Data acquisition
The study was approved by the Institutional Review Board (IRB) of the University of Illinois at Chicago (UIC) and is in adherence to the ethical standards set forth in the Declaration of Helsinki. The en face OCTA images used for this study are 6 mm × 6 mm scans collected at UIC. In this study, we have two different datasets. The training dataset which consists of 104 OCTA images and their ground truths (68 control, 12 mild DR, 11 moderate DR, and 13 severe DR scans) is planned to be used for training and validation of the CNN. The test dataset which consists of 64 OCTA images without ground truths (25 eyes from 17 control participants, 18 eyes from 13 NoDR patients, and 21 eyes from 18 mild DR patients) is planned to be used for qualitative testing of the CNN and quantitative analysis with the focus on early detection of DR. Table 1 summarizes all participant demographics and diabetes-related parameters in the test dataset. Control subjects and diabetic patients without and with DR in different stages were recruited from the UIC retina clinic. The patients present in this study are representative of a university population of diabetic patients who require clinical diagnosis and management of DR. Subjects who were 18 years of age or older met the inclusion criteria. In addition, diabetic patients having a diagnosis of Type II diabetes mellitus met the inclusion criteria for our diabetic cohort. The diabetic patients were not insulin dependent. Subjects with macular edema, proliferative DR, previous vitrectomy surgery, history of other ocular disorders other than cataracts or minor refractive error, and ungradable and low-quality OCT pictures were exclusion criterions. There is no preference between left or right eyes. A board-certified retina specialist classified the patients as NoDR or different stages of NPDR according to the Early Treatment Diabetic Retinopathy Study (ETDRS) staging system. All patients underwent a complete anterior and dilated posterior segment examination. All control OCTA images were obtained from healthy volunteers that provided informed consent for OCT/OCTA imaging. Deidentified diabetic datasets were obtained for retrospective analysis. The IRB waived the need for informed consent from the patients, as patient privacy and confidentiality were maintained according to IRB guidelines. All subjects underwent OCT and OCTA imaging of both eyes (OD and OS). One en face OCTA of each eye was used for this study.
Spectral domain (SD) en face OCTA data were acquired using an AngioVue SD-OCT device (Optovue, Fremont, CA, USA). The OCT device had a 70,000 Hz A-scan rate, ~5 μm axial resolution, and ~15 μm lateral resolution for 6 mm × 6 mm scans. Only superficial OCTA images were used in this study. After image reconstruction, en face OCTA was exported from the ReVue software interface (Optovue) for further processing.
Generating ground truths
As reported in previous publication25, readers can rely on various characteristics in OCTA images to manually detect arteries and veins accurately in the 6 mm × 6 mm dataset: (1) The presence of the capillary-free zone is associated with arteries; (2) arteries do not cross other arteries and veins do not cross other veins, physiologically; (3) vessels can be traced back proximally and distally to aid in identification; (4) arteries and veins typically alternate as each vein drains capillary beds perfused by adjacent arteries. Figure 1a, b show a representative OCTA image and corresponding manually generated AV map. For generating AVA maps for the training dataset, the k-nearest neighbor (kNN) classifier is used to classify background pixels in Fig. 1b as pixels in arterial or venous areas. Since we segmented all the visible large vessels in AV maps and used kNN only to classify background pixels as pixels in arterial or venous areas, the generated AVA maps using kNN are reliable and accurate. They were reviewed and approved by an ophthalmologist. Considering Euclidean distance as distance metric and distance-weighted voting, k values between 4 and 25 generate approximately similar and smooth AVA maps. To minimize the computation cost, the k value of 5 is selected. The output of the kNN classifier is presented in Fig. 1c with a lighter tone of blue and red comparing to arteries and veins presented in Fig. 1b. The union of the arteries and veins with corresponding arterial and venous areas generate the AVA maps represented in Fig. 1d. Generating ground truth AV maps for 3 mm × 3 mm OCTA images is discussed in our previous study25. The above-mentioned procedure can be used to generate ground truth AVA maps for 3 mm × 3 mm OCTA images.
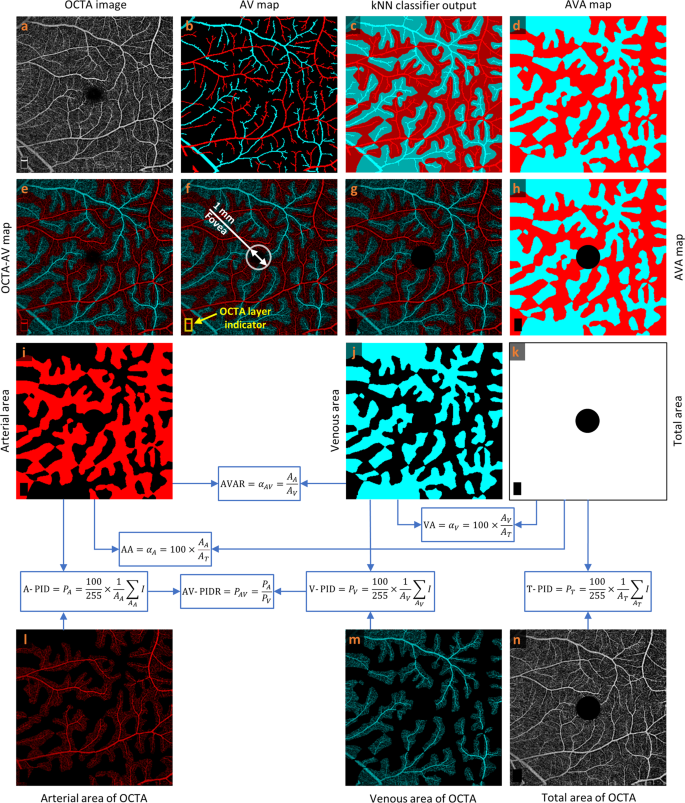

a OCTA image. b Manually generated AV map. c Output of kNN classifier. d AVA map. e OCTA-AV map. f Fovea and OCTA layer indicator in the OCTA-AV map. g OCTA-AV map excluding the fovea and OCTA layer indicator area. h AVA map excluding the fovea and OCTA layer indicator area. i Arterial area. j Venous area. k Total area shows the summation of arterial and venous areas with white color. l Arterial area of the OCTA image. m Venous area of the OCTA image. n Total area of the OCTA image excluding the fovea and OCTA layer indicator area. Calculating quantitative features is indicated by blue arrows and boxes. OCTA optical coherence tomography angiography, AV artery-vein, kNN k-nearest neighbor, AVA arterial-venous area, AVAR αAV, arterial-venous area ratio, AA αA, arterial area, VA αV, venous area, A-PID PA, arterial perfusion intensity density, V-PID PV, venous perfusion intensity density, T-PID PT, total perfusion intensity density, AV-PIDR PAV, arterial-venous perfusion intensity density ratio.
Quantitative features
By multiplying the OCTA image by the AVA map represented in Fig. 1a and d, respectively, we can have the OCTA-AV map demonstrated in Fig. 1e. To the best of our knowledge, for the first time, OCTA-AV maps are introduced in this paper as images that contain the intensity information of an OCTA image, with separate red and blue channels for arterial and venous areas. By using this method, all the vessels at different orders and scales which are visible in the OCTA images can be classified as arteries or veins. Figure 1f shows the fovea (diameter 1 mm) and OCTA layer indicator with a white circle and yellow rectangle, respectively. During image scanning of the macula, the commercial imaging device detects the fovea center automatically to keep the fovea at the center of the image. Accordingly, we can say that the center of the circle is approximately in the center of the image. Since the foveal avascular zone (FAZ) is devoid of blood vessels, the arterial and venous area segmentation in this area is artificial. As with the fovea (diameter 1 mm), the OCTA layer indicator area at the bottom left corner of the OCTA image is excluded from the OCTA-AV map as well as the AVA map presented in Fig. 1g and h, respectively.
Using the AVA maps and OCTA-AV maps, we can conduct the quantitative analysis for control, NoDR, and mild DR stages. The area of arterial or venous areas can be quantified using the AVA maps. As two novel quantitative features, the percentage of the arterial or venous areas in the total area can be defined as arterial area (AA) or venous area (VA). Therefore, AA and VA can be calculated as follows
$${\alpha }_{A}=100\times \frac{{A}_{A}}{{A}_{T}}$$
(1)
$${\alpha }_{V}=100\times \frac{{A}_{V}}{{A}_{T}}$$
(2)
where AA, AV, and AT are arterial, venous, and total area in AVA maps, respectively, and αA and αV are AA and VA, respectively. To calculate AA or VA, the number of pixels in the arterial or venous area can be divided by the number of total pixels multiplied by 100. Since the summation of arterial and venous areas are the total area, mathematically we have the following relationship between AA and VA
$$\begin{array}{c}{\alpha }_{A}=100-{\alpha }_{V}\end{array}$$
(3)
We also can define the arterial-venous area ratio (AVAR), αAV, as follows
$$\begin{array}{c}{\alpha }_{AV}=\frac{{A}_{A}}{{A}_{V}}=\frac{{\alpha }_{A}}{{\alpha }_{V}}\end{array}$$
(4)
Most commonly, the binarized OCTA images are used to calculate VAD, also known as vessel density (VD), perfusion density (PD), blood vessel density (BVD), and capillary density13,29. VAD in binarized OCTA images is the ratio of the area occupied by vessels divided by the total area converted to a percentage. In this paper, using the OCTA-AV maps that contain the OCTA intensity information, we define perfusion intensity density (PID) as a novel quantitative feature that does not require binarization with any thresholding method. The mean of the pixel intensities converted to a percentage in the total area, arterial area, and venous area can be defined as total PID (T-PID), arterial PID (A-PID), and venous PID (V-PID), respectively. So, T-PID, A-PID, and V-PID can be calculated as follows
$${P}_{T}=\frac{100}{255}\times \frac{{{{{{\rm{summation}}}}}}\,{{{{{\rm{of}}}}}}\,{{{{{\rm{intensities}}}}}}\,{{{{{\rm{in}}}}}}\,{{{{{\rm{the}}}}}}\,{{{{{\rm{total}}}}}}\,{{{{{\rm{area}}}}}}}{total\,number\,of\,pixels}=\frac{100}{255}\times \frac{1}{{A}_{T}}\mathop{\sum} _{{A}_{T}}I$$
(5)
$${P}_{A}=\frac{100}{255}\times \frac{{{{{{\rm{summation}}}}}}\,{{{{{\rm{of}}}}}}\,{{{{{\rm{intensities}}}}}}\,{{{{{\rm{in}}}}}}\,{{{{{\rm{arterial}}}}}}\,{{{{{\rm{areas}}}}}}}{total\,number\,of\,pixels\,in\,arterial\,areas}=\frac{100}{255}\times \frac{1}{{A}_{A}}\mathop{\sum} _{{A}_{A}}I$$
(6)
$${P}_{V}=\frac{100}{255}\times \frac{{{{{{\rm{summation}}}}}}\,{{{{{\rm{of}}}}}}\,{{{{{\rm{intensities}}}}}}\,{{{{{\rm{in}}}}}}\,{{{{{\rm{venous}}}}}}\,{{{{{\rm{areas}}}}}}}{total\,number\,of\,pixels\,in\,venous\,areas}=\frac{100}{255}\times \frac{1}{{A}_{V}}\mathop{\sum} _{{A}_{V}}I$$
(7)
where PA, PV, and PT are A-PID, V-PID, and T-PID, respectively, and I is intensity values in OCTA-AV maps. T-PID represents quantitative feature calculation without differentiation of the AV areas, while A-PID and V-PID represent quantitative features after AV area segmentation for differential AV analysis. The arterial-venous PID ratio (AV-PIDR), PAV, can also be defined as the division of the A-PID by V-PID as formulated bellow
$$\begin{array}{c}{P}_{AV}=\frac{{P}_{A}}{{P}_{V}}\end{array}$$
(8)
To the best of our knowledge, we defined seven new quantitative OCTA features (AA, VA, AVAR, T-PID, A-PID, V-PID, and AV-PIDR) related to AVA maps and OCTA-AV maps that can be used for quantitative analysis of diseases at different stages. The processes for calculating these quantitative features are shown in Fig. 1i–n, with blue arrows and boxes. We are going to report these quantitative features in the whole image for control, NoDR, and mild DR subjects to show their importance. These quantitative features can also be measured and reported in other regions of the OCTA images such as parafovea and perifovea, as well as their quadrants, however, that is beyond the scope of this paper.
Statistical analyses
For the statistical analysis of quantitative features, due to the limited dataset size, we treated each eye as a single observation for some subjects with images of both eyes. We performed the Shapiro–Wilk test to check if quantitative features are normally distributed. We used χ2 tests to assess the distribution of sex and hypertension among different groups. Age and quality index distributions were compared using analysis of variance (ANOVA). One-versus-one comparison of quantitative features between different groups was performed by the unpaired two-sided Student’s t test. We also applied the Bonferroni correction to compare the difference in mean values of the quantitative features. P < 0.05 was considered statistically significant.
AVA-Net architecture
For fully automated AVA segmentation in en face OCTA images, we propose the AVA-Net, a U-Net-like architecture, illustrated in Fig. 2. The input of the AVA-Net is the grayscale OCTA image. The OCTA image contains the information of blood flow strength and vessel geometry features. Since there are two classes for segmentation: arterial areas and venous areas, this is a binary segmentation problem. So, the output of AVA-Net is a single-channel grayscale image in which arterial pixels are denoted by 1 and venous pixels are denoted by 0. In this article, they are shown in blue and red colors for better demonstration. The overall design of the AVA-Net is composed of an encoder to extract features from the image and a decoder to construct the AVA map from the encoded features. The encoder includes 5 encoder blocks to reduce the image resolution by down-sampling. As shown in Fig. 2b, encoder blocks composed of two 3 × 3 convolution operations, 4 dilated convolution operations in parallel with dilation rates from 2 to 5, a concatenation operation, and a max-pooling operation with a pooling size of 2 × 2.
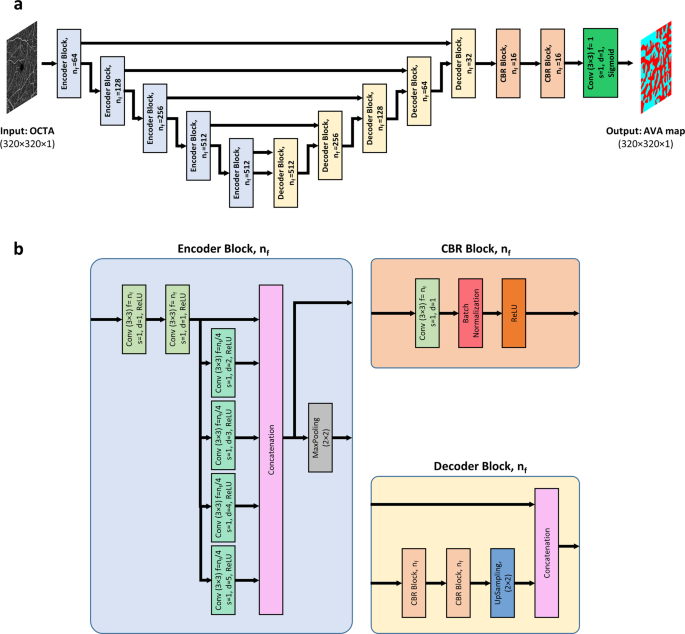
a overview of the blocks in AVA-Net architecture. b the individual blocks that comprises AVA-Net. In this figure, Conv, f, s, d, and nf stand for convolution operation, number of filters in the convolution, strides of the convolution, dilation rate of the convolution, and number of filters specified for the corresponding block, respectively. OCTA optical coherence tomography angiography, AVA arterial-venous area, CBR Conv – Batch Normalization – ReLU.
On the other hand, the decoder is composed of 5 decoder blocks, followed by 2 CBR (Conv – Batch Normalization – ReLU) blocks, and final convolutional operation with a sigmoid activation function. As shown in Fig. 2b, CBR block is composed of a 3 × 3 convolution operation, followed by a batch normalization and a ReLU activation function. The decoder blocks are composed of 2 CBR blocks, up-sampling, and concatenation operation that concatenate generated features with features coming from encoders blocks by skip connections. In all the five levels, the features prior to the max-pooling operation in the encoder blocks is transferred to the decoder blocks by skip connections. These feature maps are then concatenated with the output of the up-sampling operation in the decoder block, and the concatenated feature map is propagated to the successive layers. These skip connections allow the network to retrieve the information lost by max-pooling operations. Details of the different operations in the AVA-Net layers are presented in Fig. 2.
Loss function
In this study, the CNN was trained using IoU loss30 or Jaccard loss to directly optimize the IoU score, the most commonly used evaluation metric in segmentation31. For multiclass segmentation, it is defined by
$$\begin{array}{c}{L}_{IoU}=1-\frac{{\sum }_{c=1}^{C}{\sum }_{i=1}^{N}{g}_{i}^{c}{s}_{i}^{c}}{{\sum }_{c=1}^{C}{\sum }_{i=1}^{N}({g}_{i}^{c}+{s}_{i}^{c}-{g}_{i}^{c}{s}_{i}^{c})}\end{array}$$
(9)
where \({g}_{i}^{c}\) is the ground truth binary indicator of class label c of voxel i, and \({s}_{i}^{c}\) is the corresponding predicted segmentation probability. N is the number of voxels in the image and C is the number of classes. Since we have two classes, this is a binary segmentation problem. So, we have
$$\begin{array}{c}{L}_{IoU}=1-\frac{{\sum }_{i=1}^{N}{g}_{i}{s}_{i}}{{\sum }_{i=1}^{N}({g}_{i}+{s}_{i}-{g}_{i}{s}_{i})}\end{array}$$
(10)
Training process
AVA-Net architecture was trained using the Adam optimizer with a learning rate of 0.0001 (β1 = 0.9, \({\beta }_{2}=0.999\), \({\epsilon }={10}^{-7}\)) to have a smooth learning curve for the validation dataset. With the IoU loss function, mini-batch sizes of 28 were utilized for the training. The training process takes up to 5000 epochs when the model performance stops improving on the validation dataset. One epoch is defined as the iteration over 3 training batches. To avoid overfitting, data augmentation methods are applied on the fly during training, including random flipping along horizontal and vertical axes, random zooming, random rotation, random image shifting, random shearing, random brightness shifting. As retinal vessels in OCTA images have diverse tree-like patterns and differing vessel diameters, and because images can be taken from different locations of right and left eyes with diverse quality, the above-mentioned data augmentation methods could help improve the generalization capability of the CNN for segmenting unseen images. Since our data is limited, the 5-fold cross-validation procedure is used for CNN evaluation after shuffling the data based on the patient’s eye. Therefore, in each fold, the network was trained with 80 percent of the images, and evaluation was performed on the other 20 percent of the images.
The IoU and Dice coefficient metrics, also known as the Jaccard Index and F1 Score, respectively, are mostly used in image segmentation. Therefore, in addition to segmentation accuracy, IoU and Dice coefficient were used as the evaluation metrics for evaluating the AVA segmentation, by comparing the predicted AVA maps with manually labeled ground truths.
The training was performed on a Windows 10 computer using NVIDIA Quadro RTX 6000 Graphics Processing Units (GPU). The CNN was trained and evaluated on Python (v3.9.6) using Keras (2.6.0) with Tensorflow (v2.6.0) backend. Training every fold of AVA-Net took about 10 h with the mentioned resources.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.



