Subject characteristics
Data collected include follow-up imaging studies and clinical data of 75 BM patients from 5 different medical centers. Inclusion criteria were defined as follows: deceased adult patients with a pathologically confirmed diagnosis of BM between 1 January 2005 and 31 December 2021, at least post-contrast T1- w High-resolution sequence (pixel spacing ≤2 mm, slice thickness ≤2 mm, no gaps between slices, no noise or artifacts in images, and basic clinical data (age at diagnosis, sex, and available treatment plans, survival rates, etc. Primary tumor was non-small cell lung cancer (NSCLC) (n = 38), small cell lung cancer (SCLC) (n = 5), breast cancer (n = 22) , melanoma (n = 6), ovarian cancer (n = 2), renal cancer (n = 1), uterine cancer (n = 1).
The 75 included patients had a total of 260 BMs and a total of 637 imaging studies. Of these, 593 studies were semi-automatically segmented as described below.
image acquisition
All post-contrast T1-W sequences were obtained after intravenous administration of a single dose of contrast agent. The 593 segmented imaging sequences were acquired with a 1-T (n = 8), 1.5-T (n = 550), or 3.0-T (n = 35) MR imaging scanner. Regarding MR imaging vendors, General Electric (n = 225), Philips (n = 197), and Siemens (n = 171) medical systems were used. Other image parameters are described in Table 1.
Segmentation procedure
Segmentation was performed using an in-house semi-automated segmentation procedure26,28Tumors were automatically delineated using a gray-level threshold chosen to identify maximal contrast-enhancing tumor volume. A biomedical engineer/applied mathematician (BO-T.) then carefully modified each segmentation slice-by-slice using a brushing/pixel removal tool. The segmentation process is summarized in Figure 1. The results were cross-checked by his three investigators (DM-G., JP-B., VMP-G.) with her over 7 years of expertise in MRI, and then corrected by any I was. Study participating radiologists (BA, AOM, DA, LAP-R., EA). In this procedure he used raw medical images in DICOM format and therefore has not been modified to perform tumor segmentation.
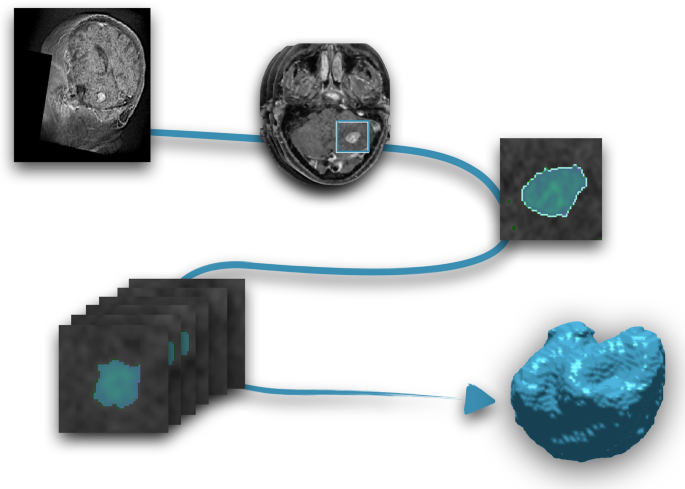
Image segmentation procedure. From the MR images (T1-W with contrast), each slice was semi-automatically segmented and manually corrected. Once all slices were segmented, the final step was her three-dimensional reconstruction of the tumor.
Clinical data and anonymization
Clinical data of 75 patients were collected. For each patient, we examined age and sex at diagnosis, primary tumor type and subtype, molecular markers (eg her EGFR, ALK, ROS1 for lung cancer), and tumor stage.Also, the GPA index is1,3, was included in the subset of institutions. For each BM, the ID (a number to distinguish it from her other BM in the same patient), location in the brain (frontal, temporal, parietal and occipital, left and right), date of appearance on MRI, and treatment received were Recorded. Recorded. For each treatment, treatment type, dose, minutes, start date and end date were recorded. Dates of available follow-up MRI studies were also included. Radiation necrosis was identified in 39 lesions.
The first step of data anonymization was performed at the institution of origin of the data. Such steps included the anonymization of patient and center data. Further de-identification was performed using the Clinical Trials Processor at the Medical Imaging Resource Center.36Within that step, all private DICOM tags, all tags containing sensitive or identifying information, and all dates were checked for all subjects, the imaging study in which the first BM was first identified was January 1900. Changed to correspond to 1 day. Anonymized times are: This means that negative numbers identified prediagnostic treatment for BM. Relative differences in the times of various events for each patient were preserved. The final step in anonymization was a falsification process that made facial reconstruction impossible. After this entire process, patient records were finally reviewed independently by her three authors (BO-T., JP-B., and JAR-R.).
Morphological parameters
Various morphological parameters were calculated from the segmentation and collected in databases such as:
volume
Three different types of volumes were calculated for each focal point.ⅤCE), necrotic (or non-enhanced) (ⅤN.) and total volume (Ⅴ=ⅤCE+ⅤN.).
Contrast enhancement spherical rim width (CE rim width)
Acquired per focus from CE and necrosis volumes
$${\rm{C}}{\rm{E}}\,{\rm{r}}{\rm{i}}{\rm{m}}\,{\rm{w}}{\ rm{i}}{\rm{d}}{\rm{t}}{\rm{h}}={}^{3}\sqrt{\left(\frac{3({V}_{CE }+{V}_{N})}{4\pi }\right)}-{}^{3}\sqrt{\left(\frac{3{V}_{N}}{4\pi } \right)}.$$
By assuming that areas of necrotic tissue and whole tumors are spherical, this function calculates the average width of CE areas. Additional information and figures for tumors with high and low CE rim width can be found at29.
water surface
It is obtained by reconstructing the tumor surface from a discrete set of voxels that characterize the tumor using Matlab’s “isosurface” command.
surface regularity
It is the dimensionless ratio between the volume of a segmented tumor divided by the volume of a spherical tumor with the same surface. For each focus it was calculated as:
$${\rm{S}}{\rm{u}}{\rm{r}}{\rm{f}}{\rm{a}}{\rm{c}}{\rm{e} }\,{\rm{r}}{\rm{e}}{\rm{g}}{\rm{u}}{\rm{l}}{\rm{a}}{\rm{r }}{\rm{i}}{\rm{t}}{\rm{y}}=6\sqrt{\pi}\frac{{\rm{T}}{\rm{o}}{\ rm{t}}{\rm{a}}{\rm{l}}\,{\rm{V}}{\rm{o}}{\rm{l}}{\rm{u}}{ \rm{m}}{\rm{e}}}{\sqrt{{({\rm{T}}{\rm{o}}{\rm{t}}{\rm{a}}{\ rm{l}}{\rm{s}}{\rm{u}}{\rm{r}}{\rm{f}}{\rm{a}}{\rm{c}}{\rm {e}})}^{3}}}.$$
This parameter ranges from 0 (for tumors with very rough surfaces) to 1 (for spherical tumors). Additional information and figures for tumors with high and low CE rim width can be found at17.
maximum diameter
This provides the maximum longitudinal measurement of the tumor and is calculated for each focal point as the maximum distance between two points on the surface of the CE tumor.
Radiomic-based features
A total of 110 different features were extracted using the open-source Python package PyRadiomics version 2.2.0.37This feature dataset contains 16 shape descriptors and various measures of intensity distribution and texture within the segmentation labels. Intensity features were derived from simple first-order statistics (19 features), gray-level co-occurrence matrix (GLCM, 24 features), gray-level run-length matrix (GLRLM, 16 features), gray-level size This includes – Zonal Matrix (GLSZM, 16 features), Neighborhood Graytone Difference Matrix (NGTDM, 5 features), and Gray-Level Dependent Matrix (14 features). Features were extracted from the original image sequence after z-score normalization, intensity scaling by a factor of 100, and subsequent shift by 300 (i.e., 3 standard deviations), and most intensity values were obtained from the primary features and geometry tolerance Check that it is positive for the value 0.04. Other specific tasks may require different feature extraction procedures18.
No voxel resampling was used before feature extraction to keep the information as unaltered as possible. The algorithm for extracting image features is shared, so users can apply resampling and redo the extraction.
Atlas location feature
All subjects were aligned to the MNI atlas space using affine registration38 Using mri_robust_register39The centroids of individual metastatic lesions are listed and can be used to efficiently identify the location and affected brain regions.
ethical approval
We have complied with all relevant ethical regulations and all subjects included in the study are deceased. Human data were obtained in the framework of the study OpenBTAI (Brain Tumor Open Database for Research in Artificial Intelligence). This is a retrospective, multicenter, nonrandomized study approved by the corresponding institutional review board: Fundación Instituto Valenciano de Oncología (2021-05), Hospital Universitario HM Sanchinarro (21.06.1858-GHM). ), Hospital Universitario 12 de Octubre (21/711), Hospital General Universitario de Ciudad Real (12/2021), Hospital Regional Universitario de Málaga (24/06/2021), Hospital Universitario y Politécnico La Fe (2021-504-1) ), MD Anderson Cancer Center (01/06/2021), Hospital Universitario de Salamanca (2021 10 879), Complejo Hospitalario Universitario de Toledo (29/9/2021-770) and Hospital Universitario Marqués de Valdecilla (14/14/2021) date – October 9, 2021).



