ChronoGauge overview
ChronoGauge is a supervised ensemble regressor developed for plant RNA-seq data that can estimate the CT of a sample based on an input vector representing transcripts-per-kilobase-million (TPM) normalized gene expression. The model includes 100 neural-network (NN) sub-predictors that are each fit to a unique set of iteratively selected gene features, with CT estimates across sub-predictors being aggregated using a circular mean. Each NN sub-predictor is a relatively simple multi-layer-perceptron developed from a proof-of-concept model17, which outputs the sine and cosine of the CT. Using a custom angular (θ) loss function, the model is able to more appropriately converge across the 24-h (1440 min) modulus compared with conventional loss functions like mean-absolute-error (MAE) (Supplementary Fig. 1). The bagging-like ensemble improves the robustness of the model across unseen data, whilst also providing a “circadian fingerprint” representing variation between sub-predictors within specific genotypes and experimental conditions. ChronoGauge was trained using Arabidopsis RNA-seq data, as this species has the largest number of suitable datasets for training and evaluating the model.
Generating the ensemble of sub-predictors requires an RNA-seq expression matrix with high time-point resolution for training. Including training samples from multiple experiments is also critical, as this reduces the extent to which ChronoGauge becomes overfit to specific datasets. In our analysis, we combined 4 separate time-course datasets from Arabidopsis (Supplementary Data 1). This included experiments under both LL conditions18,19 (measured in CT) and LD cycles19,20 (measured in ZT), giving a total of N = 56 samples. Replicates were averaged to ensure a balanced number of LL and LD samples (both are N = 28 after averaging). We note that these datasets included experiments entrained under LD cycles, including neutral-day (12:12) and long-day (16:8), but not short-day (8:16). These datasets were selected for training as they all included time-pointed samples that were derived from wild-type (WT) Col-0 whole seedlings harvested under relatively standard experimental conditions. We considered including a further LL time-course (Takeoka et al.21; N = 35) within the training set, but decided to exclude it after we found it had a significant batch effect with less robust rhythmic gene expression compared with other datasets (Supplementary Fig. 2). Time-pointed transcriptome experiments with contrasting genotypes, conditions or tissue types were not included in the training set, as we wanted to ensure these data were unseen when using ChronoGauge to generate hypotheses. Microarray datasets were not included for model training, as they are generally more prone to noise and have a smaller potential gene feature space compared with RNA-seq data.
We hypothesized that a feature set composed of highly rhythmic genes with diverse phases of expression would make intuitive biomarkers for CT prediction. However, the inclusion of LD samples in the training dataset meant that there was a risk of ChronoGauge being fit to gene features that were regulated by the LD cycle, but were not circadian-regulated. To reduce this risk, we applied the R package MetaCycle22 using only the LL dataset by Romanowski et al.18 to determine each gene’s circadian rhythmicity Q-value and phase based on the meta2d approximation (described in Supplementary Data 2). Only genes with significant circadian rhythmicity (Q < 0.05) were used to inform feature selection (N genes = 13,474) (Fig. 1a). The Romanowski et al. dataset was specifically used here as it had the largest number of rhythmic genes compared with other LL time-course datasets (Supplementary Fig. 2b), thus we assumed it was most informative.
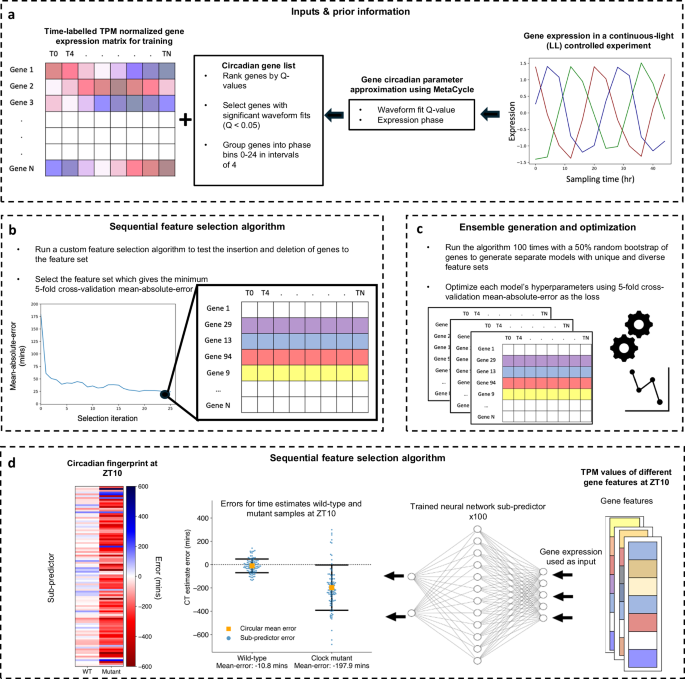
a Data input and prior information: ChronoGauge requires a gene expression matrix including observations from multiple time-course RNA-seq experiments for training and prior information regarding the circadian rhythmicity and phase of each gene’s expression across a time-course harvested under continuous light (LL) following entrainment under a light-dark (LD) cycle as determined by MetaCycle. Genes with non-significant circadian rhythmicity (meta2d Q \(\ge\) 0.05) are filtered out. Genes are grouped into 6 phase bins ranging 0–24 h in intervals of 4. b Feature selection: A custom sequential feature selection (SFS) algorithm involving both forward- and reverse-steps was used to iteratively build a gene feature set with diverse waveform phases, with the fivefold cross—validation (CV) mean-absolute-error (MAE) as a cost. The feature set giving the minimum MAE is selected from each run. c Ensemble generation and optimization: The SFS algorithm is run 100 times with a random 50% bootstrap of genes, giving unique feature sets that are used to tune and train 100 different sub-predictor neural-networks (NNs). d Single time point CT estimation: Application of trained sub-predictors to single time-pointed test samples giving 100 different circadian time (CT) estimates that are combined into a single estimate using circular mean aggregation. The errors of CT estimates can be compared across different samples, e.g., between wild-type plants and clock mutants28. Within each sample, a circadian fingerprint represents variation in errors across sub-predictors fit to different gene features. Swarmplot properties include: central orange box = mean, whiskers = standard-deviation, blue points = errors for individual sub-predictors.
Gene feature sets for each of ChronoGauge’s sub-predictors are generated using a custom sequential feature selection (SFS) wrapper method around the proposed NN architecture (Fig. 1b). This was developed to iteratively build unique sets of gene features with diverse expression phases using semi-randomised parameters. A shortlist of 150 genes was first selected using the top 25 rhythmic candidates across balanced phase groups (binned every 4 h). The gene sets are then constituted through a combined forward- and reverse selection approach using the fivefold cross-validation MAE of CT estimates as a cost function. The algorithm theoretically stops when the number of genes in the feature set reaches 40, but in practice, this value will not be reached in a scalable time-frame. We therefore only ran the algorithm within a specified duration of 6 computational hours, after which we identified the top-performing feature set across iterations. The ensemble is established by running the SFS multiple times, with a bootstrap first being applied to select 50% of genes at random from the feature space. This promotes feature set diversity across the ensemble (Fig. 1c). The optimal feature sets from each run were used to tune and train sub-predictors independently using fivefold cross-validation across the same training data. The trained sub-predictors are able to be applied to test data points, from which the CT estimates can be aggregated for each sample using their circular mean, or be analyzed as a “circadian fingerprint” (Fig. 1d).
Evaluation of ChronoGauge ensemble’s predictive accuracy
The SFS algorithm was run 100 times, giving 100 unique feature sets. Each set was used to train an individual sub-predictor within ChronoGauge’s ensemble. The limit of 100 sub-predictors was set because the cumulative number of unique genes across new feature sets appeared to plateau by 100 SFS runs (Fig. 2a). As part of model selection, we additionally evaluated two other NN-models using more biologically intuitive gene features that might be expected to be related to the circadian clock. This included a NN-model trained using 17 canonical plant circadian clock genes (genes were selected based on the fact they were used previously for circadian analyses23 and were expressed in all training datasets, Supplementary Table 1), analogous to the mammalian model BIO_CLOCK10, in addition to a NN-model fit to the top 500 rhythmic genes ranked by meta2d Q-values. The ensemble of 100 sub-predictors was ultimately selected as the optimal NN model for ChronoGauge, because it gave the lowest MAE across fivefold cross-validation (Supplementary Fig. 3).
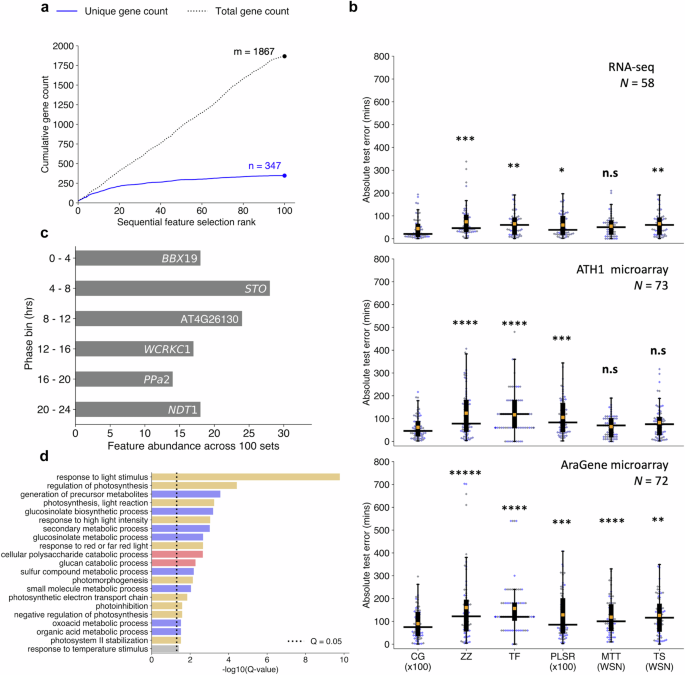
a Cumulative count of unique and non-unique genes present across sequentially selected feature sets ranked by fivefold cross-validation mean-absolute-error (MAE) of circadian time (CT) estimates. b Comparison of hold-out test set absolute errors between the ChronoGauge ensemble (CG ×100) and competing CT prediction methods including ZeitZeiger9 (ZZ), Taufisher14 (TF), partial-least-squares-regression ensemble11 (PLSR ×100), MolecularTimetable8 (MTT), and TimeSignatR12 (TS). Comparison included for an RNA-seq test set24,25,26,27,28,29 (N = 58), an ATH1 microarray set30,31,32,33 (N = 73), and an AraGene microarray set34 (N = 72). Bonferroni adjusted P-values shown from a one-tailed Wilcoxon signed-rank test. Methods using within-study-normalization (WSN) listed. Box-plot properties include: centre-line = median-absolute-error (MdAE), box limits = interquartile range (IQR), whiskers = 1.5 × IQR, orange box = mean-absolute-error (MAE), blue points = error of individual samples. c Top genes belonging to each phase bin based on the number of counts across the 100 feature sets within the ChronoGauge ensemble. d Significantly enriched biological process terms across the 347 unique selected gene features determined in TopGO38 by Benjamni–Hochberg corrected Fisher’s test using all circadian-regulated genes as a background. Processes include those related to light response/photosynthesis (orange), pest defense (blue), starch degradation (red), and temperature response (gray). n.s P ≥ 0.05, * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001, ***** P < 0.00001. Figure source data and actual P-values are provided as a Source Data file.
We benchmarked the ChronoGauge ensemble across 3 unseen transcriptome datasets composed of WT and control Arabidopsis samples (including biological replicates) from multiple studies (Supplementary Data 3) to evaluate whether ChronoGauge’s use was justified. Samples in these datasets were derived from either whole seedlings or aerial (including either leaf or shoot) tissue within an RNA-seq set24,25,26,27,28,29 (N = 58), an ATH1-based microarray set30,31,32,33 (N = 73), and an AraGene-based microarray set34 (N = 72). While ChronoGauge was developed for RNA-seq data, we felt the inclusion of microarray data would enable an expanded evaluation of its generalizability and cross-platform application. We standardized the microarray datasets independently from the training RNA-seq using z-score scaling to allow the expression values to align across platforms. ChronoGauge was compared with other single-sample CT predictors, including ZeitZeiger9, PLSR11, and Taufisher14 as well as predictors that rely on within-study-normalization, including MolecularTimetable8 and TimeSignatutR12. Hyperparameters for these models (Supplementary Table 2) were tuned using a grid-search approach (Supplementary Fig. 4). PLSR was developed as a bagging-like ensemble of 100 sub-predictors using the same SFS algorithm described for ChronoGauge, since it was non-trivial to integrate this specific model within the wrapper. We did not include TimeTeller13 because our training data was not suitable for input within this model.
All models gave CT estimates that were highly correlated (Pearson’s correlation coefficient, r > 0.9) with the true sampling times (Supplementary Figs. 5–7). However, ChronoGauge gave the smallest median-absolute-errors (MdAEs) across each set at 20.6 (+/−47.6) min in the RNA-seq set, 46.1 (+/−50.2) min in the ATH1 set, and 74.8 (+/−68.0) min in the AraGene set (Table 1). These MdAEs were significantly smaller than the other models tested (one-tailed Wilcoxon signed-rank test, Adj. P < 0.05) in all cases except against MolecularTimetable in both the RNA-seq and ATH1 microarray set, and TimeSignatR in the ATH1 set (Fig. 2b), thus demonstrating ChronoGauge’s general advantage in accuracy over other models across these hold-out test samples. MolecularTimetable was consistently placed in the top 3 models based on MdAE, which was unexpected considering it a mechanistic model that is substantially less complex than any of the ML methods. MolecularTimetable is limited however in that it requires the within-study-normalization of expression, which demands at least 2-timepoints ~12 h apart. This means it may not be suitable for predicting the CT within single time-point experiments.
In addition to using unadjusted expression data, we also tested each of the models after they were trained using RNA-seq expression data that had been adjusted by Combat-seq35 to remove batch effects between experimental groups (Supplementary Tables 3–5). While intuitively, we would expect the removal of batch effects to improve accuracy by reducing technical derived variation, we found that ChronoGauge gave marginally (and non-significantly) smaller MdAEs when trained using unadjusted expression values for all test sets (Supplementary Figs. 8 and 9). The accuracy of ZeitZeiger, PLSR, and TimeSignaturR did give smaller MdAEs for RNA-seq data following batch correction, though none were competitive against ChronoGauge when using either adjusted or non-adjusted expression. Given that Combat-seq requires a laborious process of re-integrating different datasets and produced no performance advantage, we felt justified in using ChronoGauge with unadjusted gene expression as an input.
Our training data did not include a time-course experiment entrained to short-day conditions (only neutral- or long-day), thus it was unclear whether ChronoGauge could accurately estimate the CT with said short-day entrainment. Looking at model performance within each experiment (Supplementary Table 6), we do see that experiments entrained and harvested under short-day conditions had higher MdAEs (though not unacceptable, using 120 min as a threshold) compared with all other conditions. This suggests overfitting has occurred, which on one hand may highlight differences in clock dynamics between plants entrained under short-day and neutral-/long-day photoperiods that have been observed previously36. On the other hand, it also suggests ChronoGauge may be less reliable when applied to samples harvested under short-day photoperiods. Despite the increased error, the fact that CT estimates are not random in the short-day experiments indicates that there is still some overlap in circadian gene expression compared with other experiments.
Interpretability of ChronoGauge ensemble
Understanding which gene features are most contributory towards model prediction is useful both to validate that ChronoGauge is appropriately capturing circadian variation, and to potentially uncover mechanistic insights within circadian gene regulation. However, model interpretation using conventional feature importance methods such as SHAP37 are less applicable for ChronoGauge with its ensemble of 100 sub-predictors. As ChronoGauge’s SFS wrapper is discriminative in selecting genes, we instead explored the frequency in which genes were selected across the 100 runs within each phase bin. Except for BBX19, the most commonly selected genes in each phase bin did not include clock-associated components (Fig. 2c). Other clock genes, including PRR7 and RVE8, were included in the top 5 genes of their respective phase bin (Supplementary Fig. 10). The top genes per phase bin displayed relatively consistent sinusoidal expression patterns across the training datasets with no obvious phase shifts across LL and LD experiments (Supplementary Fig. 11). While genes that were selected more often in SFS appear to be strong and logical predictors for inferring the CT, it is unclear whether they themselves give any insight into the clock mechanism or function.
To explore which biological pathways were enriched across all of ChronoGauge’s 347 selected unique gene features from the 100 SFS runs, we performed GO-term analysis using all genes determined to be circadian-regulated (meta2d Q-value < 0.05) as a background (Fig. 2d). Since ChronoGauge’s features tend to include highly rhythmic and robust genes, we believe significantly enriched processes (TopGO38 Fisher’s test, Q < 0.05) may represent pathways that are tightly regulated by the circadian clock. This included those related to light stimulus response and photosynthesis, which are pathways that are known to be particularly robust and conserved23. Multiple enriched processes were related to the synthesis of the defense secondary metabolite glucosinolate. While this defense pathway has previously been associated with the clock through both molecular and physiological observations39, its enrichment suggests a particularly robust circadian regulation. Several processes were also associated with glucan catabolism, which is involved with the starch degradation pathway. Starch degradation is critical for providing energy for plant growth in the absence of light and is clock-regulated40, thus its enrichment among our feature sets supports the suggestion that it is also a robustly regulated pathway.
Irrespective of ChronoGauge’s predictive performance lead over other models, an advantage of the ensemble is an increased ability to assess and dissect circadian function by comparing the gene features of groups of sub-predictors that give different CT estimates. To illustrate this, we demonstrated that there was substantial variation across sub-predictor outputs within each time-point of a 16:8 LD test dataset (Fig. 3a), including some that gave relatively accurate CT estimates and others that yielded advanced or delayed CT estimates. Using ZT10 as an example, we defined three groups of outputs across sub-predictors, including accurate (<60 min absolute error), advanced (> +60 min error) and delayed (< −60 min error), and performed GO term enrichment on each of these (Fig. 3b). We found that gene features within the accurate group were associated with most of the processes described using the full feature set in Fig. 2d. “Circadian rhythm” was the only process significantly enriched exclusively in the delayed group features, which included the clock components PRR7, RVE8, BBX19, and CCA1, as well as others associated with the clock. It is possible that some clock genes are not particularly reliable biomarker genes, as it has previously been shown clock gene expression values can exhibit considerable variation between plants at specific time-points20. This may also explain why our NN-based model using clock genes as features performed sub-optimally in cross-validation compared with the ensemble using multiple gene sets.
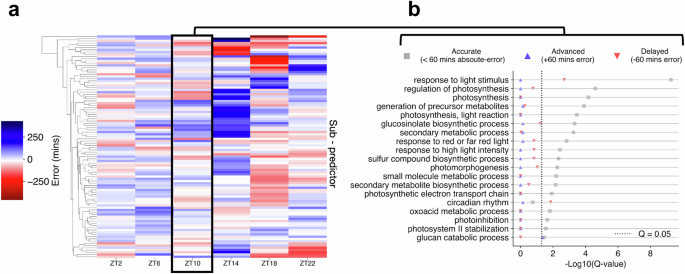
a Comparison of phase estimates made by each sub-predictor within ChronoGauge for the test accession published by Rugnone et al.24. Colour-scale represents the mean-error of phase estimates made for each timepoint. b Enrichment of biological processes terms across different sub-predictor groups in ZT10, including those with accurate (<60 min absolute-error), advanced (+60 min error) and delayed (−60 min error) CT estimates. Term enrichment was determined using Fisher’s exact test in TopGO38 (Q < 0.05) using all circadian-regulated genes as a background. Significance determined by Benjamini–Hochberg adjusted P-values. Figure source data are provided as a Source Data file.
ChronoGauge highlights dysfunction in core clock-associated genes
While ChronoGauge can make accurate CT estimates in wild-type (WT) control samples, we hypothesized that it could also detect dysfunction across the circadian transcriptional network in response to perturbations of clock components. To test this, we made CT estimates on RNA-seq datasets generated by Graf et al.28 (Supplementary Data 4), which included the clock gene mutants toc1-101 and gi-201 in a Col-0 background, and lhy-21/cca1-11 in a Ws-2 background. All samples included rosette tissue and were harvested under a 12:12 LD cycle. We compared the errors of CT estimates for each mutant with corresponding WT samples (Table 2) at both ZT0 (Fig. 4a–c) and ZT12 (Fig. 4d–f). Based on physiological assays of these mutants’ period lengths, we anticipated they would display CT estimates that were advanced or delayed compared with the WT.
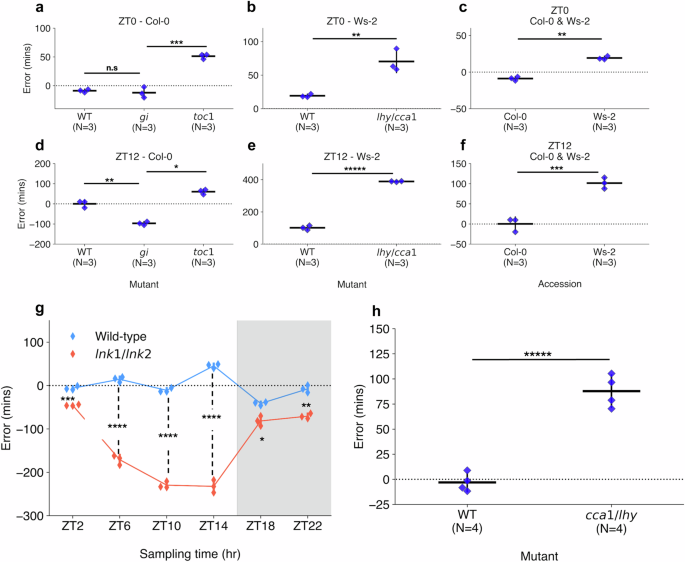
Comparison of errors of circadian time (CT) estimates across clock mutant with wild-type samples (WT) and different accessions. CT estimates made across RNA-seq data generated by Graf et al.28, including those harvested at (a–c) ZT0 and (d–f) ZT12 under a 12:12 light-dark (LD) cycle. Mutants include (a, d) toc1-101 and gi-201 in the Col-0 background and (b, e) lhy-21/cca1-11 in the Ws-2 background. Additionally, (c, f) a comparison was made between Col-0 and Ws-2 accession CT estimates. g CT estimates across RNA-seq samples published by Rugnone et al.24, including comparison of mean CT estimate errors in lnk1/lnk2 mutant (red) and WT (blue) samples harvested across a time-course under a 16:8 LD cycle. White area shows samples harvested under light, while gray area shows samples under dark conditions. h CT estimates across RNA-seq samples generated by Blair et al.50, including comparison of CT estimates in cca1-1/lhy-22 double mutant and WT samples harvested at ZT1 under a 12:12 LD cycle. All strip plot properties include: centre line = mean, whiskers = standard-deviation, blue points = individual samples. All tests performed using two-tailed independent T-test, with Bonferroni P-value adjustment for multiple tests within each experiment. n.s P ≥ 0.05, * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001, ***** P < 0.00001. Figure source data and actual P-values are provided as a Source Data file.
In the Col-0 background (Fig. 4a, d), WT samples gave a mean-error of −8.8 (+/−2.4) min at ZT0 and 0.2 (+/−14.0) min at ZT12, corresponding to a period length of ~24 h (assuming an idealized 24-h clock period). toc1-101 samples had significantly advanced CT estimates compared with the WT in both ZT0 and ZT12 with a mean-error of 51.2 (+/−3.6) min at ZT0 (Two-tailed independent t-test, Adj. P < 0.001) and 60.6 (+/−10.2) min at ZT12 (Adj. P = 0.02) based on a two-tailed independent-samples T-test. These advanced CT estimates correspond with a period length of ~23 h, which would be consistent with the previously observed phenotypes in toc1 mutants, including an earlier phase under LD cycles41 or a shorter period under LL42,43. The gi-201 samples at ZT0 gave a mean-error of −12.1 (+/−7.6) min, with no significant difference found compared with the WT (Adj. P = 1.00). However, at ZT12, gi-201 gave mean-errors of −96.9 (+/−7.2) min, suggesting a significant delay compared with the WT (Adj. P = 0.002). Previous studies have shown that gi mutants can display either long- or short-period phenotypes in LL depending on lighting conditions and the type of measurement taken across the assay44,45. The delayed CT estimate at ZT12 would correspond with a period length of ~25.5 h, indicating that these gi plants have a long-period phenotype. The relatively accurate CT estimate at ZT0 suggests that the entraining LD cycle may realign the clock in gi with the environmental cycle between dusk and dawn, with a misalignment emerging progressively through the photoperiod.
We performed GO term enrichment to find biological processes significantly associated (TopGO38 Fisher’s test, Q < 0.05) with gene features in accurate (absolute error <60 min), delayed (error < −60 min) and advanced (error > 60 min) sub-predictors across the samples under a Col-0 background (Supplementary Fig. 12). In toc1-101, GO-term processes including (but not limited to) circadian rhythm, temperature response, photomorphogenesis and red-light response were enriched in advanced error sub-predictors at both ZT0 and ZT12, suggesting a possible dysregulation to the circadian integration of light and temperature. In gi-201, most processes were significantly enriched in the accurate group at ZT0, suggesting only minor dysregulation. At ZT12 however, some enrichment was found across metabolic processes related to glucosinolate synthesis, suggesting a dysregulation of this pathway at this time-point. In both toc1-101 and gi−201 mutants, the glucan and polysaccharide catabolism processes associated with starch degradation were enriched only in the accurate groups. This suggests starch catabolism is a stably rhythmic diurnal pathway in LD, even under such clock perturbations.
In the Ws-2 background (Fig. 4b, e), WT samples gave a mean-error of 19.2 (+/−2.0) min at ZT0 and 101.5 (+/−11.2) min at ZT12. In the mutant samples, lhy-21/cca1-11 at ZT0 gave a mean-error of 70.4 (+/−13.6) min, which was significantly advanced from the WT (P < 0.01), while ZT12 samples gave a mean-error of 388.9 (+/−2.2) min demonstrating a substantial advance (P < 0.001). Based on the difference at ZT12, we suggest the clock is dysregulated, which is consistent with the damped oscillations observed in previous lhy/cca1 double mutants under LL46. The more moderate difference at ZT0 however, indicates that clock function may be somewhat rescued by the LD cycle between dusk and dawn. Following GO term analysis in lhy-21/cca1-11 samples (Supplementary Fig. 13), gene features under the advanced (error > 90 min) or delayed (error < −90 min) sub-predictors for ZT0 were enriched mainly in pathways related to light response and photosynthesis regulation. At ZT12, most processes described in Fig. 2d were enriched in the advanced error group with few sub-predictors belonging to the accurate group. We therefore suggest there is a widespread dysregulation of gene expression in the lhy-21/cca1-11 genotype under LD cycles, especially at ZT12. This is consistent with the role of both CCA1 and LHY as transcriptional repressors, which target large sets of dusk peaking transcripts47,48. We note the Ws-2 samples were significantly advanced (P < 0.01) from Col-0 at each timepoint (Fig. 4c, f). These advanced estimates likely reflect differences in circadian transcriptional dynamics between the two accessions, since ChronoGauge was trained only using Col-0 and not Ws-2, and also coincides with previous analyses that have shown Ws-2 plants display a marginally shorter period length than Col-0 plants (by ~30 min) under LL49.
We further applied ChronoGauge to samples with other circadian clock mutants. In a 16:8 LD time-course spanning 24 h of aerial tissue samples24, we observed a significant delay in CT estimates (Two-tailed T-test, Adj. P < 0.05) in lnk1-1/lnk2-1 double mutant samples compared with the WT at all time-points, with highly significant delays (Adj. P < 0.0001) between ZT0 and ZT14 (Fig. 4g). A substantial dysregulation of the circadian network would be expected within the lights-on time-points in said mutants, as the LNK genes participate in integrating light stimuli with the circadian clock. In another experiment involving plants entrained to a 12:12 LD cycle50 (including whole seedling tissue), cca1-1/lhy-20 mutant replicates harvested at ZT1 were significantly advanced relative to the WT (P < 0.001) (Fig. 4h).
We also tested ChronoGauge in datasets obtained from plants challenged with different temperature conditions (Supplementary Fig. 14). In one experiment50, Col-0 plants entrained at 22 °C were shifted to 10 °C or 37 °C one hour prior to sampling whole seedling tissue. Compared with control plants (kept at 22 °C), plants shifted to 37 °C gave significantly advanced CT estimates (Two-tailed T-test, Adj. P < 0.0001), while those shifted to 10 °C displayed no significant difference (Adj. P = 1.00). This suggests that while the circadian transcriptome is resilient to sudden drops in temperature, a sudden shift to 37 °C induces significant transcriptional differences, possibly because 37 °C is a stress condition for Arabidopsis. This is supported by the fact that the authors of the experiment reported a larger number of differentially regulated genes between 22 °C and 37 °C samples (N = 3369) compared with 22 °C and 10 °C samples (N = 426). It is difficult to associate these results with findings from independent works however, since few studies have explored the clock’s response to short-term cold or heat stress. A separate experiment27 tested differences in Col-0 plants grown at 22 °C and 27 °C under a short-day (8:16) LD cycle (including whole seedling tissue). CT estimates showed a significantly higher MAE across time-points in 27 °C samples (Two-tailed Wilcoxon signed-rank test, P = 0.03). In previous work, a subtle difference in the period lengths of Col-0 between plants that experienced long-term exposure to 27 °C compared with 22 °C49, which coincides with our findings that exposure to higher temperatures for long durations can impact clock dynamics.
ChronoGauge detects circadian variation in non-model species
The previous analyses show that ChronoGauge can detect circadian clock variation in Arabidopsis thaliana circadian mutants and plants grown under different experimental temperatures. However, we wanted to explore its versatility further by making similar analyses in non-model plant species. Because the number of samples that are suitable for training models specific to other species are minimal, we instead identified putative orthologs of ChronoGauge’s selected Arabidopsis gene features within each non-model species. Genes with no directly mapping orthologs were removed, and each sub-predictor was re-trained. Where multiple orthologous genes mapped to one Arabidopsis gene feature, their expression values were averaged. These steps ensured the feature spaces were identical across species. We tested this approach by making validation CT estimates on RNA-seq datasets, firstly including Arabidopsis halleri samples harvested from the wild51,52 (N = 383), as this dataset represents a closely related species to A. thaliana with the potential for comparison across different seasonal conditions. We also tested CT estimation in more distantly related species that contain known clock gene orthologs with time-courses that were sampled under controlled LL conditions, including Brassica rapa53 (N = 48), Glycine max15 (N = 36), and Triticum aestivum23 (N = 36) (Supplementary Data 5).
Compared to the A. thaliana test data with a MdAE of 20.6 (+/−47.6) min, other species produced less accurate CT estimates. A. halleri field samples gave a MdAE of 137.1 (+/−162.0) min, which could reflect substantial noise in expression as the samples come from natural environments. In other species, B. rapa gave a MdAE of 169.2 (+/−70.3) min, T. aestivum of 141.5 (+/−90.4) min, and G. max of 79.0 (+/−50.1) min (Fig. 5a). It is unclear why G. max displayed such high accuracy compared with other non-model species, especially considering B. rapa is a closer relative to A. thaliana. One explanation could be that like A. thaliana, G. max is a diploid, thus they may have more orthologous genes that uniquely map to one another. In contrast, the paleopolyploid B. rapa and the hexaploid T. aestivum tend to include multiple ortholog mappings to individual genes in A. thaliana. These duplicated ortholog mappings may add noise to ChronoGauge’s gene features, thus reducing its accuracy. We also noted that CT estimates were advanced in B. rapa and delayed in T. aestivum, which could be due to the clock period lengths being shorter and longer respectively in comparison with A. thaliana and G. max. Examining the period lengths of genes determined to be circadian-regulated (meta2d Q < 0.05; Supplementary Fig. 15), we found G. max genes had a significantly longer period length compared with A. thaliana genes (Two-tailed Mann–Whitney U test, Adj. P < 0.00001), with no significant difference between B. rapa and A. thaliana gene period lengths (Adj. P = 0.94). Thus, these transcriptome datasets suggest that ChronoGauge could not capture period length differences between these species. All CT estimates for non-model species displayed a high correlation (r > 0.85) with the true sampling times (Supplementary Fig. 16). The fact ChronoGauge was able to non-randomly estimate the CT across species as divergent as a monocot (T. aestivum) after training the model using expression data from the dicot A. thaliana suggests a high conservation in the circadian transcriptome that can be exploited to explore CT variation in species that lack appropriate training datasets.
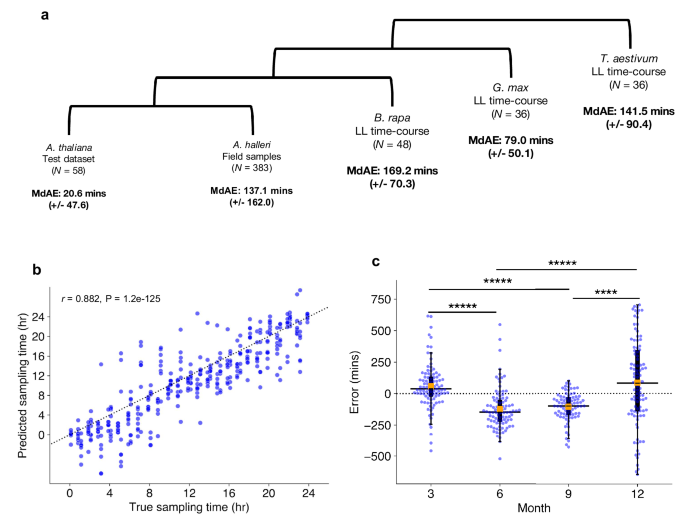
a Circadian time (CT) estimate mean-absolute-errors (MAEs) across test datasets in different species, the test dataset for Arabidopsis thalina, field samples for Arabidopsis halleri51,52, a continuous light (LL) time-course for Brassica rapa53, a LL time-course for Glycine max15 (soybean), and another LL time-course for Triticum aestivum23 (wheat). b Correlation between estimated sampling times/CTs with actual sampling times in A. halleri field data (N = 383) tested using Pearson’s correlation coefficient. Sampling time labels were adjusted to set dawn to ZT0 based on Tokyo time zone information. c Comparison of median CT estimate errors in samples with weather meta-data (N = 367) across different seasons, including Spring (3), Summer (6), Autumn (9), and Winter (12). Box-plot properties include: centre line = median, box limits = interquartile range (IQR), whiskers = 1.5 × IQR, orange box = mean, blue points = individual samples. Significant differences between group means determined using a two-tailed Man–Whitney U test with Bonferroni adjustment of P-values. **** Adj. P-value < 0.0001, ***** Adj. P-value < 0.00001. Figure source data and actual P-values are provided as a Source Data file.
The A. halleri samples51,52 were harvested from a natural plant population in Hyogo Prefecture, Japan, across different time-points of the day throughout the months of March (3), June (6), September (9), and December (12). There was a strong correlation between the true sampling time (ZT hr) and the predicted CT estimates (r = 0.88) (Fig. 5b), though the samples tended to give more accurate CT estimates around ZT12, which roughly corresponds to dusk. This may indicate a stronger synchronization of the circadian transcriptome at dusk transitions. ChronoGauge is fit to training data entrained under square-wave LD cycles, while the A. halleri field samples would have experienced a photoperiod gradient from day to night. Thus, the higher error outside of dusk might also be explained by mismatches in the shape of transcript waveforms between controlled versus natural light-dark conditions.
We wished to evaluate whether ChronoGauge could identify differences between the circadian transcriptomes of the A. halleri samples that were caused by specific seasonal environmental factors, such as changes in the photoperiod length and ambient temperature. For samples with associated weather meta-data (N = 367), we made CT estimates to compare errors across seasons (Fig. 5c and Table 3). We found that plants harvested in June (6) and September (9) (giving median errors of −147.7 (+/−164.9) and −99.8 (+/−110.6) min respectively) were significantly delayed (Two-tailed Mann–Whitney U test, Adj. P < 0.0001) compared to March (3) and December (12) (with a median errors of 38.8 (+/−180.3) and 81.4 (+/−342.1) min, respectively). On an absolute scale, CT estimates for winter samples gave the largest MdAE at 221.9 (+/−206.3) min (Supplementary Fig. 17), which was highly significantly greater (Adj. P < 0.001) than all other seasons, suggesting a less robust circadian regulation at this season. To test whether these differences might be explained by specific winter conditions, we associated sample errors with their environmental metadata including photoperiod hours (sunset–sunrise), air temperature, precipitation and wind-speed (Supplementary Fig. 18). Air temperature and photoperiod, individually, displayed modest negative correlations with the CT errors (Pearson’s correlation coefficient, r < −0.30), however, when adjusted for confounding variables using multivariate ordinary-least-squares regression, only air temperature was found to be significantly associated with error (multivariate P = 0.0009), while photoperiod was not (multivariate P = 0.13). Wind-speed and precipitation were not associated with CT error based either on individual correlation (−0.05 > r < 0.05) or when adjusted for multiple variables (multivariate P > 0.25). ChronoGauge thus suggests temperature may play a leading role in the regulation of the circadian clock across natural conditions and seasons, which is consistent with previous statistical analyses of these A. halleri field data51.
ChronoGauge predictions highlight potential genetic marker-trait-associations
Natural variation in circadian rhythms has previously been observed across Arabidopsis accessions using leaf movement assays49, luciferase imaging54, and delayed-fluorescence55. Using ChronoGauge, we generated CT estimations from RNA-seq samples (rosette tissue) of 159 Swedish Arabidopsis accessions56 to investigate variation in clock function (Fig. 6a). Each accession was grown at both 10 °C and 16 °C under 12:12 LD before being harvested for RNA-seq between ZT11 and ZT12. ChronoGauge gave a mean CT of 9.4 h (+/−1.5) in the 10 °C group and 9.7 h (+/−1.3) at 16 °C with a difference that was statistically modest (Paired samples t-test, P = 0.07). In comparison with previous phase approximations using delayed-fluorescence across a subset of accessions55 (N = 20) (Supplementary Fig. 19), our CT estimates displayed a moderate correlation at 10 °C (Pearson correlation coefficient r = 0.50), but poor correlation at 16 °C (r = 0.07), suggesting ChronoGauge’s transcriptome-based CT estimate is not necessarily consistent with other methods for quantifying clock parameters. The CT estimates for each temperature group were significantly earlier than ZT11 (the closest possible harvesting time based on mean estimates) (1-sample T-test, Adj. P < 0.001). The delayed CT estimates may indicate a long period phenotype for these ecotypes due to the lower growth temperatures compared with the training samples (22–24 °C). This is consistent with previous literature showing long-term exposure to lower temperatures are associated with longer period lengths in many Arabidopsis ecotypes49, though we note comparisons across independent experiments should not be over-interpreted due to the impact of batch effects.
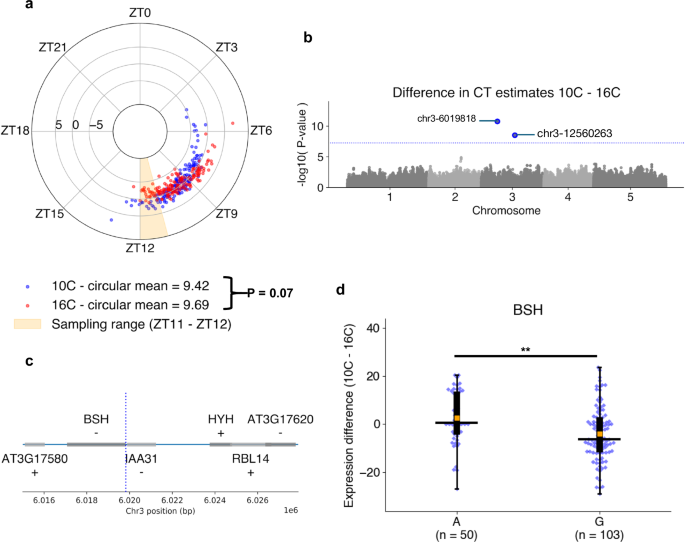
a Circadian time (CT) estimates across 159 Swedish Arabidopsis accessions56 grown under 10 °C (blue) or 16 °C (red) conditions. Polar axis determines ZT within a 24-h period. Radial axis represents the difference between temperature groups (10 °C–16 °C) for each ecotype. Statistical comparison of means between temperature groups performed using a paired T-test. b Associations between differential alleles and the difference between CT estimates 10 °C–16 °C across N = 153 accessions determined by the genome-wide-association-study (GWAS) model BLINK78 in GAPIT77. Significant marker-trait-association (MTA) sites were identified with a threshold of P < 5.68e-08 due to Bonferroni adjustment across N = 880,417 SNP sites. c Genes proximal to the chr3-6019818 A > G MTA (blue line) identified through GWAS. d Comparison of transcript-per-kilobase-million (TPM) normalized expression differences of the BSH gene between 10 °C–16 °C groups in those containing the A and G allele at chr3-6019818. Box-plot properties include: centre-line = median, box limits = interquartile range (IQR), whiskers = 1.5 × IQR, orange box = mean, blue points = individual accessions. Statistical significance determined using two-tailed Mann–Whitney U test with Bonferroni adjusted P-values for all expressed and annotated genes 30 kb up- and down-stream the MTA site. ** Adj. P-value < 0.01. Figure source data and actual P-values are provided as a Source Data file except for Manhattan plots, which are provided within an Open Science Framework (OSF) file79.
We obtained variant allele information for 153 of these ecotypes, which included a total of N = 880,417 variant sites post-processing. We used three derived circadian phenotypes (CT estimates at 10 °C and 16 °C, and a temperature compensation phenotype calculated from the difference in CT estimates between the two temperatures 10 °C–16 °C) to perform a genome-wide-association-study (GWAS) using the BLINK model to identify marker-trait-associations (MTA) for each of the phenotypes. No significant MTAs were identified for the CT estimates themselves (Supplementary Fig. 20), but two significant MTAs were found in Chromosome 3 using the phenotype that was a proxy for temperature compensation of the clock (P < 5.7e-08) (Fig. 6b). For the MTA at chr3-12560263, we found no annotated genes 30 kb up- or downstream. In comparison, the MTA at chr3-6019818 (Fig. 6c) lies within the promoter region of BSH (AT3G17590), a gene that has previously been associated with a “bushy” phenotype57, but that lacks known association to the circadian clock or temperature responses. In gene expression data, the temperature response (TPM 10–16 °C) for BSH (Fig. 6d) was significantly greater (Two-tailed Mann–Whitney-U test, Adj. P = 0.008) across ecotypes with the A allele at chr3-6019818, 2.6 TPM (+/−11.1) compared to those with the G allele at −4.0 TPM (+/−11.2), supporting the notion that this gene is associated with a response to temperature of the Arabidopsis circadian clock. Interestingly, the gene HYH is ~4 kb from the MTA and has previously been shown to be implicated in a sigma factor-mediated cold response pathway58 and regulation of a temperature-responsive miRNA59, making it a second plausible candidate for a temperature response marker. However, we detected no HYH expression in any of the 153 samples, thus were unable to support its association with the temperature response phenotype using these data. Overall, these results demonstrate that ChronoGauge can be used to generate a circadian-related phenotype within a GWAS to identify potentially novel marker genes for clock regulation that are strong candidates for functional validation.




