We present a novel method for gene discovery that can exploit information hidden in high-dimensional clinical data. Representation Learning for Gene Discovery with Low-Dimensional Embeddings (REGLE) is computationally efficient, does not require disease labels, and can incorporate information from expert-defined knowledge.
Modern healthcare systems generate vast amounts of high-dimensional clinical data (HDCD) that cannot be summarized as a single binary or continuous number (see “has asthma” or “height in centimeters”), such as spirogram measurements, photoplethysmograms (PPG), electrocardiogram (ECG) recordings, CT scans, MRI images, etc. Understanding the relationship between genome and HDCD is essential not only to better understand disease but also to develop treatments for disease.
HDCH is stored in electronic medical records and large-scale biobanking projects such as UK Biobank in the UK, BioBank Japan in Japan, and All of Us in the US, where participants' consent is obtained before anonymizing the data and sharing some of this valuable resource with qualified scientists, with the aim of enhancing the prevention, diagnosis, and treatment of a range of life-threatening diseases.
The Google Research genomics team has made progress using HDCD to characterize diseases and biological traits, such as optic disc morphology and chronic obstructive pulmonary disease (COPD). To better understand the genetic architecture of these specific traits, we previously conducted genome-wide association studies (GWAS) on trait predictions generated by supervised machine learning (ML) models. However, it is not always possible to obtain data with sufficient disease labels to train supervised ML models. Furthermore, simple disease labels cannot fully capture the biology embedded in the underlying data, and we lack statistical methods to directly utilize HDCD in genetic analyses such as GWAS.
To overcome these limitations, the paper “Unsupervised Representation Learning on High-Dimensional Clinical Data Improves Genomic Discovery and Prediction” published in 2011, Nature Genetics, We present a principled method to study the genetic factors underlying common organ functions reflected in HDCD. Representation Learning for Genetic Discovery in Low Dimensional Embeddings (REGLE) is a computationally efficient method that does not require disease labels and can incorporate expert-defined features (EDFs) when available.
Discover hidden information in HDCD
A simple way to explore the relationship between genes and HDCD is to run GWAS on each data coordinate. For example, we can look at the variation of each pixel value in a medical image. This method is computationally expensive, has high correlation between nearby coordinates, and has a huge burden of multiple testing, resulting in low power to discover significant associations. A more widely used approach is to focus on a small number of expert-defined features (EDFs) extracted from HDCD as the target traits or phenotypes for GWAS. EDFs can include clinically known features, such as forced vital capacity (FVC) and forced expiratory volume in 1 second (FEV1) in the case of spirograms, as shown in previous studies. Although these EDFs are important features discovered by experts, we hypothesized that running GWAS on them may not fully exploit the potential of HDCD, as they may not comprehensively capture the signals encoded in HDCD.
REGLE aims to overcome these limitations using a variational autoencoder (VAE) model. The method consists of three main steps: (1) learn a nonlinear, low-dimensional, separable representation (i.e., encoding or embedding) of HDCD via a VAE; (2) perform GWAS on each encoded coordinate; and (3) use the polygenic risk scores (PRS) from the encoded coordinates as genetic scores for common biological functions and combine these scores to create a PRS for a specific disease or trait (given a small number of disease labels). Notably, REGL also allows the input to the decoder to optionally include the relevant EDF in the modified VAE architecture, thus encouraging the encoder to learn only the residual signals not represented by the EDF.
Discovery of new loci related to lung and cardiovascular function
We demonstrated the capabilities of REGLE using two high-dimensional clinical data modalities: spirograms to measure lung function and PPGs to measure cardiovascular function. Both can be collected non-invasively and relatively inexpensively in the clinic or from consumer wearable devices, and both modalities have well-known features (e.g., FEV1 or FVC for spirograms, and presence or location of the dicrotic notch for PPG). Compared to genome-wide association studies of spirogram and PPG features of the same dimensionality, REGLE studies of learned encodings recover most known loci related to lung and circulatory function, while also discovering additional loci (e.g., 45% more significant loci for PPG). These loci may represent new drug targets if validated by further analysis and wet-lab experiments.
Improved genetic risk scores
A polygenic risk score (PRS) is a summary of the estimated effect of many genetic variants on a particular trait, expressed as a single number. PRSs produced by genome-wide association studies on REGLE embeddings can be combined using only a few disease labels to generate a PRS for that specific disease (step 3 above). We found that lung function PRSs produced from spirogram encodings improve prediction of COPD and asthma compared to existing methods (e.g., PRSs produced by expert-defined features, PCA, and spline fitting) and stratify risk groups more effectively than feature PRSs at both ends of the risk spectrum. As shown below, we found statistically significant improvements in multiple metrics (AUC-ROC, AUC-PR, Pearson correlation) across multiple independent datasets for asthma and COPD (COPDGene, eMERGE III, Indiana Biobank, EPIC-Norfolk).
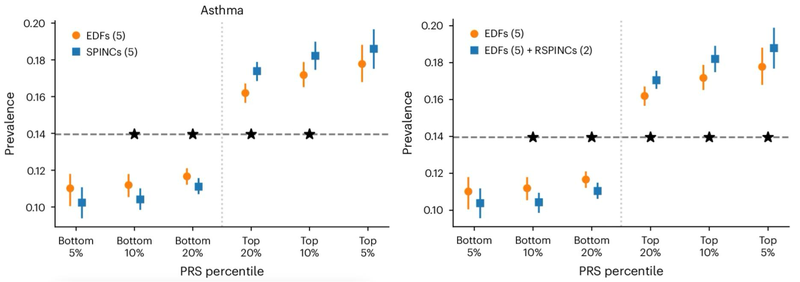
Comparison of the Expert-Defined Trait PRS for Asthma with the Spirogram Encoding (SPINC) and Residual Spirogram Encoding (RSPINC) PRS SpreadHorizontal dashed lines indicate overall prevalence; lower is better for the lower percentile and higher is better for the upper percentile. * indicates a statistically significant difference.
Similarly, PRS derived from REGLE embedding of PPG improves prediction of hypertension and systolic blood pressure (SBP). Working with academic collaborators, we evaluated hypertension and SBP PRS generated by PPG encoding and PPG features on three independent datasets (COPDGene, eMERGE III, EPIC-Norfolk) in addition to the UK Biobank held-out test set. Across multiple datasets, we found consistent trends of improvement using PPG-encoded PRS over expert-defined feature PRS for both hypertension and SBP.
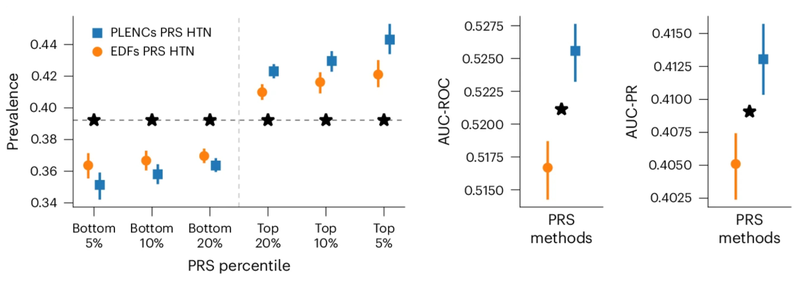
Comparison of PPG-encoded (PLENC) PRS for hypertension (HTN). Prevalence, AUC-ROCand AUC-PR Calculations are performed. * indicates a statistically significant difference.
Partially Interpretable Embeddings
use Driving force Due to the nature of REGLE, we studied the effect of encoding coordinates on the shape of the spirogram by fixing the values of expert-defined features and varying one encoding coordinate while keeping the others at zero. We then use only the decoder part of the trained model to generate the corresponding spirogram. A typical flow-volume spirogram consists of two distinct parts: (1) a relatively short part until the peak flow is reached, where the flow increases monotonically as the volume increases; and (2) the main part of the spirogram, where the flow decreases monotonically. The figure below shows that as we vary the first coordinate, the second part (negative slope) widens or narrows while keeping the first part relatively fixed. Indeed, the concavity of the second part of the curve is well known to pulmonologists. CovingIt is an indicator of airway obstruction that is not adequately represented by standard EDF.
Conclusion
We present REGLE, an unsupervised learning method that performs genetic analysis, improved discovery of novel loci, and risk prediction, overcoming limitations of previous work on ML-based phenotyping. Unsupervised learning of HDCD representations for genomic discovery is appealing because EDFs are difficult to discover manually at scale. The REGLE framework also supports the principled use of such features in modeling by modifying traditional VAE architectures. We demonstrate REGLE on two clinical data modalities, spirograms and PPGs, that can be measured routinely in clinical settings, as well as passively and non-invasively via smartphones and wearable devices.
REGLE provides a mechanism to identify genetic effects on organ function even in the absence of labeled data, and of course expert features can be incorporated into the model. It also provides a method to create disease- and trait-specific PRS with very few labels. As biobanks containing rich imaging, activity monitoring, medical records, and paired-gene data continue to grow, we expect this method, or similar methods, to be increasingly used to further elucidate the genetic basis of human traits and disease.
Acknowledgements
This work is the collaborative effort of multiple collaborators and institutions. We would like to thank all of our collaborators: Justin Cosentino, Babak Behsaz, Yuchen Zhou, Zachary R. McCaw, Howard Yang, Andrew Carroll, Cory Y. McLean (Google), Davin Hill (Northeastern University), Tae-Hwi Schwantes-An, Dongbing Lai (Indiana University), John Bates (Verily), Brian D. Hobbs, Michael H. Cho (Brigham and Women's Hospital & Harvard Medical School), Robert Luben, Anthony P. Khawaja (Moorfields Eye Hospital & University College London). We would also like to thank Nick Furlotte for peer review of the manuscript, Greg Corrado and Shravya Shetty for their support, and Annisah Um'rani for help with publication logistics.




