LFAs
LFAs are widely used diagnostic tools for POCT due to their simplicity, rapid results, and cost-effectiveness. Typically, LFAs are developed to target a single biomarker and consist of two detection lines – a test line and a control line. In recent years, LFAs have become increasingly multiplexed, covering multiple test lines for various targets. For example, COVID-19 antibody LFAs have two test lines for IgG and IgM antibodies, while carbapenemases LFAs can have up to five test lines, targeting the most prevalent carbapenemases families. This feature increases the complexity of the visual interpretation of the outcome of the test. LFAs consist of a test strip that includes a sample pad, a conjugate pad, a nitrocellulose membrane with immobilized antibodies, and an absorbent pad52. When a sample (e.g., whole blood, serum, plasma, urine, stool, saliva, or swab in liquid media) is applied to the sample pad, it moves through the strip by capillary action. If the biomarker of interest is present, it reacts with specific antibodies and binds to the capture regions, forming visible detection lines that indicate the presence or absence of the target analyte53. This straightforward design makes LFAs suitable for various applications, including infectious disease detection54,55,56, cardiac biomarker detection57, pregnancy tests58, cancer diagnostics59, and environmental monitoring60,61, among many others. From a commercial and engineering perspective, LFAs offer significant manufacturing advantages due to their simplified device architecture with low-cost materials and a long shelf life of approximately two years under ambient conditions, making them highly practical for commercial applications62. The latest progress in LFA research mainly focuses on improving the analytical performance by engineering physical components/parameters of LFAs, such as introducing sensitive conjugate labels/modalities63,64,65,66,67,68,69 and engineering advanced fluidic structures70,71,72. Additionally, distance-based LFAs have emerged as a promising alternative to traditional LFAs by using changes in visual distance as a quantitative readout instead of relying on color intensity at a fixed test line73,74. Various efforts to improve LFA performance through image/data-processing algorithms have also been introduced75.
Integration of AI and ML in LFAs presents new methodologies to improve performance. This integration has been driven by the need to further improve sensitivity, specificity, result interpretation and quantification and to shorten the testing time. A notable trend in this integration is the use of smartphone or tablet-based digital devices to read test results75,76,77. This not only enhances the usability and access to LFAs but also allows for the capture and collection of images of the test results in a digital format suitable for computational processing and interpretation without bias and reduces data loss40. Specifically, ML technologies for LFAs focus on several key areas as outlined in Table 1: (i) automating the interpretation of test results to reduce human error and improve result consistency16,17,40,41,78,79,80,81,82,83,84, (ii) improving sensitivity and accuracy through noise-tolerant analysis algorithms16,85,86,87, and (iii) transforming traditionally qualitative tests into quantitative assays for more precise diagnostics16,78,81,83,85,86.
Several studies have highlighted the critical role of ML in improving the performance of LFAs (Fig. 2); for example, a team at University College London (UCL) and the Africa Health Research Institute created an image library of over 11,000 real-world images of HIV LFAs acquired in KwaZulu South Africa (Fig. 2a)40. The ML models accurately classified tests as positive or negative, significantly enhancing the specificity and sensitivity of decision support compared to visual interpretation by nurses and community health workers. This improvement was measured at 11% (from 89% to 100%) for specificity and 2.8% (from 95.6% to 97.8%) for sensitivity, reducing the risk of both false positives and negatives. Consequently, the system elevated the positive predictive value by 11.3% (from 88.7% to 100%) and the negative predictive value by 2.3% (from 95.7% to 98%), demonstrating its increased reliability in minimizing misclassification errors in POCT settings. Given that hundreds of millions of these tests are performed annually worldwide, this improvement could have major health and economic benefits to populations. Moreover, the UCL team also applied the models to COVID-19, and evaluated over 500,000 LFA images as part of the world’s largest sero-surveillance study, REACT-2 (Real-time Assessment of Community Transmission-2) (Fig. 2b)41.
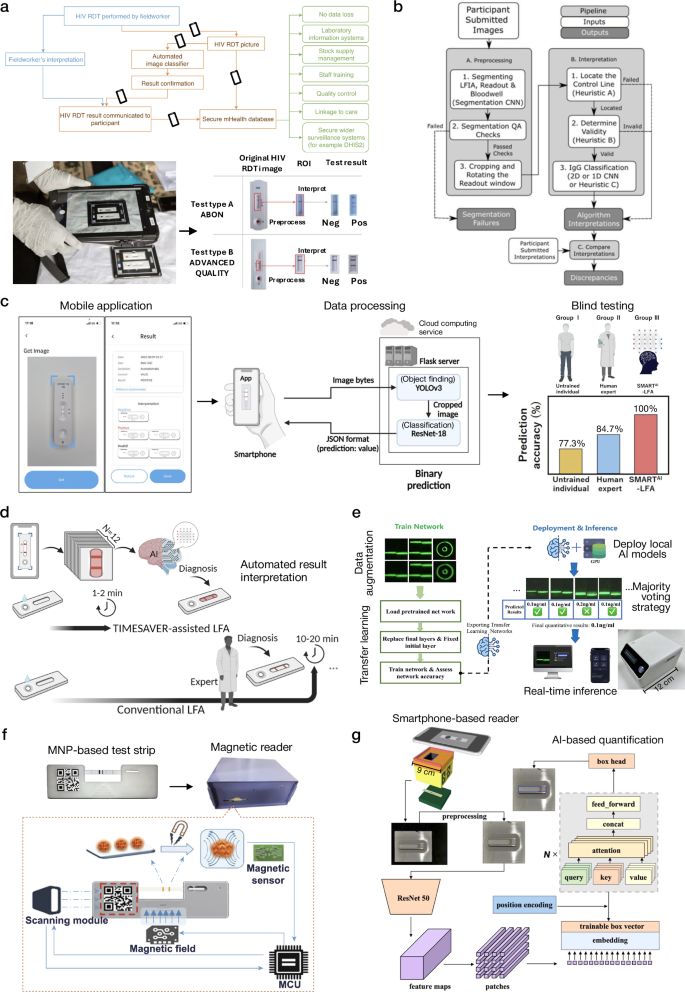
a Infographics illustrating the benefits of HIV LFA processing in field settings (top), and HIV LFA capturing and analysis procedures used to enhance sensitivity and specificity for the detection of HIV in field settings (bottom). b LFA processing pipeline used for the automated interpretation of the REACT-2 dataset. c The SMARTAI-LFA platform used deep learning for automated image processing and result interpretation, utilizing this technology to detect the presence of the SARS-CoV-2 antigen. d The TIMESAVER-LFA platform employed an AI-based verification algorithm to reduce assay time to 1–2 min. e The fluorescent UCNP-based LFA used transfer learning to enhance sensitivity and quantification capability for the detection of methamphetamine and morphine. f An ML approach was used in MNP-based LFA to improve analytical performance for the detection of hCG and multiple cardiac biomarkers. g The PDA nanoparticle-based LFA for the detection of COVID-19 neutralizing antibodies offered precise quantification of antibody concentrations through AI-based analysis. a–d, f These are adapted with permission from refs. 16,17,40,41,86, by Springer Nature; e This is adapted with permission from ref. 85 by Elsevier; g This is adapted with permission from ref. 78 by Elsevier.
As another example, for automated image processing and result interpretation, Lee et al. introduced a deep learning-assisted smartphone-based LFA (SMARTAI-LFA) platform integrating a two-step CNN model (Fig. 2c)16. This model includes automated object detection using the You Only Look Once (YOLO)v3 network and test result classification with ResNet-1888, providing accurate COVID-19 test results without the need for expert interpretation. YOLOv3 is a real-time object detection model that efficiently identifies multiple objects in a single pass through the image, ensuring high speed and accuracy. This system, which incorporates a smartphone application for cloud-based processing, achieved 98% accuracy across clinical tests captured by different users and smartphones. In the blind testing phase, this SMARTAI-LFA outperformed both untrained users and human experts, demonstrating the superior diagnostic accuracy of ML-enabled LFAs. The same authors demonstrated another approach, a time-efficient immunoassay with AI-based verification (TIMESAVER) (Fig. 2d)17. This system used a combination of neural network models to accurately identify detection regions in LFA. TIMESAVER reduced the assay time to 1–2 min compared to the traditional methods of 15 min, powered by a time-series deep learning algorithm. The system was applied to infectious diseases (e.g., COVID-19, influenza) and various biomarkers (e.g., cardiac troponin I [cTnI], human chorionic gonadotropin [hCG]), achieving high sensitivity and specificity on a spiked dataset (by spiking the target protein in running buffer). When blindly tested on clinical samples, the TIMESAVER system showed high accuracy in ~2 min of testing time, significantly faster than manual analysis by human experts, which took 15 min, showcasing the potential of ML to enhance paper-based point-of-care diagnostics.
To further improve the analytical performance (i.e., sensitivity and accuracy) and quantification capability of LFAs, ML algorithms were coupled with enhanced sensing modalities (i.e., fluorescence, magnetic nanoparticles, or optimized particles for enhanced absorption). For example, a study by Wang et al. utilized a transfer learning approach in combination with an LFA with fluorescent upconversion nanoparticles (UCNP) for the quantitative detection of methamphetamine and morphine (Fig. 2e)85. Transfer learning is an ML technique in which a pre-trained AI model applies its acquired knowledge to a new problem, rather than learning from scratch, thereby enhancing efficiency and adaptability to new datasets. In biomedical sensing-related applications, transfer learning addresses challenges such as limited annotated data, balancing computational costs with large datasets, and improving generalization across various application scenarios. In a study by Wang et al., transfer learning improved the model’s ability to detect low concentrations of analytes, reducing overfitting and improving generalization. It also ensured reliable analyte quantification in noisy environments. This approach simplified the detection process, making it feasible for use in portable POCT devices. A comparative analysis revealed that models trained without transfer learning exhibited drastically increased training times. These findings underscore the efficiency of transfer learning in biomedical POCT applications, ensuring reliable analyte quantification while optimizing computational resources.
As another important example, Yan et al. developed an approach to improve the analytical performance of magnetic nanoparticle (MNP)-labeled LFAs (Fig. 2f)86. SVMs are classification algorithms that identify the optimal decision boundary to separate data points with maximum margin. Their effectiveness with small datasets and capacity to manage complex, high-dimensional, and noisy data makes them widely applicable in medical diagnostics and image recognition. By integrating an SVM-based classifier, the study improved the sensitivity and accuracy of detecting weak signals by effectively classifying complex patterns based on MNP signals. For example, at hCG concentrations of 0.25 mIU/mL, the accuracy of the SVM classifier was 100%, while that of visual reading was 7%. In addition, a novel waveform reconstruction method was introduced to accurately restore and interpret distorted waveforms of weak magnetic signals, thereby enhancing the assay’s quantification capability. The application was validated by successfully quantifying hCG with a limit-of-detection (LoD) of 0.014 mIU/mL and a dynamic range of 1–1000 mIU/mL. This method has also been applied to analyze multiple test lines on a single test strip for several cardiac biomarkers – cTnI, creatine kinase isoenzyme MB (CK-MB), and myoglobin – demonstrating strong correlation with standard concentrations and showcasing the multiplexing potential in LFAs. For CK-MB and myoglobin, their quantification cut-offs reached 0.5 ng/mL and 5 ng/mL in diluted serum, respectively, covering clinically relevant ranges when compared to negative samples. However, for cTnI, the quantification cut-off was limited to 0.5 ng/mL, falling short of the ~0.01 ng/mL sensitivity required for clinical use.
As another method for improved LFA performance, Tong et al. utilized polydopamine (PDA) nanoparticles to enhance the colorimetric signal using a vision transformer (ViT), which was trained to quantify the test results to detect COVID-19 neutralizing antibodies (Fig. 2g)78. ViT processes images by dividing them into patches and leveraging a transformer model to capture spatial relationships throughout the entire image. Compared to CNNs, ViTs achieve high performance with less data and effectively comprehend long-range dependencies. This study employed the ViT framework to accurately predict the position of the test strip in the smartphone-captured image. The use of PDA nanoparticles improved the sensitivity of the assay, which was further advanced by using a ViT-based neural network that measured the intensity changes on the test strips captured by a smartphone-based reader. The ability of the neural network to accurately interpret these changes resulted in precise quantification of target antibody concentrations, overcoming the limitations of traditional visual assessment and making it an effective tool for evaluating vaccine efficacy in clinical and point-of-care settings.
VFAs
VFAs have recently emerged as a promising alternative testing platform to LFAs, offering shorter sample-to-answer time and improved multiplexing capabilities89. In contrast to LFAs, which rely on lateral fluid flow, VFAs utilize a vertical liquid flow through stacked paper layers within a millimeter distance range, ensuring rapid sample/reagent migration and faster assay time. This unique fluidic design also enhances multiplexing capability by compartmentalizing sensing regions with a patterned reaction membrane, thereby minimizing cross-talk between detection zones. While VFAs require a relatively more complex fabrication and assembly process compared to LFAs and involve a slightly more intricate operation, they remain highly viable for POCT applications. Moreover, both LFAs and VFAs share the same materials and fundamental assay principles, such as antibody immobilization within the paper substrate and sample interaction with optical probe conjugates90, making them complementary approaches. Typical VFAs utilize colorimetric90 or surface-enhanced Raman scattering (SERS) detection modalities91,92,93, covering various applications demonstrated so far89,90. In this section, we will primarily focus on the applications where computational models and ML were used for the interpretation of the VFA signals, the improvement of VFA diagnostic/sensing accuracy and the optimization of the VFA design (see Table 2).
Conventional ML methods, including linear and polynomial regression models, have been widely applied for VFA analysis. Some of these applications include multiplexed detection of cancer biomarkers94, quantification of metabolites in sweat95, and detection of inflammatory markers in serum96 (Fig. 3a). However, these previous approaches often fail to accurately capture the statistical features of the input signals, leading to erroneous predictions and high variability, especially for multiplexed analyses and real-life biological samples affected by various noise factors such as the sample matrix effect. In contrast, advanced ML models like neural networks can provide better consistency due to their representation power and ability to learn complex relationships between noisy assay signals and ground truth information36.
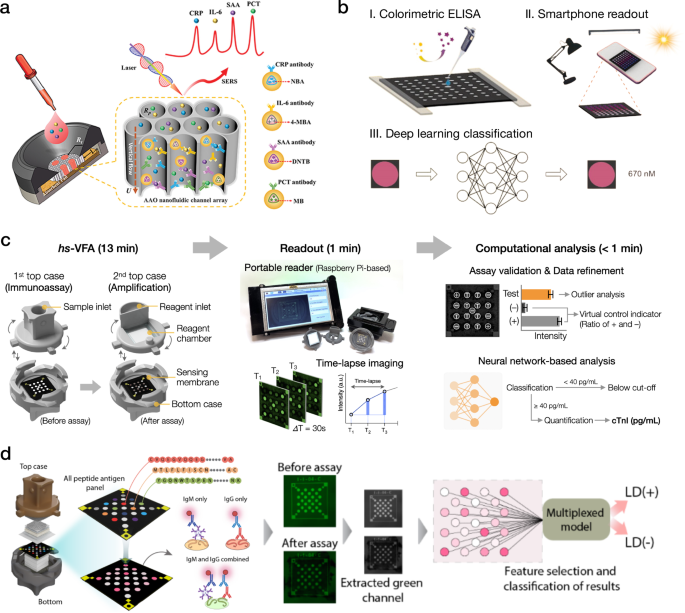
a SERS-based VFA processed by linear regression models for quantitative detection of inflammation markers. b Colorimetric ELISA (c-ELISA) processed by neural networks for accurate detection of Rabbit IgG under different illumination conditions. c Deep learning-enhanced xVFA for high-sensitivity detection of cTnI. d Peptide-based xVFA processed by deep learning models for single-tier testing of Lyme disease. a This is adapted with permission from ref. 96 by Wiley; b This is adapted with permission from ref. 97 by Elsevier; c This is adapted with permission from ref. 100 by American Chemical Society; d This is adapted with permission from ref. 102 by Springer Nature.
Neural network-based inference models (AlexNet, GoogLeNet, ResNet34, MobileNetV2) were used to enhance the adaptability and robustness of paper-based colorimetric enzyme-linked immunosorbent assays (c-ELISA)97. The activated assays were captured by a smartphone camera under three different lighting conditions and then processed by deep learning algorithms, achieving a detection accuracy of >97% in detecting Rabbit IgG in synthetic samples with GoogLeNet (Fig. 3b). As another example, Lee et al. employed a three-dimensional (3D) µPAD to detect glucose through a colorimetric assay processed by neural network models, achieving 91.2% in classifying glucose levels as low, normal, and high when tested on glucose-spiked buffer samples. The same study also proposed a microfluidic thread/paper-based analytical device (µTPAD) to detect glucose levels in artificial saliva and showed a high correlation between the predicted and the ground truth glucose levels98. These platforms applied ML methods to enhance the scalability and adaptability of VFA readout. However, the performances of these µPAD platforms were primarily demonstrated on synthetic and spiked samples, necessitating further testing on biological specimens to confirm their viability for real-life applications.
Although the fundamental principles of various VFA platforms are similar, their design and operational characteristics may differ, leading to potential inconsistencies in the output signals across different assays. These variations stem from a sophisticated interaction between assay and readout parameters, making the standardization of these processes a challenging endeavor. To address this issue, Tay et al. proposed a comprehensive physics-driven framework to model key assay operation parameters, demonstrating reliable generalization to different VFA formats and sample matrices99. The framework incorporated adaptable physical equations, covering typical paper-based assay procedures such as sample mixing, sample propagation through paper layers, analyte immobilization, and assay readout. The researchers validated their model on VFAs with varying immunoassay formation procedures, reporter label concentrations, and sample matrices (i.e., buffer, urine, saliva, and sweat), demonstrating good agreement with their experimental results. For example, the observed discrepancy between the predicted and experimental results was <10% when varying the label concentration or the washing volume, and <20% when altering the assay formation procedure. The overall high concordance of the model with experimental data and the minimal requirements for the training set size make the proposed workflow an appropriate tool for accelerated assay optimization and adoption.
In a different application, ML-driven VFAs were employed to achieve accurate cardiac biomarkers quantification in patient serum samples. For instance, Ballard et al. proposed a multiplexed VFA (xVFA) for the quantification of C-reactive protein (CRP), i.e., for high-sensitivity CRP (hs-CRP) testing, leveraging xVFA’s multiplexing capabilities and ML to improve the sensor’s dynamic range and quantification accuracy19. Researchers demonstrated accurate quantification of CRP in the high-sensitivity range (0–10 µg/mL), achieving an R2 of 0.95 compared to a Food and Drug Administration (FDA)-approved analyzer. Additionally, they were able to successfully identify elevated CRP concentrations outside the high-sensitivity range, within the acute inflammation range (up to ~1000 µg/mL), mitigating the hook effect present in singleplexed testing platforms. Furthermore, Han et al. used the xVFA platform for high-sensitivity cTnI (hs-cTnI) testing (Fig. 3c)100,101. cTnI is a gold-standard protein biomarker for diagnosing myocardial infarction (MI), and yet presents a challenge for POCT due to its extremely low clinical concentrations. To achieve high sensitivity, researchers integrated Au-ion amplification chemistry into the xVFA and demonstrated a high-sensitivity VFA (hs-VFA) with a colorimetric modality, which achieved trace-level cTnI detection by increasing the diameter of the optical absorption labels (AuNPs). During the amplification stage, the AuNP diameter increased from 15 nm to >200 nm, leading to a significantly higher signal. The hs-VFA showed a LoD of 0.2 pg/mL, which is >10 times lower than the clinical cutoff concentration in hs-cTnI testing. Fully connected neural network (FCNN) models were employed to exclude outlier sensors, enhance the precision of cTnI testing, and accurately quantify cTnI concentration in serum. The computationally optimized hs-VFA demonstrated a high correlation with an FDA-approved analyzer (Pearson’s r > 0.96) and good repeatability with a <6.2% coefficient of variation (CV) between duplicate repeats of the tested samples, meeting clinical requirements for hs-cTnI testing.
The xVFA platform was also used for the multiplexed quantification of cardiac markers in serum. Researchers integrated fluorescence-based detection into the xVFA platform and applied this fluorescence-based xVFA (fxVFA) for the parallel quantification of three cardiac markers (i.e., myoglobin, CK-MB, and heart-type fatty acid-binding protein [h-FABP]) in serum samples30. The assay contained a total of 17 immunoreaction spots/channels, including 6 testing spots with 2 spots per biomarker type and 11 control spots. The activated fxVFA was captured by a hand-held and cost-effective fluorescence reader, and the signals of the immunoreaction channels were processed by neural network models for the multiplexed quantification of the biomarkers. Three distinct FCNN models were developed, one model per cardiac marker, and in addition to inferring the biomarker concentration in the sample, the models were also used to optimize the subset of immunoreactions needed for each biomarker. The optimal spot configurations included spots specific to the target biomarker, as well as cross-reactive spots and control spots. The neural networks successfully learned from complex cross-reactive patterns between different immunoreactions, achieving superior performance compared to standard linear and polynomial regression models that were applied to the same patient samples. Quantification on the blind testing set, composed of 16 patient serum samples, showed a high correlation (R2 > 0.9) with ground truth ELISA measurements for all three cardiac markers.
In addition, the xVFA platform processed by ML algorithms was applied for various serological testing applications, providing binary diagnostics of bacterial and viral diseases. For example, ML was employed to achieve accurate Lyme disease diagnostics using multiplexed serological testing on an xVFA platform18,102,103. Conventional Lyme diagnostics involves a laborious two-tier testing procedure and has sensitivity limitations, particularly at the early stages of Lyme disease. To facilitate Lyme disease diagnostics without compromising the accuracy of two-tier testing, Joung et al. proposed an xVFA processed by a smartphone-based reader and FCNN models for the serological testing of Lyme disease. This xVFA requires only 20 µL of patient serum and takes 15 min to operate18. The assay leveraged multiplexed detection of IgG/IgM antibodies across a panel of Lyme antigens, with ML utilized for two distinct purposes: (1) computational optimization of the assay design and (2) diagnostics of Lyme disease from patient serum samples. Computationally optimized assay achieved 90.5% sensitivity and 87.0% specificity on a blind testing set. Importantly, in addition to the improvements in diagnostics performance, computational assay optimizations also reduced the per-test cost, achieving 44% cost reduction when implementing the optimal subset of immunoreactions at the xVFA. Further improvement of xVFA performance for Lyme disease was achieved by transitioning to a peptide panel assay (Fig. 3d), resulting in 95.5% sensitivity and 100% specificity despite the presence of early-stage Lyme samples within the testing set102.
Another serological test implemented using the same xVFA platform was used to monitor human immunity levels in response to SARS-CoV-2 infection and vaccination, accurately identifying unprotected, protected, and infected cases104. The multiplexed design of the xVFA platform allowed for the incorporation of multiple structural proteins of the virus within a single cartridge. The combined responses from these proteins were processed by FCNN models to classify patient immunity levels. Prior to training the final models, the assay was computationally optimized by selecting an optimal subset of the proteins for both IgG and IgM panels, resulting in the highest testing accuracy. The blind testing on 31 serum samples from 8 individuals showed an accuracy of 89.5%, enabling reliable tracking of the immune response dynamics over time. The integration of diverse SARS-CoV-2 proteins into a unified multiplexed panel represents a robust alternative to singleplex tests, with the potential to encompass broader populations, including those vaccinated with different vaccine types and exhibiting diverse immune responses.
Recent work has also demonstrated electrochemical-based VFAs, showcasing an example where the assay signals are processed by a field-effect transistor (FET) and deep learning for cholesterol detection in patient serum105. In this setup, the cartridge comprises a paper membrane placed over an ion-sensitive sensing electrode, connected to the FET gate. When the sample reaches the membrane, cholesterol-specific enzymes produce protons in response to the cholesterol concentration, which is recorded in FET transfer curves captured repeatedly over a 5-min interval. These transfer curves are further converted into a two-dimensional (2D) heatmap reflecting the kinetic details of the enzymatic reactions. Lastly, a shallow FCNN model optimizes the interval of kinetic data carrying a concentration-specific response and quantifies the cholesterol concentration in serum using the optimal transfer curve signals. Quantification results on 30 blindly tested serum samples exhibited a strong correlation (R2 > 0.976) with a CV of <7% when compared against the ground truth results from a CLIA-certified clinical laboratory. In the same work, the authors have also adopted this VFA platform to immunoassay format, potentially offering a wide range of applications in POCT105.
As a broader category encompassing VFAs, array-based sensors represent advanced bio/chemical sensing platforms that consist of miniature arrays of distinct sensing elements systematically arranged on a substrate106. These systems enable high-throughput and multiplexed detection, allowing simultaneous measurement of multiple analytes. VFAs can be considered a subset of array-based sensors, as they also employ reaction compartmentalization or multiple-spot arrays on the sensing membrane. However, a key distinction lies in the reagent delivery mechanism—while array-based sensors typically involve direct exposure of reagents in liquid or gas form to the sensor surface107,108,109,110, VFAs rely on a controlled fluid delivery through stacked paper layers to transport samples and reagents to the sensing region. Each microdot within the array can independently capture and detect specific analytes, producing complex, high-dimensional data outputs ideal for analysis with ML techniques. The combination of multiple dot arrays and ML leverages the array’s capability for simultaneous, multi-analyte detection and enhances data interpretation by efficiently processing subtle variations in signal intensity, spatial distribution, or optical changes.
One example of ML-enhanced array-based sensing for POCT is a recent study by Kim et al., which introduced a fluorescent microarray designed for mobile reader-based high-throughput volatile organic compound (VOC) sensing110. This system employed 75 different fluorescent Kaleidolizine derivatives, immobilized on a wax-printed cellulose substrate, where each fluorophore exhibited unique fluorescence intensity and color shifts upon VOC exposure. To decode the complexity of these fluorescence variations, the study utilized a random forest algorithm to analyze hue differences in fluorescence patterns, achieving 97% classification accuracy across five VOCs. This work highlights the necessity of ML in multiplexed sensing applications, as the subtle fluorescence shifts could not be reliably interpreted using conventional threshold-based approaches.
Expanding upon this approach, Yang et al. applied a VOC-based array sensor for bacterial identification, leveraging the distinct metabolic signatures emitted by different bacterial species109. Their study introduced an ML-enabled paper chromogenic array (PCA), consisting of 23 chromogenic dyes and dye combinations, which underwent colorimetric shifts upon exposure to bacterial VOC emissions. Unlike traditional microbiological techniques requiring extended culturing, this method enabled rapid and non-destructive bacterial detection. A neural network model was trained to classify bacterial strains based on digitized color changes, achieving 91–95% strain-specific identification accuracy. Notably, this system successfully distinguished between viable E. coli, pathogenic E. coli O157:H7, and Listeria monocytogenes, demonstrating its potential for food safety applications. While the imaging step in this study was performed using a desktop scanner, the system can be readily adapted for smartphone-based imaging, presenting strong potential for future POCT applications.
NAATs
NAATs detect the target genetic material of interest by amplifying it using methods such as polymerase chain reaction (PCR)111, loop-mediated isothermal amplification (LAMP)112, recombinase polymerase amplification (RPA)113, and rolling circle amplification (RCA)114,115. These methods have become increasingly prevalent for COVID-19 detection and have been essential for molecular diagnostics over the past decade116, enabling precise detection and quantification of deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). Importantly, reverse transcription PCR (RT-PCR) is widely adopted as the gold standard for diagnosing viral infections and analyzing gene expression111, due to its high accuracy and specificity. Alternative isothermal methods, such as LAMP, RPA, RCA, and CRISPR-mediated117 nucleic acid amplification, eliminate the need for thermal cycling and expensive instrumentation116,118, making them more suitable for bedside or in-field settings119. Moreover, the integration of paper-based LFAs and microfluidic devices further enhances the accessibility and practicality of NAATs120.
Despite the merits of NAATs summarized above, these technologies have been hindered by relatively lengthy assay times, low signal intensities at low copies of target sequences, lack of objective and automated result interpretation, and reliance on benchtop equipment121. To overcome these challenges and streamline the deployment of testing in point-of-care settings, NAATs have been adopted to include four main steps: (i) the collection of sample swabs (e.g., from nasopharyngeal [NP] cavities), (ii) the introduction of samples to portable diagnostic devices, (iii) the amplification of nucleic acids using custom thermal modules, and (iv) the prediction or classification of positive and negative samples through ML-integrated optical mobile devices122,123. Integration of AI/ML into these portable readout devices has been crucial for advancing NAATs and decentralizing their applications through POCT124.
The integration of AI/ML into NAATs can address the limitations mentioned above by significantly reducing assay time, eliminating subjective result interpretation, and maintaining high accuracy124. Many POCT NAAT diagnostic tools are equipped with comprehensive AI models such as CNN122, long short-term memory (LSTM)20, recurrent neural network (RNN)21, transformer125, and gated recurrent unit (GRU)126. These AI models enable advanced image processing, sample classification, early result prediction, and high-throughput automated screenings127. The deployment of these AI models enables the analysis of diagnostic results at lower signal intensity levels, conversion of qualitative metrics into quantitative data for early prediction, and achievement of high accuracy, sensitivity, and specificity within much shorter assay times (e.g., <20 min125,126,128). The specific examples are outlined in Table 3 and in the next paragraphs of this section.
ML-integrated POCT NAAT systems have significantly advanced after the COVID-19 pandemic, driven by the critical need for cost-effective, accurate, sensitive, and high-throughput screening of viral infections118. Recent studies have demonstrated the successful incorporation of AI/ML into these systems with various sensing modalities (e.g., colorimetric122,127, fluorescent20,125,126, electrochemical129, and SERS21) for SARS-CoV-2 RNA detection (Fig. 4), showcasing the versatile and general learning capabilities of modern AI and ML models130. For example, Rohaim et al. developed a hand-held AI-LAMP system for SARS-CoV-2 RNA (RdRP gene) detection, which automated image acquisition and processing and reduced assay time and subjectivity of colorimetric LAMP (Fig. 4a)122. This system utilized CNN to classify colors more accurately, achieving 98% accuracy and significantly higher sensitivity than the gold standard quantitative RT- PCR (qRT-PCR). Furthermore, Jaroenram et al. developed a dual RT-LAMP assay that utilized both colorimetric analysis and a deep learning detection transformers (DETR)-integrated analysis tool (RT-LAMP-DETR) on a smartphone for ultrasensitive COVID-19 detection (Fig. 4b)127. In addition to pH-sensitive indicators that provided a visual indication of results, high-throughput analysis of these colorimetric results was supported by RT-LAMP-DETR through cross-comparison. This result interpretation achieved 100% accuracy, sensitivity, and specificity, validating a scalable method for the screening of COVID-19 suitable for low-resource settings.
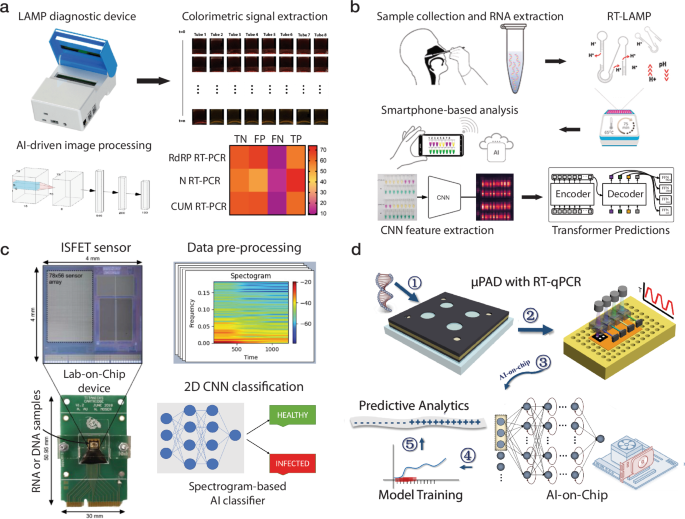
a A hand-held AI-LAMP device for rapid detection of COVID-19 with AI-based image analysis reduced the sample-to-answer time and improved signal interpretation. b A one-step smartphone-based colorimetric RT-LAMP COVID-19 screening method enabled by pH-sensitive dyes and a transformer AI model. c A lab-on-chip nucleic acid amplification device that utilized ISFET arrays and a spectrogram-based CNN to classify COVID-19 and three cancer biomarkers, featuring a compact size and improved accuracy. d An AI-aided on-chip µPAD for COVID-19 detection using three neural network models (i.e., RNN, LSTM, and GRU) for qualitative analysis. a This is adapted with permission from ref. 122 by MDPI; b–d These are adapted with permission from refs. 126,127,129, respectively, by Elsevier.
Additionally, Tripathi et al. introduced a low-cost lab-on-chip platform that transformed ion-sensitive field-effect transistor (ISFET) data into spectrograms compatible with CNNs to identify nucleic acid amplification (Fig. 4c)129. This method, in addition to efficiently identifying infectious diseases and cancer biomarkers, achieved 84.84% accuracy with a 30kB-sized CNN model, facilitating its deployment on edge computing devices. Similarly, Sun et al. developed a portable optoelectronic system interfaced with paper microfluidics and deep learning for the real-time detection of SARS-CoV-2 (Fig. 1f)20. This device transfers real-time data from fluorescent signals of amplified viral sequences to RNN, LSTM, and GRU models, enabling early predictive analysis and reducing assay time by 45%. This model demonstrated robust outcomes for NAATs, with AI-integrated early prediction achieving 98.1% accuracy, 97.6% sensitivity, and 98.6% specificity.
Moreover, a system reported by Sun et al. integrated deep learning with µPADs for the rapid and accurate detection of SARS-CoV-2 RNA (ORF1ab gene), using quantitative PCR (qPCR) data and AI predictive analysis (Fig. 4d)126. The analysis was driven by RNN, GRU, and LSTM models to derive qualitative forecasting from real-time PCR analytics of patient samples. The GRU was found to be the most accurate in predicting end-point values and trends of qPCR curves, with a mean absolute percentage error of 2.1%. The accurate predictions made as early as after 13 amplification cycles accelerated the NAAT procedure, reducing the assay time to 12 min and allowing better preparedness for future disease outbreaks.
Similarly, Yang et al. combined SERS sensors with deep learning algorithms for the rapid detection of SARS-CoV-2 RNA in human NP swab specimens (Fig. 1g)21. A silver nanorod (AgNR) array sensor functionalized with DNA probes was used for detecting viral RNA. Specifically, SARS-CoV-2 RNA selectively hybridized with complementary DNA sequences immobilized on the AgNR substrate, inducing spectral changes detectable by SERS. These spectral shifts were then analyzed using an RNN model, which classified the results with an accuracy of 97.2% for positive specimens and 100% for negative specimens, providing results within 25 min. As an integration of paper microfluidics with NAATs, Sun et al. introduced an approach that combined paper microfluidics with deep learning and cloud computing for accelerated SARS-CoV-2 RNA analysis125. Real-time amplification of synthesized RNA templates was performed on paper materials, and the time-series data were transmitted to a cloud server with preloaded deep learning models for predictive analysis, achieving clinical accuracy, sensitivity, and specificity of 98.6%, 97.6%, and 99.1%, respectively.
Instead of generating numerous copies of target genes to amplify signals, CRISPR-mediated nucleic acid detection leverages the collateral cleavage activity of CRISPR-associated enzymes, where abundant non-target signal reporters are cleaved to produce a detectable signal131. A notable advancement in this field was reported by Roh et al.132, who developed a CRISPR/Cas12-based multiplexed nucleic acid detection platform utilizing spatially encoded hydrogel microparticles (HMPs). This study integrated ML algorithms to automate the analysis of individual HMPs, employing neural networks to recognize/classify coded HMPs and perform segmentation for fluorescence detection. Upon binding of Cas12a/gRNA complexes to human papillomavirus RNA in cervical brushing samples, the HMPs fluoresced and subsequently captured within a microfluidic device, where a Mark R-CNN model automatically identified them. The trained model achieved a high accuracy of 97.9% and an F1 score of 97.8% in the validation tests, successfully differentiating four distinct HMP types. This rapid assay demonstrated an attomolar LoD of 2 aM (equivalent to 1.2 copies/µL), highlighting its high sensitivity.
Beyond CRISPR-mediated approaches, nanoparticle-based virus detection has also gained attention. Draz et al.133 developed a CNN-enabled smartphone system for detecting intact viruses on a microchip using the nanocatalytic activity of platinum nanoparticles (PtNPs). PtNPs catalyze the decomposition of hydrogen peroxide into oxygen gas, generating visual bubble patterns upon immuno-capturing virus particles (e.g., Zika virus [ZIKV], hepatitis B virus, and hepatitis C virus) on a microchip. Building upon these advancements mentioned above, Shokr et al.134 integrated CRISPR-based detection with nanoparticle technology by conjugating an anti-Cas9 antibody with PtNPs, enabling ZIKV RNA detection. This system facilitated reliable bubble generation within the microchip, achieving a LoD of 400 aM for ZIKV detection. Furthermore, this platform incorporated adaptive adversarial learning and generative adversarial networks to enhance its adaptability to emerging pathogens, allowing rapid diagnostic reconfiguration without extensive retraining. Additionally, the system augmented real smartphone-taken image datasets to improve generalizability, making it highly suited for low-cost, smartphone-based diagnostics. Collectively, these ML-integrated NAAT advancements provide highly sensitive, portable, and scalable molecular diagnostic solutions, significantly contributing to epidemic preparedness by offering low-cost, rapid, and accurate nucleic acid detection methods.
In addition to SARS-CoV-2 RNA detection, ML-integrated NAAT technologies have been applied to detect other targets such as bacteria DNA22, cancer biomarkers135, and λ DNA136. For example, Guo et al. proposed a smartphone-based DNA diagnostic tool for malaria that integrated CNN models for local decision support and blockchain technology for secure data management and reporting (Fig. 1h)22. This system used streptavidin-labeled red particles to detect target sequences for a colorimetric readout and employed a CNN model to interpret the results, achieving 97.83% accuracy. A key feature of this system was its blockchain-enabled data security, ensuring tamperproof record-keeping and controlled access to diagnostic results. Unlike traditional cloud-based methods, blockchain technology provides a low-power, cost-effective alternative for securely transmitting medical data in resource-limited settings. This approach eliminated the need for specialized infrastructure, making it particularly suitable for decentralized healthcare applications. The system was successfully field-tested in rural Uganda, where it exhibited high diagnostic accuracy (correctly identifying over 98% of tested cases), reliable data transfer, and seamless integration into local healthcare workflows. By combining ML-driven image analysis with blockchain-based security, this platform provides a scalable, privacy-preserving framework for real-time disease surveillance and secure diagnostics in underserved regions.
Furthermore, AI/ML technologies have also been used for digital NAATs. Digital NAATs have the unique advantage of extremely high sensitivity (usually 1–10 copies/µL) to their target sequences and absolute quantification without the need for standard curve-based calibration137. For example, Li et al. developed an all-in-one OsciDrop digital PCR/LAMP system that can perform multiplexed fluorescent quantification of cancer biomarkers (HER2 and EGFR genes)135. U-Net and MobileNet models were used for ML-integrated droplet image analysis, enabling highly accurate and consistent nucleic acid concentration quantifications (CV < 1.5%). Another example of digital NAATs is Fractal LAMP, which utilized a computer vision algorithm to achieve high accuracy for the detection of amplified DNA by recognizing the LAMP byproducts that form fractal structures136. A Bayesian model and bootstrapping method facilitated high-accuracy concentration measurements over 3 orders of magnitude. Although these assays and detection procedures may still require benchtop equipment, their potential for miniaturization into portable devices renders them as emerging POCT technologies. In the future, combined progress in device engineering, along with the high sensitivity and absolute quantification ability offered by digital ML-integrated NAATs will substantially expand the application of precision medicine, even in rural areas and low-income countries.
Imaging-based point-of-care sensors
Recent advancements in AI and ML have significantly enhanced image-based diagnostics for various diseases138,139,140,141,142,143,144,145. For example, deep learning models, particularly CNNs and ViTs, have achieved dermatologist-level accuracy in skin cancer detection and lesion classification, leveraging large, annotated datasets to improve diagnostic precision138,139,140. As another example, AI is being integrated into echocardiography, cardiac magnetic resonance imaging, and coronary computer tomography angiography, enabling automated segmentation, disease detection, and risk stratification with performance comparable to human experts143,144,145. These advancements and many others underscore the growing role of AI-powered image analysis in enhancing diagnostic accuracy, improving workflow efficiency, and enabling early disease detection, demonstrating its transformative impact on clinical decision-making. Similarly, AI-driven imaging technologies are also revolutionizing point-of-care diagnostics. Integrating AI with imaging technologies at the point-of-care has significantly transformed diagnostics and treatment within healthcare environments, especially in scenarios where rapid decision-making is essential, such as emergency rooms, rural clinics, and field medical assessments146. In recent years, point-of-care applications in microbiology and pathology, among others, have been rapidly expanding, benefiting from the integration of AI tools147. The primary applications of AI in imaging at the point-of-care include shortening of the diagnostic time, image quality enhancement, and increasing the accuracy and portability of the imaging hardware. These advancements not only make imaging technologies more accessible to underserved populations but also reduce the cost associated with conventional benchtop equipment31. Some of the specific examples of imaging-based point-of-care sensors are outlined in Table 4.
Bacteria detection
In a recent study, Wang et al. proposed a deep learning-based imaging system for rapid detection of bacteria colonies, enabling real-time monitoring and early detection of bacteria species148. This computational microscopy system periodically captured holographic images of bacteria growth in a petri dish and analyzed these time-lapsed holograms using CNN models, accurately identifying early-stage bacteria colonies (Fig. 5a). In a proof-of-concept demonstration, the device detected three bacteria species, namely E. coli, K. aerogenes, and K. pneumoniae, achieving >95% sensitivity in identifying bacteria colonies and shortening the detection time by >12 h compared to standard Environmental Protection Agency (EPA)-approved methods. In addition, the proposed platform showed a LoD of approximately 1 colony forming unit (CFU)/L within only 9 h of testing, making it an appealing tool for microbiology research and related sensing applications.

a Image of the holographic microscopy system for bacteria detection (left), whole agar plate image of mixed E. coli and K. aerogenes colonies (middle), and amplitude and phase images of the individual growing colonies (right). b Image of the holographic system for PFU imaging (left), whole plate comparison between the stain-free viral plaque assay after 15 h and the traditional plaque assay after 48 h (right). c Image of the field-portable lens-free imaging flow cytometer (left), which can be used for water quality monitoring and to detect parasites in bodily fluids; the reconstructed images of different microplankton species captured using this portable imaging cytometer (right). d Image of the smartphone-based fluorescent microscope with the disposable sample cassette (top), and schematic illustration of the emission and excitation paths (bottom). e Schematic illustration of the Sight OLO hematology analyzer (top), and false-colored micrographs of different anomalous cell types and formations captured by OLO, red channel: hemoglobin absorption; green channel: nuclear DNA fluorescence; blue channel: cytoplasmic staining (bottom). f Schematic illustration of the miniaturized microscope for automated blood analysis (top), and AI-driven quantification pipeline for FWD, RBC, and MCH count (bottom). a This is adopted with permission from ref. 148 by LSA; b This is adapted from ref. 23 with permission from Nature BME; c This is adapted with permission from ref. 155 by LSA; d This is adapted with permission from ref. 156 by Lab on a Chip; e This is adapted with permission from ref. 157 by Wiley; f This is adapted with permission from ref. 158 by Analyst.
Virus detection
Furthermore, a compact label-free live plaque assay was developed to provide a significantly faster plaque-forming unit (PFU) detection for the quantification of viruses23. A compact lens-free holographic imaging system reconstructed phase images of the target PFUs during the incubation period. The imager scanned the entire area of a six-well plate every hour and utilized a DenseNet neural network-based classifier to convert the phase images of the samples into PFU probability maps, identifying the locations and sizes of the PFUs within the well plate. This stain-free method was capable of automatically detecting the first cell-lysing events caused by vesicular stomatitis virus (VSV) replication as early as 5 h after incubation and achieved a PFU detection rate exceeding 90% in <20 h, providing significant time savings compared to traditional plaque assays, which typically require over 48 h for testing (Fig. 5b).
Parasite imaging
Point-of-care imaging applications have also been developed for diagnosing significant tropical diseases such as schistosomiasis and malaria, which require precise diagnosis typically through microscopy techniques149. The diagnosis of schistosomiasis relies on bright-field microscopy to identify Schistosoma haematobium (S. haematobium) eggs in urine samples, where the operator’s skill is crucial, especially for mild infections. Similarly, malaria diagnosis involves imaging parasites in blood. Automated miniaturized digital microscopes such as the Schistoscope have been developed to identify S. haematobium eggs in real-life biological samples. In recent work, the Schistoscope was applied to automatically detect and quantify S. haematobium eggs in urine samples using ML-based algorithms, achieving over 80% sensitivity150. Furthermore, the EasyScanGo system was utilized for the detection of malaria and employed a CNN-based algorithm for the detection of malaria parasites in blood smears, achieving 91.1% sensitivity and 75.6% specificity, thereby matching the accuracy of experienced microscopists151. These AI-enhanced tools can be applied for automated screening of tropical diseases in underserved populations and low-income countries that lack highly qualified personnel to perform manual diagnosis.
Another platform for detecting parasites in bodily fluids was developed by Zhang et al.152 Unlike traditional methods, this technique exploited the movement of self-propelling parasites as a natural biomarker and contrast mechanism. The sample was illuminated with a coherent light source, and a CMOS image sensor placed underneath the sample captured the time-lapse holographic speckle patterns. These recorded patterns were analyzed using a custom computational motion analysis (CMA) algorithm, which employed holography to create 3D contrast map highlighting the parasites’ movements in the sample. A deep learning-based classifier based on a CNN model was then used to detect and count the parasites from the reconstructed 3D locomotion map. The proposed platform was applied for the detection of trypanosome parasites and showed a strong correlation between detected and spiked parasite concentrations with a LoD of only 3 parasites per 1 mL of biological fluid.
Waterborne parasites, including Giardia lamblia, affect 200 million people yearly, causing diarrheal illnesses such as Giardiasis153. A study by Göröcs et al. introduced a portable, label-free imaging flow cytometer to detect and enumerate Giardia lamblia cysts in real-time in liquid samples154. This device uses lens-free on-chip holographic microscopy to analyze continuously flowing samples at a throughput of 100 mL/h. As the samples flow through the channel, holograms are captured and reconstructed in real-time on a laptop, providing color intensity and phase images. These images are then automatically processed by a CNN model, which digitally sorts and counts the images containing Giardia lamblia cysts. This field-portable imaging flow cytometer can detect less than 10 cysts per 50 mL of sample and can be applied for water quality monitoring in low-resource settings and to detect parasites in bodily fluids. The same imaging flow cytometer was further used for the identification of different plankton types in ocean water. (Fig. 5c)155. Additionally, Koydemir et al. developed a mobile phone-based fluorescence microscopy system using a custom ML algorithm based on bootstrap aggregating to detect Giardia lamblia cysts156. This system consisted of a smartphone coupled with an opto-mechanical attachment weighing only 205 grams (Fig. 5d). It utilized a hand-held fluorescence microscope aligned with the smartphone’s camera unit to image custom-designed disposable water sample cassettes. This mobile phone-based microscopy technique had a LoD of 12 cysts per 50 mL of water sample and showed ~84% sensitivity and >76% specificity for the detection of Giardia cysts.
Hematology analysis
Some AI-based blood sample imaging techniques for point-of-care settings involve coupling miniaturized microscopes with benchtop computers for sample analysis. For instance, complete blood counting (CBC) faces challenges in differentiating between cell sizes and morphologies due to variations in cell maturity. To address this issue, Sight OLO (Sight Diagnostics, Israel) developed a compact microscopy system for hematology in point-of-care settings, providing differentiated five-part CBC through computer vision and AI. This machine-vision technology differentiates cells by extracting their unique peculiarities through CNN models (Fig. 5e)157. The platform was compared with a standard Sysmex XN hematology analyzer and demonstrated strong concordance with the ground truth measurements (Pearson’s r > 0.95) for all major CBC parameters. Another challenge in the field of blood analysis is identifying abnormal erythrocytes, platelets, and leukocytes in the blood smear. For this reason, Chen et al. introduced a label-free contrast-enhanced defocusing imaging (CEDI) and machine vision for instant and on-site diagnostic applications158. The proposed platform identified abnormal blood components through CNN-based algorithms applied to both bright-field and fluorescence microscopy images of the smears (Fig. 5f).
Several blood analysis platforms powered by deep learning were developed using flow cytometry techniques159,160. For instance, Zhang et al. utilized a microfluidic imaging flow cytometer coupled to a CNN to detect COVID-19 by analyzing platelet formation in patient blood samples159. The flow cytometer continuously captured bright-field images of blood cell suspensions, and the CNN further classified captured samples between single platelets, platelet aggregates, and white blood cells. Subsequently, a random forest model extracted cell features from the images and a linear discriminant analysis-based classifier used these features to distinguish between COVID-19-related and non-COVID-19 thrombosis, achieving an accuracy of 75%.
Furthermore, a UNet-based imaging platform for blood analysis was presented by de Haan et al. for automated screening of sickle cells in blood smears using a smartphone-based microscope161. This system comprises two distinct but complementary deep neural networks. The first network enhances and standardizes the blood smear images taken with the smartphone microscope to match the spatial and spectral quality of laboratory-grade microscope images. The second network then uses the enhanced images to perform semantic segmentation, differentiating between healthy and sickle cells in the blood smear. This image processing pipeline takes <7 s per blood smear slide and allows for accurate detection of sickle cells within the sample, achieving ~98% accuracy for the diagnosis of sickle cell disease.
Antimicrobial susceptibility testing
A few AI-driven imaging systems have been proposed for antimicrobial susceptibility testing (AST), which quantifies the efficiency of antibiotics against bacterial infections in patients162,163. AST procedure relies on turbidity measurements of individual wells within a 96-well microplate, often requiring expensive hardware with specialized objectives and mechanical scanning components to capture the entire microplate, limiting applications of AST in resource-limited settings. Recently, fiber optics-based platforms have been proposed as a more cost-effective alternative. These systems utilize LED modules to illuminate the entire well plate and further employ optical fiber bundles to couple transmitted light from the wells to a camera, reducing both the footprint and cost of the device. For instance, Feng et al. proposed a smartphone-based microplate reader that uses a fiber bundle to deliver transmitted light from individual wells to a smartphone camera162. In this work, an ML-based algorithm was applied to determine the optimal threshold for well turbidity detection and to conduct AST in a blind-testing manner. The reader was tested for AST against Klebsiella pneumonia bacteria, achieving an average well turbidity detection accuracy of 98.21% and a minimum inhibitory concentration accuracy (MIC) of 95.12% when blindly tested on 39 patient isolates. In addition, Brown et al. developed a compact fiber-based system for accelerated AST, reducing the incubation time by at least 8 h compared to the gold standard manual inspection method163. The well plate was periodically illuminated by an LED array with a 15-min time interval and a fiber bundle coupled the light transmitted by individual wells to a compact camera, connected to a Raspberry Pi computer. Each well was processed by 21 fibers, capturing the spatial distribution of turbidity within the well, which provided more accurate turbidity estimations compared to configurations with a single fiber per well162. FCNN models processed signals from all 21 fibers to detect antibiotic response to Staphylococcus aureus bacteria over time, achieving 90% detection accuracy after 7 h and 95% accuracy after 10.5 h, considerably faster than the gold standard method, which typically requires 18–24 h. In another application, an automated fiber optics-based microplate reader was applied to accelerate the detection of bacterial growth in water164. This reader combined fluorescent modality for E. coli with colorimetric modality for total coliform and used a Raspberry Pi camera to periodically capture fluorescent and colorimetric images of a 40-well microplate. Bacterial growth was detected by monitoring the intensity increase over time, based on an empirically optimized threshold, enabling the detection of E. coli and total coliform within <16 h, 8 h faster than the conventional approach, with a sensitivity of 1 CFU per 100 mL.
Performance benchmarking of ML models in POCT
Studies to date demonstrate that integrating ML with POCT devices can substantially improve diagnostic performance, but the optimal choice of the ML algorithm to be used often depends on the dataset size, number of features, and task complexity (Supplementary Table S1 in Supplementary Information). When datasets are small, and the feature space is limited, traditional ML models, including random forests, SVMs, and logistic regression, often remain the preferred choice due to their simplicity, faster training times, and reduced risk of overfitting. For instance, Kim et al. demonstrated that random forests outperformed neural networks in classifying prostate cancer using urinary biomarkers in a POCT setting165. However, as the number of input features increased, neural networks surpassed random forests in accuracy, highlighting their superior ability to model complex, nonlinear relationships. This trend highlights a key principle: while traditional models perform well in simple, low-dimensional applications, deep learning models become more effective as data complexity increases, especially in assays with multiple biomarkers or complex feature interactions.
The relationship between feature complexity and ML model performance also applies to image-based POCT, where data are inherently high-dimensional. In these applications, each pixel can serve as an individual input feature, creating complex, nonlinear relationships that can favor deep learning models over traditional ML approaches. Davis et al. demonstrated that CNNs consistently outperformed simpler models, including random forests, in classifying LFAs from smartphone-captured images166. Their study found that while random forests performed well on low-resolution images with fewer features, CNNs excelled in high-resolution datasets, where the increased feature space allowed them to extract more nuanced visual patterns. Additionally, CNNs maintained a strong performance even in the presence of Gaussian noise. This highlights the advantage of CNNs in handling real-world imaging variability, making them a preferred choice for external validation and deployment in POCT applications requiring robustness and generalizability.
The advantages of deep learning models become even more apparent in VFAs, where the 2D sensing membrane provides a more complex spatial distribution of information. This trend is well-illustrated in the study by Han et al. on myocardial infarction diagnosis using a chemiluminescent VFA-based cTnI assay101. While both logistic regression and neural networks achieved nearly identical overall accuracy (95.5% in the case of neural networks and 95.6% in logistic regression), a key distinction emerged in their error distributions. The neural network-based approach had no false negatives compared to logistic regression101, making it clinically preferable, as failing to diagnose a myocardial infarction due to a false negative could have severe consequences for patients. These findings underscore the importance of neural network models in applications where diagnostic sensitivity is crucial, even when traditional models appear to perform at a comparable level. Additionally, random forests had inferior performance compared to neural network models101, suggesting that decision tree-based models may struggle to perform reliable classification near biomarker cut-off values, where subtle variations in signal intensity may have a crucial impact on diagnostics outcomes.
A similar trend can be observed in the work by Eryilmaz et al., which examined ML-based classification of COVID-19 immunity status based on multiplexed serological assays104. Unlike single-biomarker tests such as troponin detection, serological assessments of immune protection involve multiple antibody titers targeting different viral proteins, such as spike and nucleocapsid, as well as different antibody classes, including IgG and IgM. In this study, logistic regression and random forest both demonstrated suboptimal classification accuracies (83.1% and 77.4%, respectively) compared to the performance of neural networks (89.5%). The superior performance of neural networks highlights their ability to capture nonlinear relationships between biomarker levels and immune status, which traditional ML models struggled to represent accurately.
Taken together, these studies and various others in the literature reinforce a key principle in POCT machine learning model selection: when an assay provides only a small number of input features, traditional ML models such as logistic regression and random forest can perform highly competitively. However, as the dimensionality of the assay increases, whether due to multiplexed detection, image-based inputs, more complex continuous biomarker gradients or test-to-test variations, deep learning models begin to offer clear advantages. Their ability to model nonlinear interactions and extract meaningful features from high-dimensional data allows them to outperform simpler models, particularly in cases where diagnostic sensitivity is critical.
Despite these advancements, ML integration into POCT remains an emerging field, and most existing studies focus on evaluating a single ML algorithm or reporting the best-performing model for their specific test rather than conducting direct comparative analyses across multiple ML techniques. As research in this area continues to expand, a growing number of studies are expected to explore more comprehensive benchmarking methodologies. For instance, meta-analyses comparing ML performance across different POCT modalities, the development of standardized datasets for multi-platform ML evaluation, and domain adaptation studies that assess ML transferability across biomarkers will allow for a more rigorous assessment of ML models in POCT. These efforts will further refine best practices for ML model selection in POCT applications, ultimately improving diagnostic accuracy, robustness, and real-world utility.




