Identification of key EET regulators and their prognostic significance in endometrial cancer
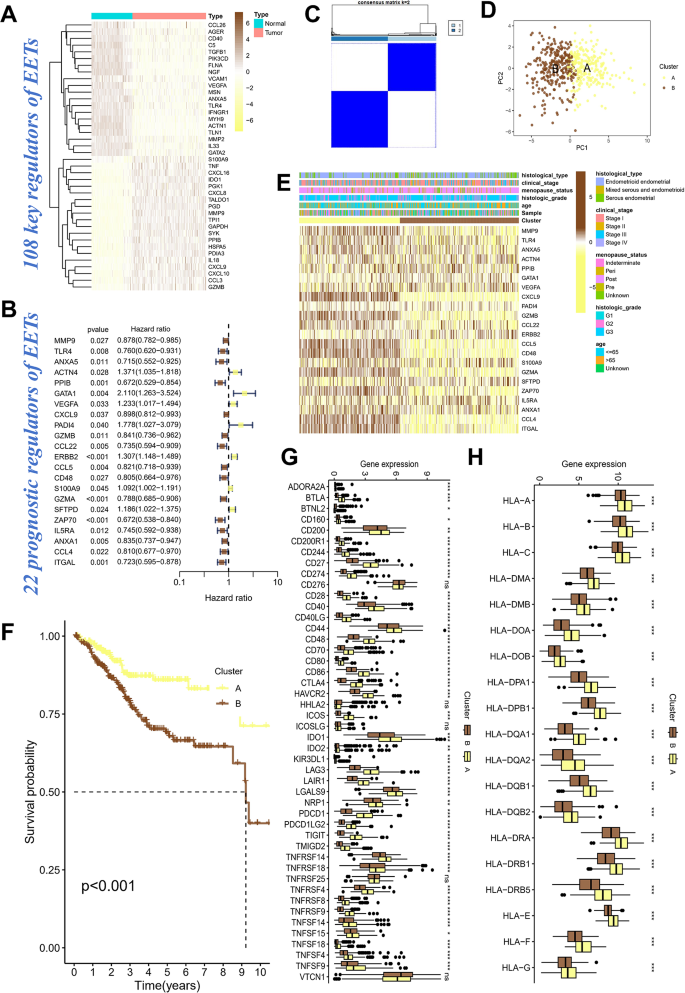
Using differential analysis, we identified a set of 108 key eosinophil extracellular trap (EET) regulators, whose expression was significantly altered in tumor tissues compared to normal tissues (Fig. 1A). The heatmap clearly demonstrates distinct expression patterns of these genes, with many showing upregulation in tumor tissues, such as CCL26, AGER, and CD40. The prognostic significance of 22 EET-related genes was further explored through a forest plot, which assessed hazard ratios (HR) and p-values in relation to patient survival (Fig. 1B). Several genes, including MMP9, TLR4, and VEGFA, were found to be significantly associated with poor prognosis (p < 0.05), with hazard ratios greater than 1, indicating their involvement in tumor progression. Based on unsupervised clustering analysis (Fig. 1C), the matrix revealed strong correlations among genes within the tumor samples, suggesting shared regulatory pathways or interactions, which led to the identification of two distinct subgroups. Principal component analysis (PCA), shown in Fig. 1D, further confirmed the existence of these distinct tumor subtypes. The separation of these clusters suggests the presence of different molecular subtypes of endometrial cancer, which may be associated with distinct clinical characteristics and prognostic outcomes. The relationship between these clusters and clinical factors such as histological type and clinical stage was further explored in Fig. 1E, with Cluster A predominantly showing upregulation of EET regulators. Kaplan-Meier survival analysis revealed significant differences in survival outcomes between Cluster A and Cluster B, with Cluster A demonstrating poorer survival (p < 0.001) (Fig. 1F). This finding indicates that the gene expression profile associated with Cluster A is predictive of worse prognosis, reinforcing the clinical relevance of these results. Gene expression differences between tumor clusters A and E were further examined through boxplots (Fig. 1G and H). Notably, immune-related genes such as HLA-A, HLA-B, and HLA-C exhibited significant differential expression across clusters, with elevated expression in the more aggressive Cluster A. The upregulation of these immune-related genes in Cluster A may reflect an adaptive response of the tumor to evade immune surveillance.
Identification and prognostic implications of Eosinophil Extracellular Trap (EET) regulators in endometrial cancer. Panel (A) Heatmap showing the differential expression of 108 EET regulators in tumor vs. normal tissues. Brown represents high expression, while light yellow indicates low expression. Panel (B) Forest plot illustrating the prognostic significance of 22 EET-related genes. Panel (C) Matrix of EET regulators, showing strong correlations within tumor samples and the identification of two distinct tumor subgroups. Panel (D) Principal component analysis (PCA) of tumor samples, confirming two distinct clusters based on gene expression profiles. Panel (E) Heatmap showing the association of tumor subtypes with clinical factors, with Cluster A showing predominant upregulation of EET regulators. Panel (F) Kaplan-Meier survival analysis showing poorer survival in Cluster A compared to Cluster B. Panel G and H. Boxplots of gene expression differences between clusters A and E for immune-related genes, with elevated expression in Cluster A
Immune infiltration profiles and tumor microenvironment characteristics
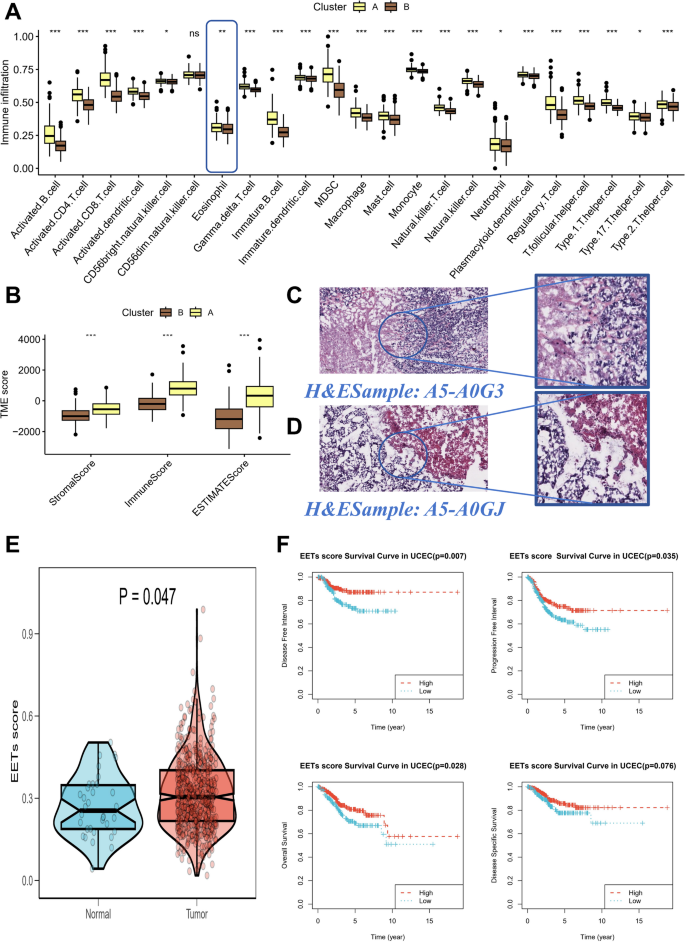
Using differential analysis, we assessed immune infiltration profiles in tumor clusters A and B (Fig. 2A). Significant differences in the infiltration of various immune cell types were observed. Notably, eosinophil infiltration was significantly higher in Cluster A (p < 0.05), suggesting a pivotal role for eosinophils in this tumor subtype. Other immune cell types, including activated CD4 + T cells, activated CD8 + T cells, and natural killer cells, also exhibited significant differences in their infiltration levels between the clusters, highlighting distinct immune microenvironments across tumor subtypes. Tumor microenvironment (TME) characteristics were further evaluated using stromal, immune, and ESTIMATE scores (Fig. 2B). Cluster A exhibited significantly higher immune and ESTIMATE scores compared to Cluster B, indicating a more immune-enriched environment in Cluster A. Figure 2C-D presents representative H&E-stained images of Cluster A samples, showing dense immune cell infiltration, particularly eosinophilic staining cells. The histological images further support the immune infiltration findings from Fig. 2A, revealing substantial immune cell aggregation in Cluster A. In Fig. 2E, EET scores were significantly higher in tumor tissues compared to normal tissues (p = 0.047). This suggests that EET activation is a prominent feature in endometrial cancer, particularly in tumor tissues, where elevated EET scores may contribute to disease progression. Kaplan-Meier survival analysis (Fig. 2F) demonstrated that higher EET scores were associated with significantly poorer clinical outcomes. Patients with high EET scores had reduced disease-free survival (p = 0.007), progression-free survival (p = 0.035), and overall survival (p = 0.028), underscoring the prognostic value of EET activation in endometrial cancer.
Immune infiltration and tumor microenvironment in endometrial cancer.Panel (A) Immune infiltration profiles in tumor clusters A and (B) Panel B. TME scores in tumor clusters A and B. Panels (C) Representative H&E-stained images of Cluster A samples, showing dense immune cell infiltration, especially eosinophils. Panels (D) Representative H&E-stained images of Cluster B samples. Panel (E) EET scores in tumor vs. normal tissues. Tumor tissues showed significantly higher EET scores (p = 0.047). Panel (F) Kaplan-Meier survival analysis of EET scores. High EET scores were associated with worse disease-free survival, progression-free survival, and overall survival
Molecular pathway activation and functional enrichment in tumor subtypes
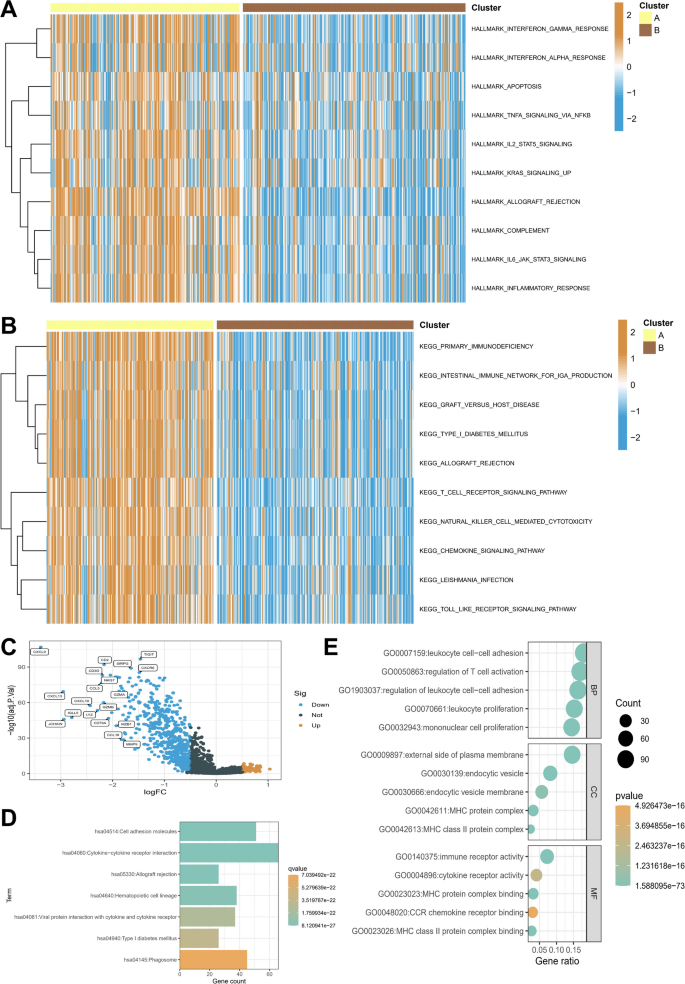
We further investigated the molecular pathways in tumor clusters A and B. Figure 3A presents a heatmap of hallmark gene sets, revealing distinct patterns of gene expression between the two clusters. Cluster A exhibited higher expression of genes associated with immune responses, such as interferon gamma response and complement, suggesting a more active immune-related signature. In Fig. 3B, KEGG pathway analysis highlighted differential activation of various immune-related pathways between the clusters. Cluster A showed significantly higher expression of pathways associated with primary immunodeficiency, immune network for IgA production, and T cell receptor signaling, indicating distinct immune and inflammatory responses in the two subtypes. Figure 3C displays the results of differential gene expression analysis of 975 genes, with the volcano plot highlighting significant upregulation of immune-related genes in Cluster A (Supplementary File 1). Notably, genes involved in leukocyte cell adhesion and T cell activation were significantly upregulated in Cluster A, supporting the higher immune infiltration observed in this group. Based on the upregulated differentially expressed genes in Cluster A, Fig. 3D shows that functional enrichment analysis revealed significant enrichment of genes related to immune receptor activity and cytokine receptor activity in Cluster A, further corroborating the immune-enriched profile of this subtype. Figure 3E summarizes the Gene Ontology (GO) terms, with leukocyte cell-cell adhesion and T cell activation identified as key biological processes in Cluster A.
Molecular pathways and functional enrichment in tumor clusters A and B. Panel (A) Heatmap of hallmark gene sets, showing higher immune-related gene expression in Cluster (A) Brown represents high pathway enrichment scores, while blue indicates low pathway enrichment scores. Panel (B) KEGG pathway analysis highlighting significantly higher expression of immune-related pathways. Brown represents high pathway enrichment scores, while blue indicates low pathway enrichment scores. Panel (C) Volcano plot of differential gene expression analysis. Panel (D) Functional enrichment analysis of upregulated genes in Cluster A. Panel (E) GO term summary
Machine learning-based model evaluation and prognostic biomarker discovery
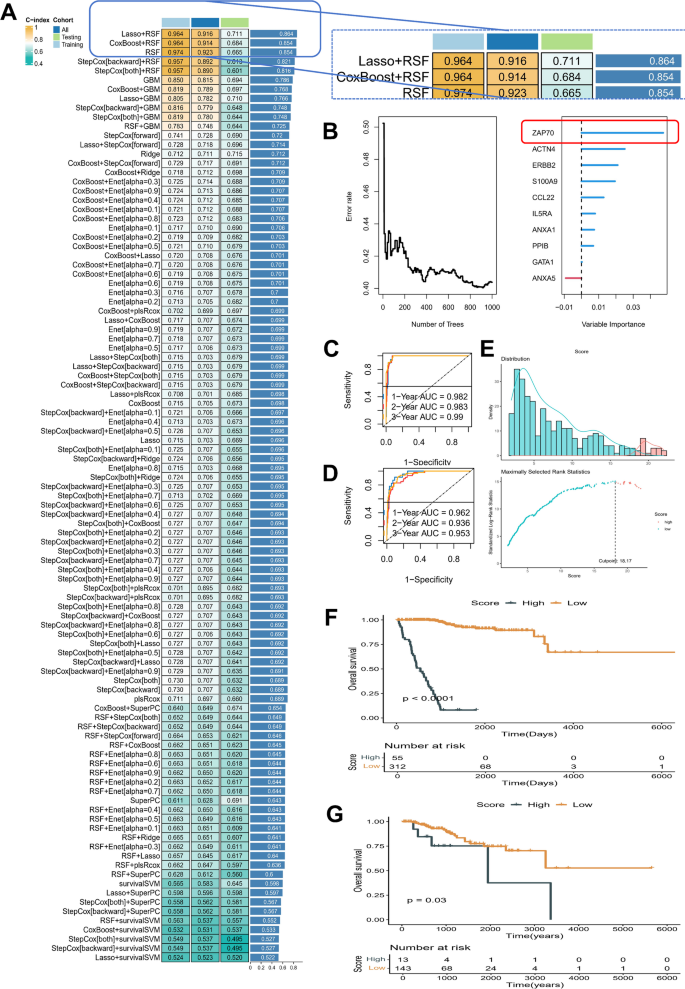
Based on eosinophil extracellular trap (EET) regulators with prognostic value, we evaluated the performance of 101 combinations of machine learning algorithms to identify the optimal model for endometrial cancer prediction. Figure 4A presents a heatmap of performance metrics (C-index) for the different model combinations. The Lasso + RSF models achieved the highest C-index values of 0.854, indicating superior predictive performance. The detailed C-index values of each model can be found in Supplementary Files 1. Figure 4B shows the variable importance ranking, with S100A9 emerging as the most important feature for predicting patient outcomes, followed by ACTN4, ERBB2, and ZAP70. In Fig. 4C-D, the performance of the Lasso + RSF model is evaluated using receiver operating characteristic (ROC) curves for 1-year, 2-year, and 3-year survival predictions. The AUC values for these time points were all greater than 0.9 in both the test and training cohorts. Figure 4E shows the distribution of the survival score, where a higher score correlates with worse prognosis. The optimal cutoff for the risk score was 18.17, which divided patients into high-risk and low-risk groups. The model’s ability to discriminate between these two groups is clearly reflected in the survival curves. Figure 4F also presents Kaplan-Meier survival curves, with high-risk and low-risk categories significantly associated with overall survival. The high-risk group exhibited significantly worse survival outcomes (p < 0.0001), further validating the clinical utility of the model. Finally, Fig. 4G shows the survival curves for the Lasso + RSF model applied to a validation cohort, confirming its predictive value over a longer follow-up period. The model’s performance remained significant, with a p-value of 0.03, demonstrating its reliability across different datasets.
Evaluation and Validation of the Lasso + RSF Model. Panel (A) Heatmap of C-index values for 101 machine learning algorithm combinations. Panel (B) Variable importance ranking, with S100A9 identified as the most important feature for predicting patient outcomes. Panels (C-D) ROC curves for 1-year, 2-year, and 3-year survival predictions using the Lasso + RSF model. Panel (E) Distribution of survival scores, with an optimal cutoff of 18.17 to classify patients into high-risk and low-risk groups. S Panel (F) Kaplan-Meier survival curves for high-risk and low-risk groups in training cohort. Panel (G) Kaplan-Meier survival curves for the Lasso + RSF model applied to a validation cohort
Clinicopathological feature correlations with risk scores and mutational analysis
We assessed the association between the risk score and various clinicopathological features. Figure 5A shows the distribution of risk scores across different age groups, with patients aged ≤ 65 years exhibiting significantly lower risk scores compared to those aged > 65 years (p < 0.001). Similarly, Fig. 5B demonstrates that advanced disease stages (III and IV) are associated with higher risk scores, whereas early-stage disease (I and II) is linked to lower risk scores (p < 0.05). Figure 5C further highlights that patients with higher tumor grades (G3) exhibit significantly higher risk scores compared to those with lower grades (G1 and G2) (p < 0.0001). Figure 5D presents the risk scores across different histological types, with serous tumors showing significantly higher risk scores compared to endometrioid and mixed tumors (p < 0.01). Next, we evaluated the correlation between immune cell infiltration and risk scores. Figure 5E-F shows a correlation matrix, indicating that immune cell types such as myeloid dendritic cells, T cell CD4+, and macrophages exhibit significant correlations with the risk score across multiple methods (XCELL, TIMER, QUANTISEQ, EPIC, and CIBERSORT). Figure 5G illustrates somatic mutation data in the high-risk group, with alterations in PTEN, PK3CA, and TP53 being most common (p < 0.0001). These mutations were significantly associated with higher risk scores, with PTEN and PK3CA mutations being particularly prevalent in the high-risk cohort (32% and 29%, respectively). Finally, Fig. 5H provides a detailed overview of somatic mutations, showing that the low-risk group exhibited significantly lower mutation rates in these genes (69%, 53%, and 54%, respectively), further reinforcing the association between genetic alterations and risk stratification.

Association of risk scores with clinicopathological features and immune cell infiltration. Panel (A) Distribution of risk scores across different age groups. Panel (B) Distribution of risk scores across different disease stages. Panel (C) Risk scores across different tumor grades. Panel (D) Distribution of risk scores across different histological types. Panels (E) Heatmap showing the correlation between immune cell types and risk scores. Panels (F) Correlation matrix showing the correlation between immune cell types and risk scores. Panel (G) Somatic mutation data in the high-risk group. Panel(H) Overview of somatic mutations in the low-risk group
S100A9 as a key prognostic biomarker in endometrial cancer
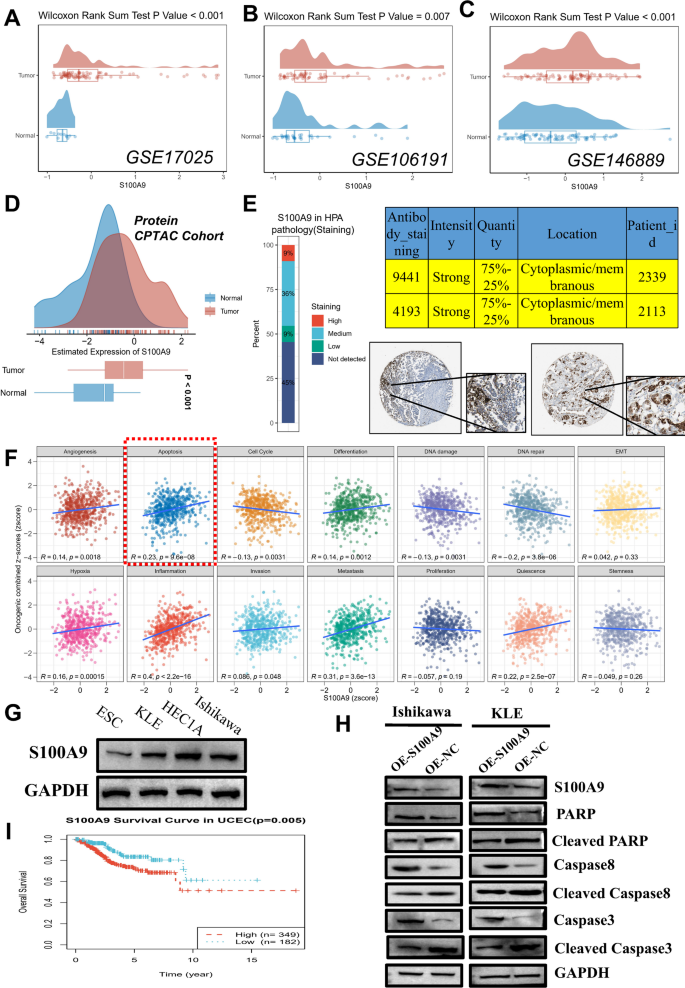
As one of the most important eosinophil extracellular trap (EET) regulators in machine learning models, we assessed the expression of S100A9 across various datasets to investigate its potential as a prognostic marker in endometrial cancer. Consistent with the TCGA-UCEC results shown in Figs. 1A and 6A-C presents the expression levels of S100A9 in different datasets (GSE17025, GSE106191, and GSE146889), demonstrating significantly higher expression in tumor samples compared to normal tissues, with Wilcoxon rank sum test p-values of < 0.001, 0.007, and < 0.001, respectively. Figure 6D provides a broader overview of S100A9 expression in the CPTAC protein cohort, showing a distinct shift toward higher expression levels in tumor samples. Protein expression of S100A9 in tumors is further supported by data from the Human Protein Atlas (HPA), where the majority of tumor samples show strong cytoplasmic and membranous staining (Fig. 6E). To explore the functional implications of S100A9 in cancer progression, we examined its correlation with various biological processes. The CancerSEA database, which categorizes different functional states of 14 tumor cell types, provided additional insights. Figure 6F shows the correlation between S100A9 expression and multiple oncogenic pathways, with strong positive correlations observed in the apoptosis pathway (R = 0.23, p < 1e-08). To validate these findings, we investigated the expression of S100A9 in different endometrial cancer cell lines. Figure 6G shows that S100A9 is expressed in multiple endometrial cancer cell lines, with significantly higher expression in tumor cell lines (Ishikawa, KLE, and HEC1A) compared to the normal ESC cell line. We selected KLE, which had relatively low expression, and Ishikawa, with the highest expression, for overexpression experiments. Figure 6H presents the results of these experiments, showing that S100A9 overexpression in Ishikawa and KLE cells leads to decreased levels of apoptotic markers, including cleaved PARP, cleaved caspase 3, and cleaved caspase 8, suggesting that S100A9 may influence apoptosis pathways in endometrial cancer cells. Finally, in light of the potential activation of the aforementioned oncogenic pathways, survival analysis in Fig. 6I demonstrates that S100A9 expression is significantly associated with patient outcomes in endometrial cancer. Patients with high S100A9 expression exhibited worse overall survival compared to those with low expression (p = 0.005).
Expression and Functional Implications of S100A9 in Endometrial Cancer. Panel (A–C) Expression levels of S100A9 in different datasets (GSE17025, GSE106191, and GSE146889). Panel (D) S100A9 expression in the CPTAC protein cohort. Panel (E) S100A9 protein expression in tumors, as shown by Western blot data from the Human Protein Atlas (HPA). Panel (F) Correlation between S100A9 expression and oncogenic pathways from the CancerSEA database. Panel (G) Western blot showing S100A9 expression in endometrial cancer cell lines (Ishikawa, KLE, HEC1A) compared to the normal ESC cell line (n = 3; quantified by ImageJ, normalized to GAPDH). Panel (H) Western blot results from overexpression experiments in Ishikawa and KLE cells, showing increased levels of apoptotic markers (n = 3; quantified by ImageJ, normalized to GAPDH). Panel (I) Kaplan-Meier survival analysis showing that high S100A9 expression in TCGA-UCEC is significantly associated with poorer overall survival compared to low expression
Pathway enrichment, and immune microenvironment analysis
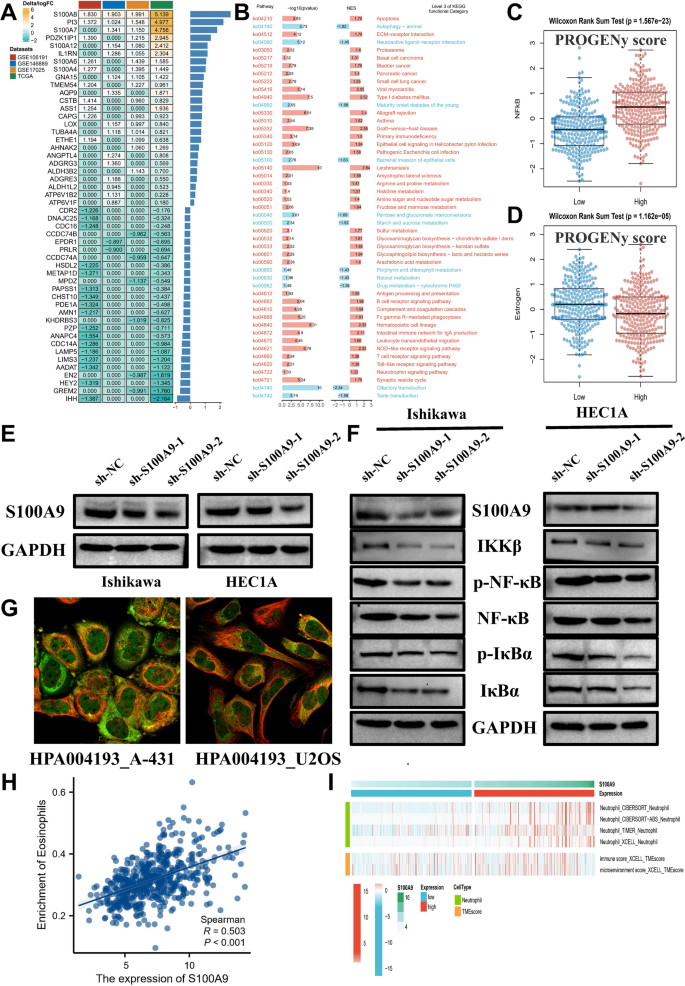
To identify key genes associated with endometrial cancer, we evaluated the consensus expression of the top 25 genes across multiple datasets. Figure 7A presents a heatmap showing the expression differences of these genes across various datasets. Positive values indicate higher expression in the high-expression group, while negative values reflect higher expression in the low-expression group. Based on these consensus genes, we conducted an enrichment analysis. Figure 7B shows significant activation of key oncogenic pathways, as inferred by the KEGG analysis, highlighting pathways associated with apoptosis and inflammatory response in the high-expression group, suggesting a link between these pathways and higher gene expression levels. To further validate these findings, we assessed the association between PROGENy scores and various oncogenic pathways in Figs. 7C-D. Figure 7C shows a significant relationship between NF-kB pathway activity and risk groups, with the high-expression group demonstrating significantly higher NF-kB pathway activity (p = 1.6e-23). Similarly, Fig. 7D illustrates that estrogen signaling pathway activity is significantly higher in the high-expression group (p = 1.6e-6), reinforcing the association between these pathways and higher gene expression levels. In Fig. 7E, we confirmed that S100A9 knockdown in endometrial cancer cell lines effectively reduced its expression. Figure 7F further validates the disruption of the NF-kB pathway in knockdown conditions, providing experimental support for the pathway activation observed in our computational analysis. Figure 7G presents immunofluorescence staining of A-431 and U2-OS cells from the HPA database, showing prominent cytoplasmic and membranous localization of S100A9, indicating its potential role in signaling processes within the cell. Finally, Fig. 7H shows a significant positive correlation between S100A9 expression and eosinophil infiltration (based on ssGSEA scores, R = 0.503, p < 0.001). Figure 7I further highlights the consensus immune cell infiltration differences of S100A9 across various algorithms (XCELL, TIMER, QUANTISEQ, EPIC, and CIBERSORT), suggesting that S100A9 may not only regulate eosinophil activity but also have a close relationship with neutrophil extracellular traps (NETs) or neutrophil functions in the tumor microenvironment.
Consensus Expression and Functional Implications of S100A9 in Endometrial Cancer. Panel (A) Heatmap displaying the consensus expression differences of the top 25 genes across multiple datasets. Panel (B) Enrichment analysis of key oncogenic pathways using KEGG in the high-expression group. Panel (C) Boxplot showing the relationship between NF-kB pathway activity and risk groups. Panel (D) Boxplot illustrating estrogen signaling pathway activity, with significantly higher activity in the high-expression group. Panel (E) Western blot analysis confirming S100A9 knockdown in endometrial cancer cell lines, effectively reducing its expression in these cells. Panel (F) Western blot analysis validating the disruption of the NF-kB pathway following S100A9 knockdown (n = 3; quantified by ImageJ, normalized to GAPDH). Panel (G) Immunofluorescence staining images from the HPA database, showing the cytoplasmic and membranous localization of S100A9 in A-431 and U2-OS cells. Panel (H) Scatterplot showing the positive correlation between S100A9 expression and eosinophil infiltration based on ssGSEA scores. Panel (I) Heatmap presenting the consensus immune cell infiltration differences of S100A9 across various algorithms (XCELL, TIMER, QUANTISEQ, EPIC, and CIBERSORT)




