dataset
For pre-training, we selected 189 compounds from 10 MoA classes (ATPase, Aurora Kinase, HDAC, JAK, HSP, PARP, protein synthesis, topoisomerase, tubulin polymerization inhibitor, and retinoid receptor agonist, see distribution in Table SM1) at 10uM, similar to Harrison et al.52 They were randomly distributed into eight 384-well plates seeded with human osteosarcoma U2OS cells. Each plate contained all compounds, with a median of 2 wells (technical replicates) per compound per plate. Images were acquired at five time points (4, 8, 12, 16, and 20 hours post-dose) in four fields of view (FOV) and three Z-planes (3.6u step size) per field of each well using an Opera Phenix microscope (20x air, 2160 × 2160 pixels). Define. replica As one complete experimental run (seeding, compound administration, and imaging of all plates at all time points) duplicate Each well receives the same treatment. To account for variation in biological replicates, seven replicas of the pre-training set were collected, resulting in a total of 430,080 raw images. The size of the pre-training dataset after pre-processing was 5.7 million input images of 224 × 224 × 3 pixels (see Supplementary Material for details). For multiple dose evaluation, the same protocol was used to acquire one replica at each of the three doses (0.156, 0.625, 2.5 μM) and two replicas at the 10 μM dose, resulting in a total of 307,200 raw images. In the holdout set, 89 additional compounds belonging to the same class were distributed across four 384-well plates, of which 81 data passed quality control (122,880 raw images). The supplementary material contains details regarding cell preparation, data collection, data preprocessing, and data volume.
Training a self-supervised model
we chose DINO18 With a small backbone of ViT53 It is used as the main algorithm based on three main advantages: Effectiveness in spatial clustering embedding.18the capabilities of zero-shot learning, and its successful application in fluorescent cell imaging23, 24, 25, 38. Our implementation builds on recent advances in addressing the unique challenges of microscopy, incorporating batch-to-batch loss and sampling strategies proposed by Haslum et al.38 Mitigates batch effects and Barlow Twins loss functions present in high-throughput fluorescence microscopy data.22,38combines self-supervised methods of sample contrast and feature contrast.54the robustness of feature learning is enhanced.
To evaluate the contribution of each loss component, we conducted an ablation study across four model variants trained for 60 epochs with a standard DINO training schedule.18 If the starting learning rate is 0.001 on the same pre-training dataset:
-
Standard DINO loss (DINO)
-
DINO + Barlow Ross (DINO + Barlow)
-
DINO + cross-batch loss (DINO + XB)
-
DINO + Barlow + Cross-Batch Loss (DINO + Barlow + XB)
CellProfiler baseline
Additionally, we performed an experiment to create a baseline from CellProfiler.43 Features extracted at the single cell level. Single cell segmentation was extracted using the latest CellPose-SAM.14 model. The Cell Profiler pipeline shares the parameters used in the code repository that accompanies the paper. We extracted 284 features at the single-cell level and aggregated the median values for each image. The following steps (normalization, phenotypic activity, MoA classification) were identical for all self-supervised models and the CellProfiler baseline.
Aspect-agnostic vision transformers as a data augmentation strategy
While being a standard DINO,18 We observed that brightfield Z-stacks naturally provide varying focal sharpness due to the use of Gaussian blur for feature stability. In this study, we used three Z planes (-3.6, 0, and +3.6 μm) covering the depth of field range for training. A similar approach can be used with other objective lens configurations. We perform ablation studies to confirm that variation in focus, and not deblurring alone, causes changes in performance (see Supplementary Material, Table SM2).
Evaluate the expansion strategy using the best performing loss function configuration.
Plane independent DINO + Barlow + cross-batch loss
(PA + Dino + Barlow + XB)
(See Figure 1 for model architecture).
Loss function ratio and Barlow Twins uncorrelated hyperparameters
When training with Barlow twin loss, we experimentally found that strong decorrelation helps training, so the coefficient of the off-diagonal Barlow twin loss element was set to 0.5. In the total loss function, the loss ratio for DINO: Barlow Twins was 1:1. In cross-batch training, two global crops were sampled from two different experimental replicas of the same compound at the same dose and time point. Three local crops were sampled for each image, which served as a source of global crops. For airplane-independent training, airplanes were randomly selected for each crop.
Downstream nuclear detection
Recent work55,56 We demonstrated that features learned by a self-supervised model can be beneficial for completely unsupervised object discovery tasks. A study by Simeoni et al.57 We demonstrated the possibility of segmenting and detecting salient objects from the background using a relatively simple unsupervised algorithm based on pre-trained attention maps and features and a computer vision segmentation smoothing algorithm (bilateral solver).58). We tested whether this algorithm can perform fully unsupervised live nuclear detection from bright-field images using the patch-level features of our best-performing self-supervised training model and validated the algorithm against Hoechst staining (see Results section for training and validation results and Supplementary Materials for details on biological experiments).
Evaluation of self-supervised models
Self-supervised learning can provide a universal feature extraction function for determining cell states for various tasks. We validated this with live-cell bright-field microscopy using a five-step evaluation pipeline for each dose and time point combination.
-
1.
Feature extraction: Generation of embeddings from evaluation and holdouts set at each time point and dose.
-
2.
Normalization: Application of embedded normalization techniques.
-
3.
Phenotypic activity assessment: Evaluating feature space properties using mean average precision (mAP)39 and mAP-ES) metrics to estimate the activity of each compound.
-
4.
Benchmark phenotypic activity against nuclear number as a traditional assay: segment and count the nuclei on each image and compare the activity detected by nuclear number to the activity based on mAP and mAP-ES metrics.
-
5.
Validation of downstream tasks: Evaluation of learned representations with MoA classification using linear probes on extracted features.
The MoA classification for each dose and time point was then aggregated stepwise for MoA prediction (see “MoA classification” section).
Embedded normalization approach
Batch effect normalization is well studied in fluorescence imaging;59,60,61its effectiveness for live bright-field embedding remains to be elucidated. Based on the paper by Arevalo et al.59 For our analysis, we selected the best performing methods from three different families under comparable settings (single microscope, comparable amounts of compounds, multiple replicas).
-
Fast MNN62nearest neighbor based method
-
harmony63a mixed model-based method
-
Baseline, Arevalo et al. baseline method (using median absolute deviation per plate normalization)59.
Furthermore, a new approach (“MAD+Harmony/FOV”) was proposed and evaluated as described below.
New batch modification approach: MAD+harmony/field of vision
After the features were extracted, they were concatenated within a pandas dataframe with all features and corresponding metadata per tile. As a first step, each measurement (plate imaged at a specific time point) was normalized individually to the DMSO well using the MAD + Robustize method, then all normalized embeddings were concatenated and normalized using Harmony grouped only at the level of experimental replica and field of view. The pseudocode for the algorithm is provided in Algorithm SM1.
Evaluation of phenotypic activity
To quantitatively assess the quality of embedding across different models and normalization techniques, we employed the mean mean precision (mAP), a common measure of a compound’s phenotypic activity.39. In the new approach, enabled by multiple replicates of compounds, the mAP of each compound is treated as an experimental distribution rather than a single value, and a null distribution is constructed based on the DMSO sample. Phenotypic activity is then defined as the strength of the difference between the mAP per compound and the null distribution as measured by Cohen’s d effect size.40 (mAP-ES), a compound is defined as active if mAP-ES ≥ 0.8 (corresponding to a large effect size in the statistical literature).40).
Mean Average Precision Effect Size (mAP-ES)
map39 Quantify how clearly a compound’s phenotypic response can be distinguished from a negative control in the embedding space. To calculate this, we need to define, for each query well containing the perturbation, its positive and negative pair.39. For compound treatments, we defined positive counterparts as three wells with the same treatment, but taken from different randomized plates in different experimental replicas (probably the strongest batch effect difference). A negative pair was defined as 31 DMSO wells randomly sampled from the same plate as the query well (probably minimal batch effect difference).
Similarly, a null distribution of mAPs in DMSO wells was generated. A query DMSO well was randomly selected, and three DMSO wells from different plates and experimental replicas were defined as a positive pair. 31 DMSO wells from the same plate were selected as negative pairs. A distribution of mAPs for each compound and a null distribution of mAPs for the negative control were created by randomly sampling the query wells, positive, and corresponding negative pairs 1000 times.
Phenotypic activity was defined as the strength of the difference between mAP per compound and the mAP null distribution as measured by Cohen’s d effect size.40 (mAP-ES). This approach treats the mAP per compound as a random value, and the null distribution of mAP in DMSO wells by design accounts for batch effects present in the dataset. An example of mAP distribution is shown in Figure 6.
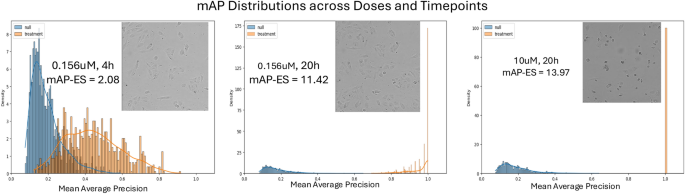
Example of mAP distribution of null and treatment (compound TAK-901) at three doses and time points with different activity levels.
Benchmarking phenotypic activity against nuclear number
Cell number is a recently proposed baseline to assess the added value of phenotypic profiling. This is because many activity benchmarks are strongly correlated with viability and can be predicted from a single cell count characteristic.41. A nuclear detection algorithm was used to extract the number of live cells per field of view for each image. Normalized cell counts were calculated as the Z-transform of cell counts based on the statistics of the DMSO wells of each plate at each time point, and for each perturbation, the 95% confidence interval of the median of all replicates (without a priori aggregation) was calculated using the bootstrap method. A compound was considered active if the upper bound of the confidence interval was less than zero (for DMSO wells, meaning normalized cell number equal to zero). The number of compounds considered active based on cell number was then compared to the number of active compounds based on mAP-ES.
MoA classification
Representation quality was further assessed by MoA classification using linear probes.33a standard approach for self-supervised learning that tests whether an embedding contains linearly separable class information. Five-fold cross-validation was used for the pre-training compound set, and the classifier was trained on the active compounds from the pre-training set for the holdout set.
To address potential polypharmacology49 To account for different dose responses, we trained separate MoA classifiers for active compounds for each condition at each dose time point and generated the final MoA prediction by taking a weighted average of the softmax outputs. The contribution of dose was assessed by incrementally adding dose using a classifier trained on activity-filtered data for each condition.



