This paper focuses on optimizing the performance of the OLDPSO-SVM algorithm for AD diagnosis. The diagnostic tasks involve distinguishing between AD, NC, and MCI, specifically in AD vs. NC, NC vs. MCI-NC, NC vs. MCI-C, MCI-NC vs. MCI-C, MCI-NC vs. AD, and MCI-C vs. AD. First, different TFs are compared to enhance classification accuracy and stability. Next, the performance of GM, WM, and their combination is evaluated across various diagnostic tasks. Following that, OLDPSO-SVM is compared with other machine learning and deep learning models, highlighting its superior performance. Finally, GM and WM feature selection results are presented, identifying key brain regions for classification.
Performance comparison of different TFs in OLDPSO-SVM
The objective of this experiment, which was based on the combined use of GM and WM data, was to identify the most suitable TF for different classification tasks to optimize the performance of the OLDPSO-SVM algorithm. Specifically, the objective was to compare the performance of various TFs in terms of accuracy AvgR and StdR, and determine the optimal TF for each classification task.
As shown in Table 11, in the AD vs. NC classification task, the V3 performed the best, achieving an accuracy of 0.99111 with a StdR of 0.018754, indicating both high accuracy and remarkable stability. For the NC vs. MCI-NC task, the S1 was the most effective, with an accuracy of 0.8976. Although the S3 had a slightly lower StdR (S3: 0.047054 vs. S1: 0.049048), S1’s higher accuracy made it the superior choice. In the NC vs. MCI-C task, the V2 excelled, achieving an accuracy of 0.99069 and a StdR of 0.019628, showcasing both excellent classification performance and robustness.
For more challenging tasks, such as MCI-NC vs. MCI-C, the S1 emerged as the best performer, with an accuracy of 0.88382. However, its relatively high StdR (0.079168) suggests greater variability in classification outcomes. In the MCI-NC vs. AD task, the S1 again showed strong performance, achieving an accuracy of 0.94688 and a StdR of 0.035789, demonstrating both high accuracy and stability in this task. Lastly, in the MCI-C vs. AD task, the V2 delivered the best performance, with an accuracy of 0.87956 and a StdR of 0.038667. Despite the inherent difficulty of this classification task due to overlapping features between MCI-C and AD, the V2 effectively captured the subtle distinctions between the two groups, leading to reliable performance.
Figure 14 provides a clear visualization of the performance of different TFs across multiple classification tasks, combining both a heatmap and line plot with error bands to depict the accuracy and uncertainty of each TF. The heatmap highlights differences in accuracy, with darker shades indicating higher classification performance, while the line plot further reveals the trends and stability of the TFs across different tasks. Compared to Table 11, the error bands in the line plot offer a clearer representation of the variability in classification outcomes. This is particularly evident in more challenging tasks, such as NC vs. MCI-NC and MCI-NC vs. MCI-C, where the performance of the TFs shows greater fluctuation. In contrast, the AD vs. NC task demonstrates consistently high accuracy with minimal variability.
Figure 15 illustrates the fitness convergence of the OLDPSO-SVM algorithm across six different classification tasks, using eight different TFs. The objective is to identify the optimal TF for each classification task. The horizontal axis indicates the number of algorithm iterations, while the vertical axis shows the average fitness values obtained after ten independent runs. The results indicate that V3 performs best in the AD vs. NC task, with the fastest convergence and the lowest fitness value. In the NC vs. MCI-NC, MCI-NC vs. MCI-C, and MCI-NC vs. AD tasks, S1 demonstrates the lowest fitness value, making it the most effective TF. For the NC vs. MCI-C and MCI-C vs. AD tasks, V2 achieves the fastest convergence and the lowest fitness value, making it the optimal TF in these cases. In subsequent experiments, we will adopt the most effective TF identified for each classification task.
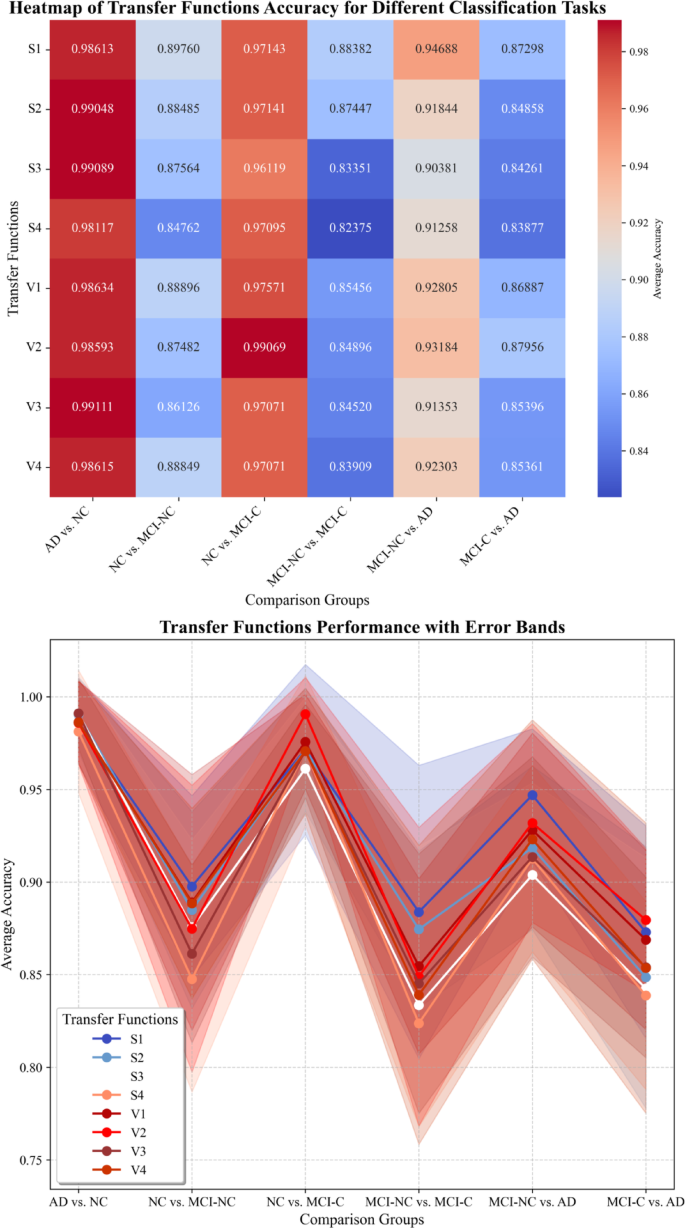
Performance comparison of TFs across multiple classification tasks.
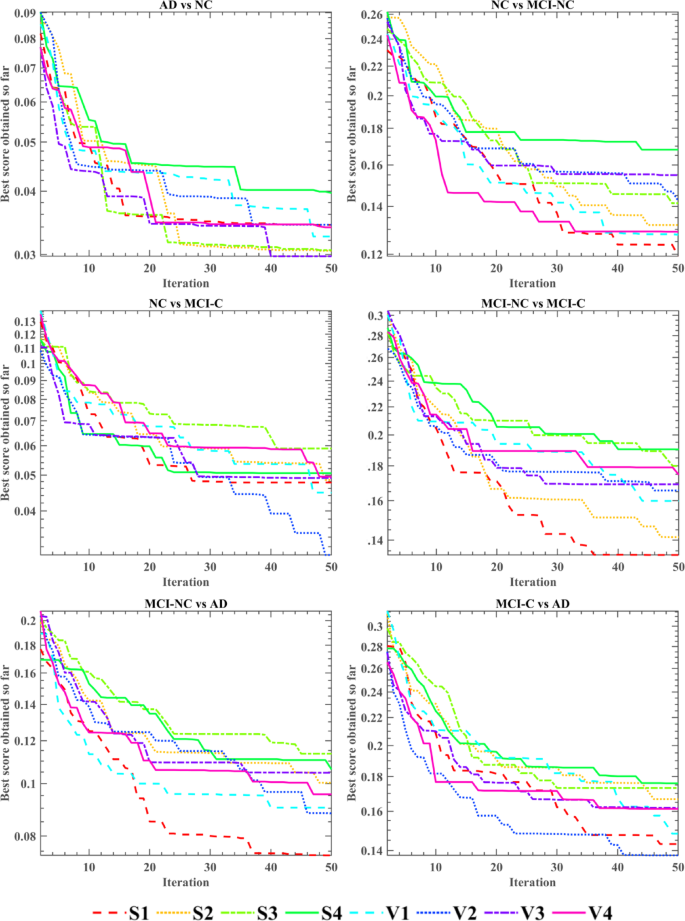
Fitness convergence of OLDPSO-SVM algorithm using different TFs across six classification tasks.
Performance evaluation of GM, WM, and combined models
This study aims to assess the classification performance of GM and WM in distinguishing between AD, NC, and individuals with MCI. The study compares the effectiveness of using GM, WM, and their combination (GM + WM) across various classification tasks. As shown in Table 12, the GM + WM combined model demonstrates superior performance in all classification tasks.
In the classification between AD and NC, the GM + WM model achieved an accuracy of 99.11%±1.88%, with sensitivity and specificity around 99%, outperforming the GM (98.10%±3.33%) and WM (93.96%±6.19%) models. For NC vs. MCI-NC, the GM + WM model also performed best, with 89.76%±4.90% accuracy, 91.82%±10.00% sensitivity, and 87.36%±12.62% specificity, surpassing the individual GM and WM models. In NC vs. MCI-C, the GM + WM model achieved an accuracy of 99.07%±1.96%, again outperforming the GM and WM models. For the more challenging MCI-NC vs. MCI-C task, GM + WM achieved 88.38% ± 7.92% accuracy, delivering the best performance despite the subtle differences between the groups. In MCI-NC vs. AD, the GM + WM model showed strong results, with 94.69% ± 3.58% accuracy, outperforming the GM and WM models. Finally, for MCI-C vs. AD, the GM + WM model achieved 87.96% ± 3.87% accuracy, again surpassing the individual models.
In Fig. 16, the mean value and standard deviation for various classification metrics across different classification tasks (AD vs. NC, NC vs. MCI-C, and MCI-NC vs. AD) are presented. The results demonstrate that the combined GM and WM model (GM + WM) consistently outperforms the individual GM and WM models, with mean values approaching 1 in key metrics such as accuracy, specificity, Precision, and F-measure, indicating superior classification performance. Moreover, the standard deviation results highlight the stability and consistency of the GM + WM model. In several metrics, the GM + WM model exhibits significantly lower standard deviations compared to the individual GM and WM models, particularly in MCC and F-measure, indicating less variability and more consistent classification outcomes. In contrast, the individual GM and WM models display larger standard deviations, especially in Sensitivity and other metrics, with the WM model showing particularly high variability, indicating less stable performance across different experiments.
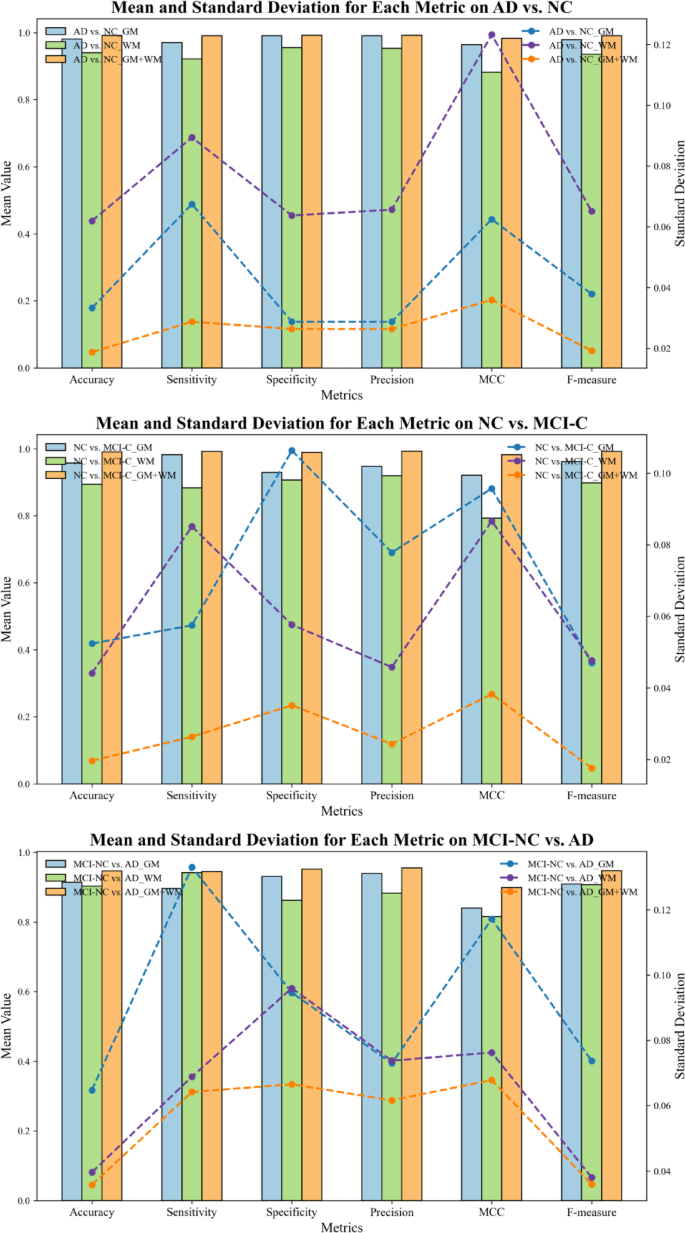
Mean and standard deviation of classification metrics for GM, WM, and GM + WM models in distinguishing AD, NC, and MCI.
Overall, the GM + WM model not only achieves excellent performance in terms of classification metrics, but its lower standard deviations suggest higher robustness, with more stable and reliable results across experiments. These results suggest that combining GM and WM provides a more comprehensive representation of brain pathology, leading to significantly improved accuracy in disease classification.
Comparison with classical machine learning algorithms
The objective of this experiment is to evaluate the performance of the proposed OLDPSO-SVM algorithm in comparison with other commonly used machine learning models, specifically Back Propagation Neural Network (BP), Classification and Regression Tree (CART), Random Forest (RandomF), AdaBoost, and Extreme Learning Machine for Feature Selection (ELMforFS), for classifying individuals with AD, MCI, and NC. The aim is to identify a more effective method for early diagnosis. As shown in Fig. 17, the results demonstrate that the proposed OLDPSO-SVM algorithm outperforms these baseline models across various classification tasks, particularly in key metrics such as accuracy, sensitivity, specificity, precision, MCC, and F-measure. OLDPSO-SVM exhibited superior stability and classification capability in addressing AD-related classification problems, highlighting its potential for significant application in early diagnostic processes.
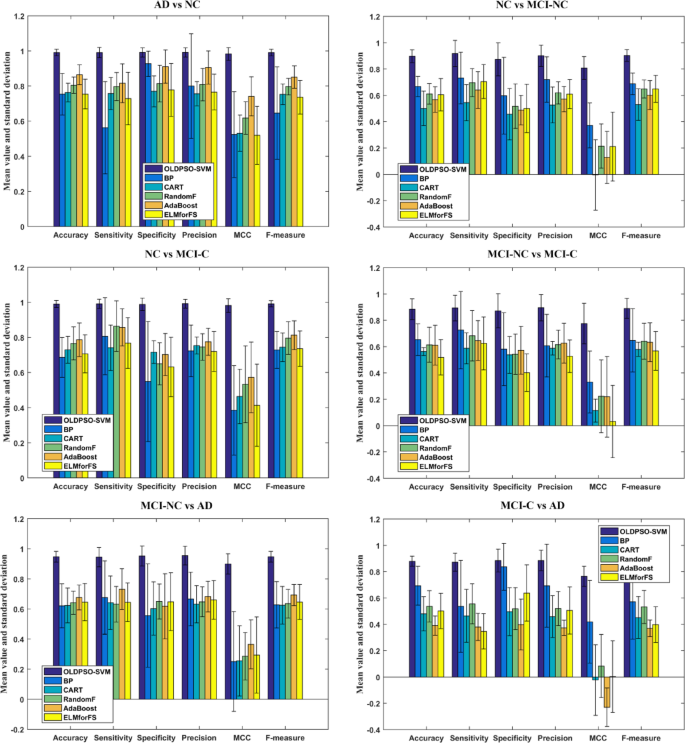
Performance comparison of OLDPSO-SVM and baseline models for AD classification.
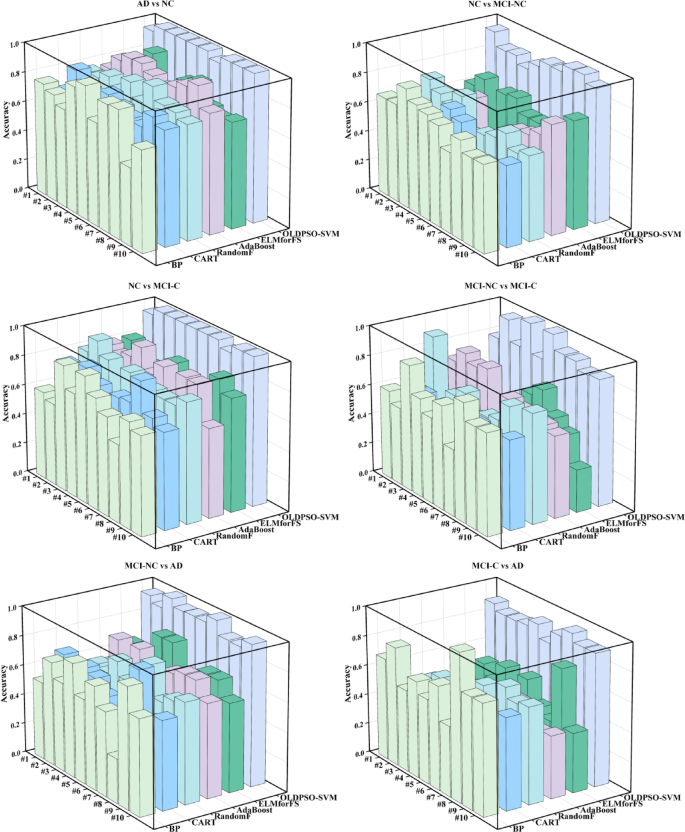
Accuracy performance of various machine learning models across 10-fold cross-validation for different AD classification tasks.
Figure 18 presents the accuracy performance of various models across each fold in a 10-fold cross-validation setting for different classification tasks. Each color and position in the bars represent a distinct machine learning model. The figure reveals that the OLDPSO-SVM model maintains a consistently high and stable accuracy across the majority of folds in each task, significantly outperforming the other machine learning models. In contrast, the accuracy of the other models exhibits greater variability, particularly in more complex tasks, such as MCI-NC vs. MCI-C and MCI-C vs. AD, where the performance differences between models become more pronounced. Overall, the results indicate that OLDPSO-SVM demonstrates robust classification capability and reliability in handling AD-related classification tasks.
Comparison with other existing methods
To evaluate the performance of the proposed OLDPSO-SVM in AD recognition, we conducted a comparative analysis against previous deep learning methods, such as CNNs and Transformers, on the ADNI dataset, as summarized in Table 13. It is important to note that due to differences in data processing protocols across studies, this comparison serves as a reference. Despite these variations, our results demonstrate that the proposed OLDPSO-SVM achieved a satisfactory accuracy, second only to Liu et al.56 (99.4%) and outperforming most existing methods reported in the literature. Although CNNs and Transformers have advantages in end-to-end feature learning, the core strength of OLDPSO-SVM lies in its unique feature selection capabilities and interpretability. By precisely identifying key brain regions associated with AD, this model can provide biologically meaningful insights for medical diagnoses. Furthermore, it demonstrates robust classification performance on small sample datasets, significantly outperforming deep learning models that rely on large-scale data training, such as the CNN developed by Yagis et al.50, which achieved only 73.4% accuracy. This broader comparison further underscores the effectiveness and potential advantages of OLDPSO-SVM in AD recognition.
GM and WM feature selection for AD diagnosis using OLDPSO-SVM
Figure 19 presents the GM and WM feature selection results across AD, MCI, and NC groups, derived from the OLDPSO-SVM algorithm using 10-fold cross-validation, repeated 10 times. The vertical axis represents the total number of times each brain region was selected, while the horizontal axis (R1 to R90) corresponds to the 90 regions of interest (ROIs) defined by the AAL template, as described in Table 2. These feature selection results underscore the importance of specific brain regions in the classification tasks, with regions frequently selected indicating their crucial role in distinguishing between AD, MCI, and NC groups. To offer a more intuitive depiction of the results shown in Figs. 19 and 20 provides 3D visualizations of the top five most frequently selected brain regions for each classification task. These 3D brain maps vividly illustrate the structural differences in both GM and WM across the AD, MCI, and NC groups.
In the AD vs. NC classification task, the top five GM regions selected were the left and right hippocampus (R37, R38), the left supplementary motor area (R19), the left middle frontal gyrus (R8), the left and right inferior temporal gyrus (R89, R90), and the right heschl gyrus (R80). The top five WM regions selected were the left parahippocampal gyrus (R39), the right inferior occipital gyrus (R54), the left insula (R29), the right pallidum (R76), the left precentral gyrus (R1), and the right amygdala (R42).
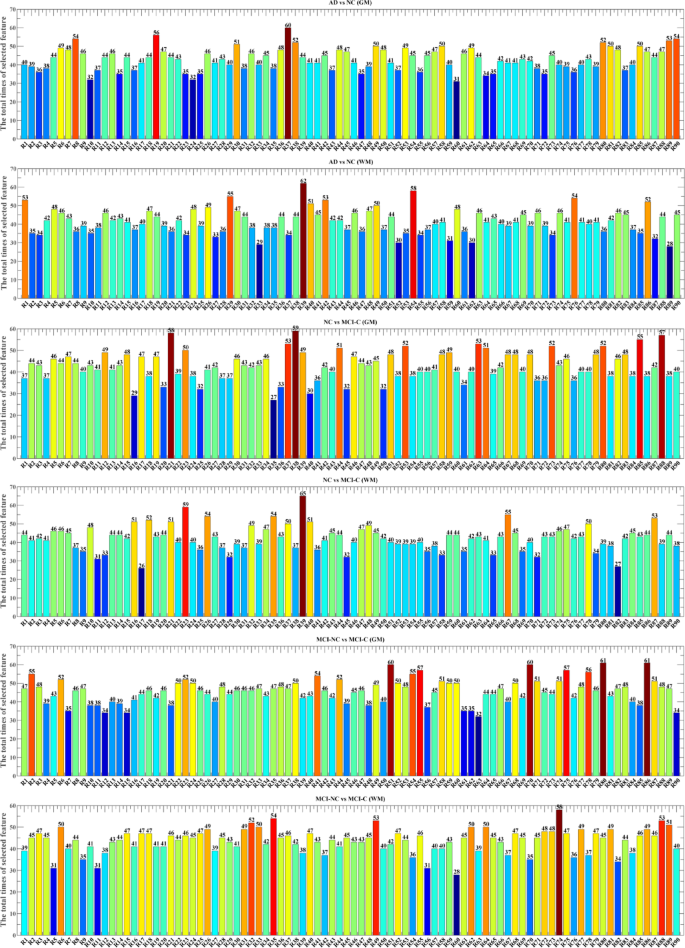
Comparison of GM and WM feature selection in AD, MCI, and NC.
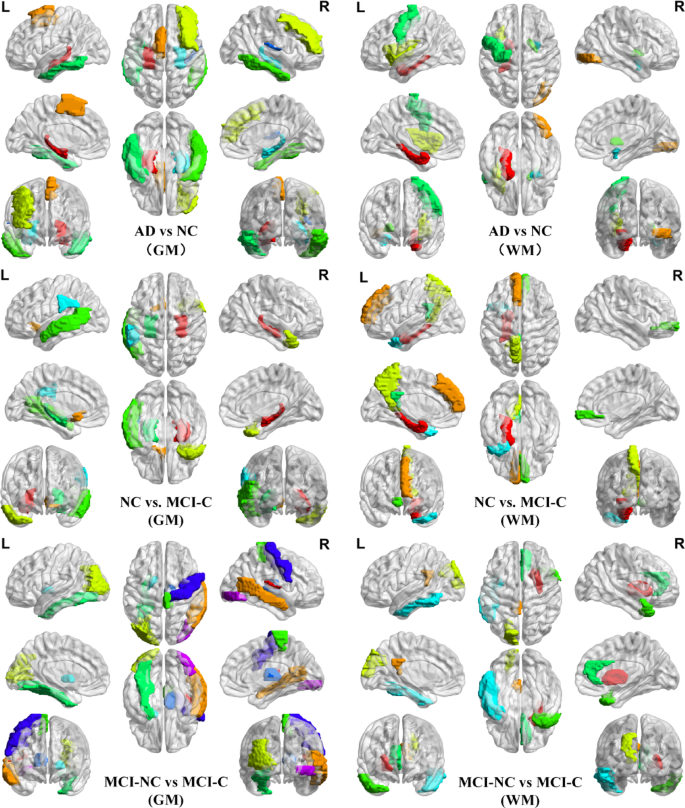
Comparative analysis of structural differences in GM and WM among AD, MCI, and NC. The structural differences were visualized using BrainNet Viewer with version 1.7 (http://www.nitrc.org/projects/bnv/).
In the NC vs. MCI-C classification task, the top five GM regions selected were the left and right hippocampus (R37, R38), the left olfactory (R21), the right temporal pole (R88), the left middle temporal gyrus (R85), and the left supramarginal gyrus (R63). The top five WM regions selected were the left parahippocampal gyrus (R39), the left medial superior frontal gyrus (R23), the left precuneus (R67), the right medial orbitofrontal cortex (R26), the left posterior cingulate gyrus (R35), and the left middle temporal pole (R87).
In the MCI-NC vs. MCI-C classification task, the top five GM regions selected were the right heschl gyrus (R80), the right middle temporal gyrus (R86), the left middle occipital gyrus (R51), the right paracentral lobule (R70), the left fusiform gyrus (R55), the left pallidum (R75), the right thalamus (R78), the right precentral gyrus (R2), and the right inferior occipital gyrus (R54). The top five WM regions selected were the right putamen (R74), the left posterior cingulate gyrus (R35), the left superior occipital gyrus (R49), the right middle temporal pole (R88), the right anterior cingulate gyrus (R32), and the left inferior temporal (R89).
This detailed selection of GM and WM regions highlights the key structural changes in the brain across the different classification tasks, emphasizing both hemispheric and regional involvement in the progression from NC to MCI and AD.




