Study design and ethical considerations
Data for this study were collected from the records of patients at the Department of Pediatric Surgery, Split University Hospital, who underwent appendectomy for suspected acute appendicitis. Data were collected from January 2019 to July 2023. The Ethics Committee of Split University Hospital approved the study protocol (approval number 500-03/22-01/188, approval date 28 November 2022), which complies with the 1975 World Health Organization Declaration of Helsinki, as revised in 2013, and the International Conference on Harmonization guidelines for good clinical practice. Patient anonymity was strictly maintained.
Inclusion and Exclusion Criteria
All pediatric patients (aged 0-17 years) referred for emergency surgical treatment for suspected acute appendicitis were included in the study. Exclusion criteria were age >17 years, presence of significant comorbidities (chronic cardiac, renal, or gastrointestinal disease), and body mass index (BMI) >35 kg/m.2incidental appendectomy, and PHD that was neither appendicitis nor histologically normal appendix (e.g., neuroendocrine tumor or intestinal parasitosis), or when histopathological reports were not available.
Research objectives and data preparation
We set three research objectives:
-
Development of a predictive model for negative and positive cases of acute appendicitis based on PHD.
-
Comparison of appendicitis prediction model performance with acute (AIR) score.
-
Differentiate cases of complicated from uncomplicated appendicitis and negative appendectomy.
The initial feature set included patient data, complete and differential blood count features, biochemistry measurements (e.g., sodium concentration and C-reactive protein (CRP)), and laboratory features (presence or absence of abdominal pain, rebound tenderness, or guarding). All patients underwent surgical treatment, with the majority of cases involving a three-port laparoscopic appendectomy. In a minority of cases, a standard open appendectomy was performed. The surgical technique was selected based on surgeon preference.
Twenty-two features were included for model training and analysis: age, sex, duration of symptoms, height, weight, BMI, temperature, white blood cell (WBC) count, CRP, neutrophil percentage, lymphocyte percentage, platelet/lymphocyte ratio (TLR), neutrophil/lymphocyte ratio (NLR), mean platelet volume (MPV), mean corpuscular hemoglobin concentration (MCHC), sodium concentration, and presence or absence of signs and symptoms such as rebound tenderness, vomiting, nausea, and pain migration. The outcome feature was PHD. Patients with a confirmed histopathological diagnosis (PHD) of acute appendicitis were classified as having uncomplicated appendicitis (catarrhal or celluloid) or complicated appendicitis (gangrenous or gangrenous perforating) based on the histopathological report.
Features were excluded if more than 30% of the data were missing or if they were highly correlated with some of the selected features, such as hemoglobin count, hematocrit, and mean corpuscular hemoglobin.
In total, the dataset consisted of 614 pediatric patients. Patients were excluded if they had two or more missing values in features that were determined to be highly important based on previous studies (features neutrophil percentage, lymphocyte count, white blood cell count, CRP, and sodium concentration). After applying the above exclusion criteria, 551 patients were included in the final analysis. Of the 551 patients, 47 were appendicitis negative, 252 were uncomplicated appendicitis, and 252 were complicated appendicitis, indicating that the dataset was unbalanced. The final dataset contained missing values as shown in Table 1.
Train, optimize, and validate predictive models
We tested three ML algorithms: Random Forest, eXtreme Gradient Boosting (XGBoost), and logistic regression. The first two algorithms were selected based on their known effectiveness in handling tabular data and imbalanced datasets, and logistic regression was chosen as the baseline model.16,17,18.
A nested cross-validation approach was used to train and validate the model, with 5-stage inner and outer cross-validation repeated 10 times (Figure 1). As a result, each outer fold was split into a training set (80%) and a test set (20%), and the target variable was stratified due to the unbalanced nature of the dataset. Inner cross-validation was performed on the training set of each outer fold, to tune the hyperparameters and shift the threshold. Missing values were imputed utilizing the Bagged Trees algorithm (using the 'step_impute_bag' function in the 'recipes' package in the R programming language). To avoid data leakage, imputation of the validation and test datasets was performed only based on the estimates calculated from the training dataset.
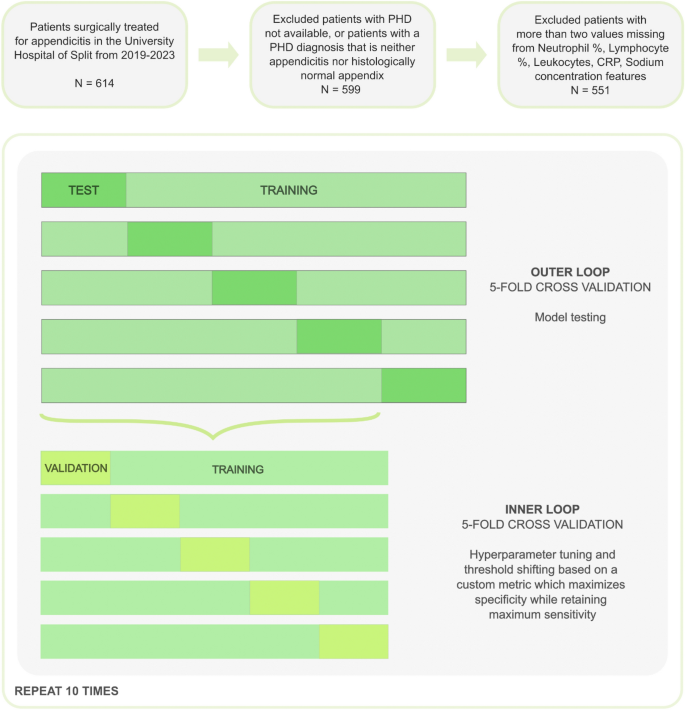
Research workflow (feature selection, model training, validation)
To address data imbalance, a threshold shift in the receiver operating characteristic curve (ROC) was used. Because our goal was to build a model that would improve the detection of negative cases without sacrificing the ability to detect positive cases, we chose to perform the threshold shift based on a custom metric that tunes the model's hyperparameters and identifies the most specific point on the ROC curve where the sensitivity remains at 1 (max(specificity/sensitivity) = 1).
The threshold applied to the test set is determined as the second lowest threshold obtained from internal cross-validation. It is a more conservative approach than adopting the average threshold in terms of maintaining a high sensitivity on the test set. In other words, we impose a constraint that the model must accurately diagnose true appendicitis for 100% of the patients in the training data. In this way, when the model outputs a negative diagnosis, we can be as certain as possible that it is a true negative and not a false negative.
The mean and standard error of the target metric were obtained by averaging the results of 50 outer-fold test sets (10 repetitions of 5-fold outer cross-validation). The best model was selected based on the average specificity score while maintaining the highest sensitivity.
Modeling was performed using Python (version 3.9.5, Python Software Foundation, Wilmington, DE, USA) and R programming languages (R Core Team, 2023, Vienna, Austria). The Python packages used were “numpy”, “pandas”, “scikit-learn”, and “xgboost”. Within the R ecosystem, the “dplyr”, “tidyr”, “purrr”, “ggplot2”, “plotly”, “tidymodels”, “ranger”, “xgboost”, and “fastshap” libraries were used. For the random forest models, the hyperparameters tuned were the number of variables available for splitting at each node and the minimum node size to split. For XGBoost, besides the two hyperparameters already mentioned, the learning rate, gamma, number of trees, subsample ratio, and maximum tree depth were also tuned.
Feature importance and model explainability
The importance of individual features was assessed using Shapley Additive Explanation (SHAP) values.19These values were obtained using 10,000 simulations using the “fastshap” package. The Shapley values measure the impact of each feature on the shift in prediction from the average model prediction to the final model output by focusing on a specific feature within a union of many features. As a result, these values help explain predictions from ML models.
Statistical analysis
The normality of data distribution was tested using the Kolmogorov-Smirnov test. For comparison of two groups, the significance of the difference was tWe use the -test for normally distributed data and the Mann–Whitney test for deviations from normality. When comparing two or more groups, we use ANOVA if data from all groups are normally distributed, otherwise we use the Kruskal–Wallis test. We use the chi-square test for non-numeric features. We use 0.05 as the p-value threshold when considering statistical significance. We use the R programming language for all statistical analyses and visualizations.
Ethical approval
The Ethics Committee of Split University Hospital approved the study protocol (approval number: 500-03/22-01/188, approval date: 28 November 2022). Informed consent was not applied for this study.