demographic information
Clinicopathologic information for 348 (64.9%) men and 188 (35.1%) women included in this study was downloaded from TCGA and summarized in Table 1. The average age was 65.3 years, and approximately 250 (46.6%) had advanced gastric cancer.
Correlation analysis
Of the clinicopathological data, only stage was significantly associated with cancer. An important criterion for measuring relevance is p– < 0.05 value (Figure 2A).
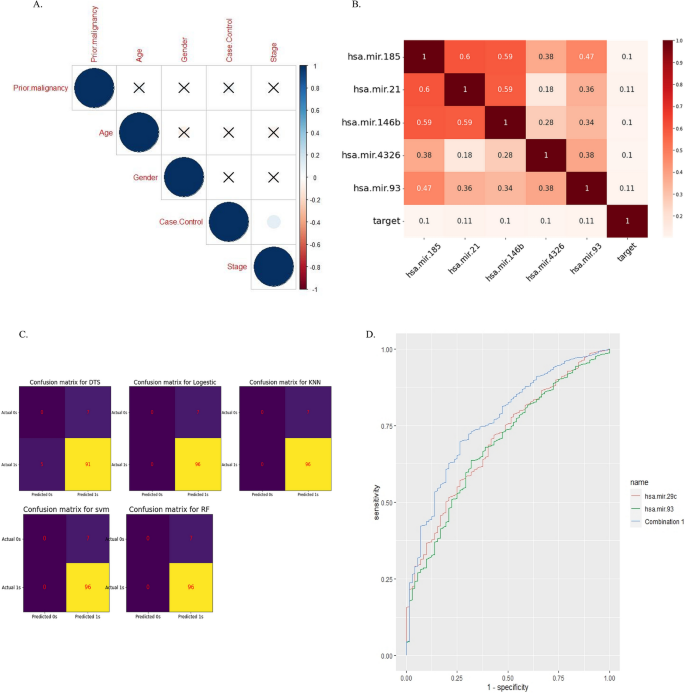
(a) Correlation analysis using the ggcorpot package, R software v 4.2. (B.) Heat map analysis used to show important features. (C.) confusion matrix was used to compare different machine learning algorithms. Figure B&C was plotted using Python v3.7. (D.) ROC curve analysis revealed biomarker potency of miR-29c alone and in combination with miR-93 using the combioROC package in R 4.2.2.
Data collection
As mentioned in the Materials and Methods section, the source of raw data from clinical information and sequencing was the TCGA database. Based on the criteria mentioned, 536 samples were selected for further study, of which approximately 465 were related to GC patients and 72 to age- and sex-matched controls.
Data preprocessing and identification of differentially expressed miRNAs (DeMiRs)
The dataset contained 1882 miRNAs, which was reduced to 220 miRNAs after normalization using the Limma package, R software. In a processing step using heat maps, the most significant features were selected (Figure 2B) and classified using machine learning algorithms. Then 5 algorithms (SVM, dts, rf, logistic regression, and knn) were tested with 4 different metrics (accuracy, f1score, ROC_curve, and confusion matrix), and finally obtained from these 4 metrics According to the scores, the SVM algorithm was chosen as the most accurate algorithm. (DTS, Accuracy: 88%, AUC = 47%; random forest, Accuracy: 93%, AUC = 39.5%; SVMs, Accuracy: 93%, AUC = 88.5%; KNN, Accuracy: 93%, AUC = 41.7%; logistics, Accuracy: 93%, AUC = 88%). The confusion matrix can also be seen in Figure 2C. As a result, a list of 29 miRNAs with 5 significant up and 24 significant down expression in gastric cancer was selected for further analysis (Table 2) Figure 3.

29 miRNAs obtained from logFC-based feature selection-based machine learning.
ROC curve analysis for identification of diagnostic biomarkers
The results of ROC curve analysis were hsa-miR-93 (combined AUC was 0.76, sensitivity was 0.69, specificity was 0.73, cutoff was 0.86) (Figure 2D).
Survival analysis of demiR
Survival analysis of demiR was performed using SPSS version 20, p– Values were considered < 0.05. As a result, 13 miRNAs (Hsa-miR-21, Hsa-miR-146b, Hsa-miR-185, Hsa-miR-1.1, Hsa-miR-1.2, Hsa-miR-143, Hsa-miR-4652, Hsa-miR-1911, Hsa-miR-29c, Hsa-miR-3170, Hsa-miR-139, Hsa-miR-5683, Hsa-miR-133a.2) have prognostic function (Figure 4).

Kaplan Meier visualization of identified prognostic biomarkers using Survival, survminer, and ggplot2 R packages in R software v4.2.2.
Validation of candidate microRNAs in the dataset
Of the 29 candidate microRNAs obtained from the machine learning algorithm using the online web server described in the Materials and Methods section, hsa-miR-21, hsa-miR-133a, hsa-miR-146b, hsa-miR-146b, hsa- miR-29c, and HSA-MIR-204 can be found in (gse54397)、exp004526(gse54397)で高度に検証されていました。 GSE106817)、EXP00405、EXP00118(GSE28700)、EXP00406、EXP00666、EXP00444 (GSE78775)、EXP00476 (GSE99415)、EXP00316 (GSE77380)、および EXP00175(GSE33743 )) was performed using the supplemental file (httpsR miPath heatmap server 1, run online/mpd.bioinf.uni-sb.de/, Figure 5A).
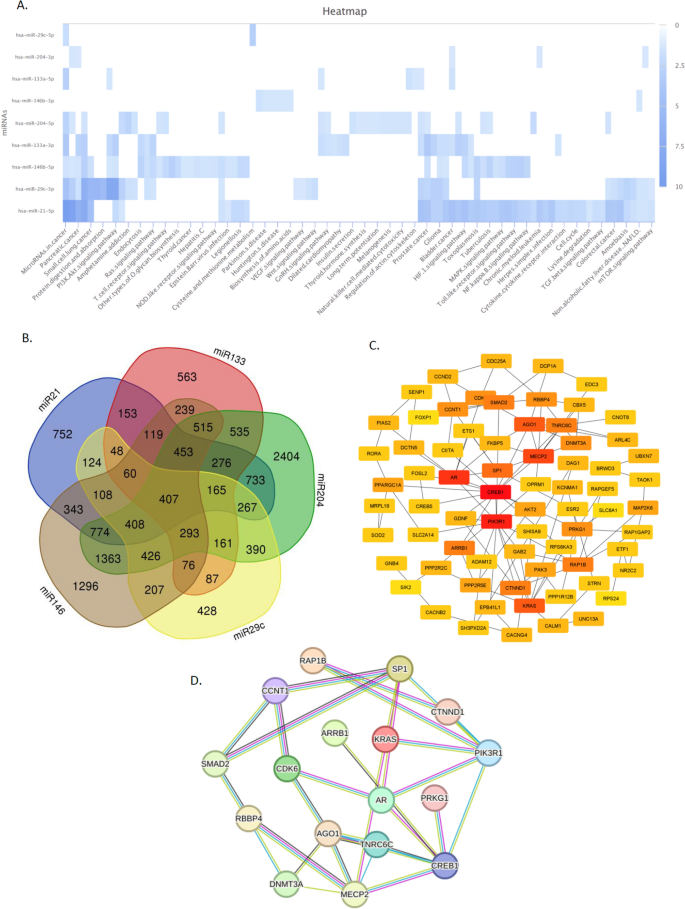
(a) using miRPathDB v2.0 (https://mpd.bioinf.uni-sb.de/heatmap_calculator.html?organism=hsa) to generate a heatmap of highly validated miRNAs and the pathways in which they are involved (B.) candidate miRNAs and their common target genes using the Venn diagram online database (https://bioinformatics.psb.ugent.be/webtools/Venn/). (Ha) We used the Cytohubba tool (https://cytoscape.org/ cytoscape version3.9.1) to select 100 high-scoring genes based on degree (D.) miRNA target genes. The hub gene reparsed by the string database consists of 100 nodes and 223 edges. Set the highest trust score of 0.9 and hide disconnected nodes in the network (https://string-db.org/).
miRNA target prediction
Prediction of miRNA targets was achieved using several databases including miRwalk, miRdb and Targetscan. Using the Venn diagram online database, a list containing 407 common genes was identified (Figure 5B).
Protein-protein interaction network analysis
The candidate genes predicted in the previous step were submitted to the STRING database to build a PPI network based on the criteria described in Materials and methods. To obtain hub genes with important roles, PPI networks were imported and visualized by Cytoscape software. We used the Cytohubba tool to select 100 high-scoring genes based on degree (Fig. 5C). Finally, we imported the hub genes into the string database and re-analyzed the PPI network (Figure 5D).
Functional analysis
To reveal the role of selected hub genes, we performed enrichment analysis using R software. Results demonstrate hub gene transcription factor binding, enzyme binding, RNA polymerase II cis-regulatory region sequence-specific DNA binding, protein binding, double-stranded DNA binding, arrestin family protein binding, sequence-specific DNA binding, and chromatin binding. bottom. In terms of molecular function, most genes are involved in miRNA-mediated inhibition of translation, positive regulation of viral transcription by the host, regulation of gene expression by gene imprinting, production of miRNAs involved in gene silencing by miRNAs, Gene silencing, enriched Wnt signaling pathways, calcium regulatory pathways, regulation of cellular senescence, negative regulation of gene expression, and epigenetic gene silencing on biological processes. Chromatin, euchromatin, nucleoplasm, non-membrane-bound organelles, and cytosol were the most enriched cellular components. Identification of key signaling pathways using the KEGG database indicates that candidate hub genes are primarily associated with glioma, melanoma, prostate cancer, non-small cell lung cancer, renal cell carcinoma, GnRH secretion, aldosterone-regulated sodium reabsorption, and pancreatic It has been shown to be involved in cancer (Fig. 6).

Target gene enhancement analysis based on gene ontology fora) molecular function, (B.) cell component (C.) biological processes. (D.) KEGG pathway analysis. All analyzes were plotted using SRplot http://www.bioinformatics.com.cn/srplot, an online platform for data analysis and visualization.



