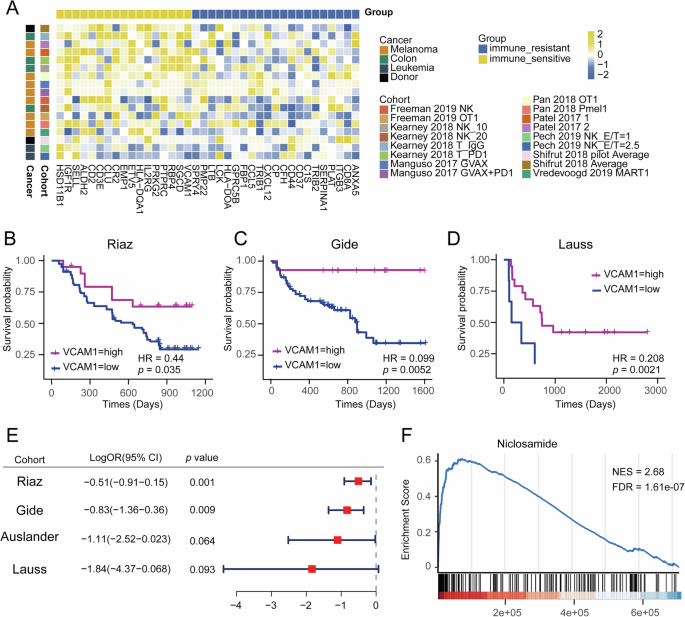
A Heatmap of z scores of 77 genes across 17 CRISPR Screen datasets. Negative (positive) z scores indicated better (worse) immune response after knockout of a specific gene. B–D Kaplan–Meier survival curves generated for the expression of VCAM1 in Riaz et al. (B), Gide et al. (C), and Lauss et al. (D) cohorts. E Forest plot of univariate logistic regression analysis of VCAM1 in melanoma datasets. F Enrichment plot of VCAM1 fold change values under Niclosamide treatment.
Then, we developed an algorithm to identify drugs capable of increasing the expression of VCAM1 using LINCS data (Fig. S6). For a specific drug, we evaluated the enrichment of its instances among the entire set of drug instances (details in “Drug screening” section). Among the 32,730 drugs evaluated, we selected those demonstrating both significant enrichment (FDR < 0.01) and strong upregulation effects (high normalized enrichment score (NES)). Notably, Niclosamide showed a significant enrichment at the fifth of top 10 small molecule instances (Fig. 7F, FDR < 0.0001, Supplementary Data 6), indicating its ability to promotes the expression of VCAM1. Niclosamide, an FDA-approved anthelmintic drug, has been demonstrated potential in enhancing cancer immunotherapy. A study by Luo et al. reported that Niclosamide can promote the activity of cytotoxic T cell and boost antitumor immune responses when combined with PD-1/PD-L1 antibody, both in vitro and in vivo, for the treatment of NSCLC32. Similarly, Zhang et al. indicated that Niclosamide enhances T cell-mediated killing of cancer cells and significantly improve the efficacy of anti-PD-1 immunotherapy in two syngeneic animal tumor models33. Additionally, Niclosamide has been shown to enhance the anti-tumor activity of NK cells against K562 and A375 tumor cell lines34. In summary, these findings suggest that Niclosamide is a promising repurposed drug to overcome resistance to immunotherapy and enhance antitumor immune responses.




