Dataset
One aim of this work was to curate a large, high-quality dataset of molecular tastants from publicly available data. Every molecule was assigned a taste label: sweet, bitter, sour, umami, or undefined. The category of “undefined” contains molecules, not clearly assignable to one of the other categories, explicitly including compounds labeled as salty or tasteless. The dataset utilizes a standard text-based representation of molecules, called SMILES20. Every molecule is labeled with a taste: sweet, bitter, sour, umami, and undefined. Where the last category encompasses compounds that had either previously been established as tasteless or compounds that were present in the source databases but for which no clear taste could be associated, particularly compounds with odor rather than taste labels. Salty was excluded as a taste category because only a very small number of molecules actually produce this taste apart from sodium and chloride ions21. Data curation was handled with the cheminformatics package RDKit38.
The FART dataset encompasses small molecules with an average molecular weight of 374 Da ±228, see Fig. 3c. To visualize the distribution of the taste labels over the chemical space covered by the dataset, a t-SNE plot (Fig. 3a) was generated from Morgan fingerprints27. Figure 3b highlights the strong imbalance of the taste classes of the dataset.
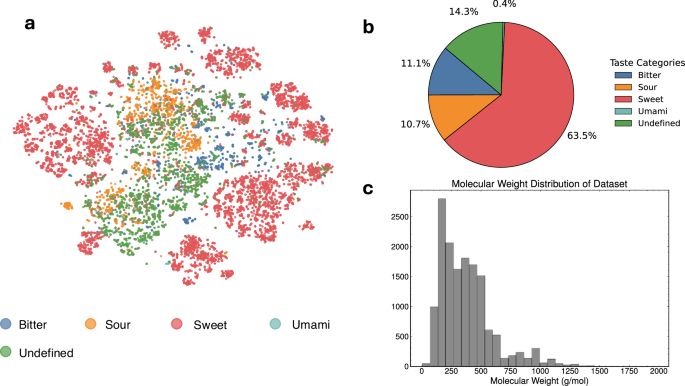
a t-SNE plot of the chemical space covered by the dataset. b Distribution across taste classes for the dataset, showing a strong imbalance for umami. c Molecular weight distribution for the dataset (mean 374 g/mol, std. deviation 228 g/mol).
The FART dataset aggregates experimental data from six sources39,40,41,42,43, see also Supplementary Table 1 in the SI. ChemTasteDB is one of the largest public databases of tastants and contains 2944 organic and inorganic tastants from which 2177 were used to train FART. The database was curated from literature39. Biological macromolecules were generally excluded, notably longer peptides, which we do not consider in our approach. Previous work has explored this chemical space particularly in the context of umami and bitter prediction44,45,46.
FlavorDB aggregates data on both gustatory and olfactory sensation from a number of sources40. The FART database uses the “flavor profile” given by FlavorDB as most molecules do not have a specific entry for taste. Data from FlavorDB will thus be more heterogeneous given that some of these flavor profiles will actually be based on smell, not taste. Care was taken to only include compounds with unambiguous taste adjectives in the dataset. From the 25,595 total molecules, 10,372 could be clearly attributed to one of the four taste categories. FlavorDB is dominated by sweet molecules and is also the source of the data imbalance in the final dataset. PlantMolecularTasteDB contains 1,527 phytochemicals with associated taste of which 906 were used for this dataset41. The database is based on both literature and other databases, some of which overlap with other sources used for FART. To obtain more data on bitter compounds, a database of ligands that bind to the human bitter receptor (TAS2) was also considered which yielded 53 previously unseen bitter compounds42. Combined, these datasets represent the largest publicly available dataset of unique, labeled molecule-taste pairs. However, there remains significant heterogeneity in how these datasets reference results, how experimental data was collected, and certain biases in terms of cultural or genetic preferences may persist. Thorough curation is thus needed to guarantee standardized data and to reduce noise as much as possible.
Water-soluble, acidic molecules (pKA between 2 and 7), assumed to taste sour47, were collected from an ongoing project based with the International Union of Pure and Applied Chemistry (IUPAC) digitizing three high-quality sources of pKA values in the literature48,49,50. Sour taste is influenced by other factors such as cell permeability, which is the reason why organic acids taste more acidic than inorganic acids such as HCl at the same pH. Nonetheless, acidic molecules can be assumed to also taste sour47. A total of 1,513 acids could be obtained in this way although it should be noted that sour taste, as all tastes, is concentration-dependent and that some of the weaker acids may not be picked up by humans. The pKA values refer to the most acidic proton and are all measured between 15 and 30 ∘C in water, i.e. around physiological temperature, excluding any acids that are not water-soluble. Lastly, 19 umami-tasting molecules were collected from the literature43 of which 11 were not found in any other database.
The combined dataset was reduced to the taste label associated with a canonicalized SMILES representation. The open-source cheminformatics package RDKit38 was used to further curate the dataset. First, all SMILES that did not allow the generation of a valid molecular graph were excluded. To avoid solvent-containing molecules, all entries with multiple uncharged fragments were removed. Charged molecules were additionally excluded to prevent substances with missing counter ions. Labels were visually inspected and manually corrected for around 100 molecules. All SMILES were standardized with the default RDKit standardization procedure including canonicalization. Duplicates could be removed with the help of these canonicalized SMILES, see Supplementary Fig. 1 in the SI for a summary.
While only very few entries with invalid SMILES (21) or charged molecules (342) needed to be removed, the number of entries containing multiple neutral fragments (3783) was more significant. These are typically SMILES that contain the tastant as well as a solvent which needs to be removed. Filtering for molecules below 2000 Dalton in molecular weight (1) was introduced to avoid SMILES strings that are longer than what the context window of ChemBERTa allows. The dataset was further inspected by eye and some mislabeled entries were manually corrected. The duplicate removal (14,685) reduced the dataset by almost half to a final size of 15,025 entries, see Supplementary Fig. 1. The large number of duplicates underlines the significant overlap among the databases used. When duplicate entries existed from different sources, which source would be given in the final dataset was arbitrarily determined based on the index. The final dataset exhibits a strong data imbalance, where sweet represents over 60% and umami less than 1% of the data.
The curated dataset was further enriched by general information (PubChemID, IUPAC name, molecular formula, molecular weight, InChI, InChIKey), accessed through the PubChem API51. The dataset, FartDB, was published in agreement with the FAIR principles14 and can be accessed through several different interfaces to encourage its use by other research projects.
Visualization
The t-SNE plot, see Fig. 3a, (perplexity = 30) was generated using 1024-Morgan fingerprints (radius = 2) based on PCA initialization and the Jaccard distance metric. Heatmap plots for interpretability were generated using custom code utilizing the SimilarityMaps functionality of RDKit, see Fig. 2.
Tree-based classifiers: XGBoost, random forest
Tree-based ensemble models such as random forest (RF) or XGBoost26 are often considered strong baseline models in classification tasks for more complex model architectures, such as transformers, given their robust performance, efficient training, and low model complexity. A RF model is an ensemble learning method that trains multiple, in this case 150, decision trees during the training process, combining their predictions to improve accuracy and mitigate overfitting on the training data. A XGBoost model is an ensemble learning method that builds multiple decision trees sequentially, optimizing each tree to correct errors from the previous ones, thereby improving accuracy and mitigating overfitting through regularization techniques.
In this work, three different tree-based classifiers were trained including hyperparameter optimization. The two XGBoost models were trained either using 1024-Morgan (radius = 2) fingerprints or these same fingerprints concatenated with 15 descriptors calculated using Mordred52. These predictors had been previously found to be particularly correlated with taste28. The models were evaluated with a multi-class logarithmic loss function, results are given in Table 1.
FART models
For our transformer-based model, we utilized ChemBERTa18,19, which has been pre-trained on 77 million SMILES strings using a masked language modeling approach. Here, a percentage of the input is randomly masked, and the model is trained to predict the masked parts of the SMILES string. This allows the model to learn contextualized representations of chemical structures in a self-supervised manner, capturing both local and global molecular features. We fine-tuned ChemBERTa on our taste prediction dataset using a categorical cross-entropy loss function with a learning rate of 10−5 and a batch size of 16. We experimented with three different training configurations: one model was trained for 20 epochs on the original dataset and a second model was trained for 2 epochs on a 10-fold augmented dataset. The performance of all models is summarized in Table 1. The Chemprop model were trained as described in the original publication using default hyperparameter optimization25.
To evaluate multi-class classification performance, macro and weighted averages are commonly used to summarize metrics across all classes. The macro (unweighted) average is computed with
$${\rm{Macro}}=\frac{1}{C}\mathop{\sum }\limits_{i=1}^{C}{M}_{i},$$
(1)
where C is the number of classes and Mi is the metric (e.g., precision, recall, F1-score) for the ith class. The weighted average is given by
$${\rm{Weighted}}=\mathop{\sum }\limits_{i=1}^{C}\frac{{n}_{i}}{N}\cdot {M}_{i},$$
(2)
where ni is the number of instances in class i, N is the total number of instances across all classes, and Mi is the metric for the ith class.
All transformer models were trained on multiple NVIDIA T4 GPUs in Google Cloud using the HuggingFace Transformers library53. For all experiments, the ChemBERTa checkpoint seyonec/SMILES_tokenized_PubChem_shard00_160k on HuggingFace was used, consisting of 6 layers and a total of 83.5 million parameters. Training on the unaugmented dataset was run for 20 epochs, while training on the augmented dataset was run for 2 epochs. A weight decay of 0.01 was applied, and a batch size of 16 was used. Both parameters follow standard values for fine-tuning and have not been optimized for this problem. Standard values for fine-tuning were also used for all other necessary parameters. Training was continued until overfitting was observed, as indicated by the loss function on the evaluation dataset, or until the loss had saturated. At this point, the best model checkpoint, corresponding to the lowest evaluation loss, was selected for further analysis.
Confidence metric
Similar to linguistic synonymy, where multiple words share the same meaning, distinct SMILES representations can map to the same underlying molecular structure. To leverage this property, we generated an ensemble of 10 synonymous SMILES for each molecule in our dataset. The exact number of SMILES augmentation has been arbitrarily set and could alternatively be treated as an optimizable hyperparameter. The exact number of augmentations is arbitrary and can be defined by the user while ten is a reasonable number to expect even for smaller molecules. We then performed inference on the entire ensemble, obtaining individual predictions for each SMILES variant.
To aggregate these results, we employed a voting procedure across the ensemble’s predictions. Using the strictest threshold, where all 10 predictions had to agree for a label to be assigned by the model, we still retained predictions for 94% of the dataset while boosting the model’s accuracy to above 91%. Both the transient augmentation and subsequent prediction on these SMILES adds to the computational time. The confidence metric suggested here is straightforward to implement and provides a robust indication of prediction reliability in addition to the interpretability framework.
Interpretability framework
Integrated gradients23 is a method for attributing a deep neural network’s prediction to its input features. The core idea is to integrate the gradients of the output taken along a linear path from a baseline input to the input at hand, see Eq. (3). Mathematically, for a neural network F(x), an input x and baseline input \({x}^{{\prime} }\) (e.g. the zero input), the attribution for the ith feature is:
$${{\rm{IntegratedGrads}}}_{i}(x)=\left({x}_{i}-{x}_{i}^{{\prime} }\right)\times \mathop{\int}\nolimits_{0}^{1}\frac{\partial F\left({x}^{{\prime} }+\alpha \times \left(x-{x}^{{\prime} }\right)\right)}{\partial {x}_{i}}\,{\rm{d}}\alpha .$$
(3)
The method satisfies important axioms like sensitivity (if inputs differ in one feature but have different predictions, that feature should receive attribution) and implementation invariance (attributions are identical for functionally equivalent networks). The method is readily available for Hugging Face Transformer models through the transformers-interpret package54.




