Data augmentation approach to enable deep learning analyses
Augmenting nucleotide sequence datasets
An innovative augmentation strategy was implemented to address the challenges inherent to limited gene representation in deep learning applications, generating overlapping subsequences and shared nucleotide motifs to expand a dataset of the sequences without modification of any nucleotides. Indeed, each 300-nucleotide gene sequence was decomposed into overlapping k-mers of 40 nucleotides using a variable overlap range (5–20 nucleotides), with a requirement that each k-mer shared a minimum of 15 consecutive nucleotides with at least one other k-mer. This method generated variable overlaps, ensuring a high degree of data diversity without redundancy. The comprehensive coverage of overlapping subsequences across a single gene sequence has been illustrated in Fig. 1. The figure provided a visual representation, showcasing the structural integrity of the augmented sequences while also reflecting the increased complexity introduced by the overlapping strategy. According to the current configuration of the augmentation method, between 50% and 87.5% of each sequence was designated as invariant, preserving conserved regions that emphasize significant differences among subsequences to support effective model training. Conversely, a portion ranging from 12.5% to 50% at the left, right, or both ends of each sequence was treated as variable, introducing diversity into the subsequences and enriching the training dataset. This approach resulted in the generation of 261 subsequences from each original sequence, representing a substantial augmentation. The level of diversity, along with the controllable conserved regions, can be easily adjusted based on the specific characteristics of different datasets. For more detailed information on the augmentation approach, please refer to Sect. 5.2 of the Materials and Methods.
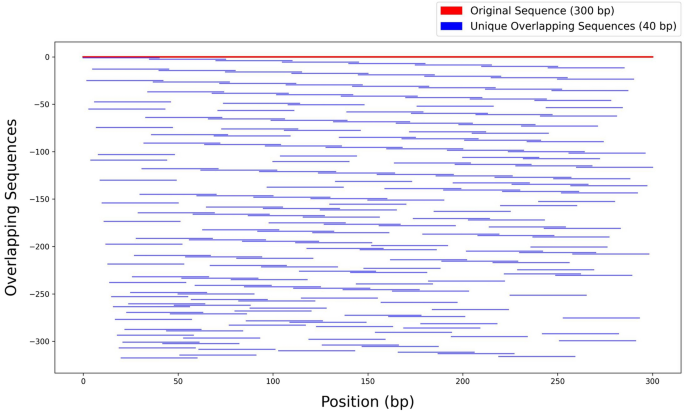
Graphical visualization of the augmentation process. A single original sequence and its corresponding generated overlapping subsequences were plotted. The red line represents the original 300-bp sequence, while the blue lines indicate each unique overlapping subsequence (40 bp) and their positional relationships. The graph highlights how variable overlaps and common base constraints create a complex yet non-redundant dataset.
Applying this approach, the original dataset of 100 sequences— e.g., for the chloroplast of C. reinhardtii dataset— was transformed into a larger dataset of 26,100 subsequences (261 subsequences per gene sequence). This approach provided a robust data foundation with high-quality representatives for each original sequence, enabling subsequent model training. Each overlapping subsequence provided a slight variation from the others, which increased the amount of data available for training deep learning models without altering the fundamental information contained in the original sequences. The nucleotide sequence augmentation approach was applied to the sequence datasets from eight different chloroplast genomes from microalgae and higher plants. These diverse datasets were used for further analysis to investigate the efficiency of the approach.
CNN-LSTM model performance on the augmented nucleotide sequences
The results of the CNN-LSTM hybrid model were independently evaluated across multiple genome datasets — C. reinhardtii, C. vulgaris, A. thaliana, T. aestivum, Z. mays, G. max, N. tabacum, and O. sativa—with and without the data augmentation (Fig. 2). The model performance on non-augmented data showed no accuracy, indicating its inability to make reliable predictions. However, the model performance showed significantly higher accuracy with data augmentation across all datasets (Fig. 2). For instance, accuracies for augmented data were highest for A. thaliana (97.66%), followed closely by G. max (97.18%) and C. reinhardtii (96.62%), demonstrating the model’s ability to generalize well across higher plant and algal genomes. The standard error analysis further supported the robustness of the augmented approach, as shown by the low error rates for genome datasets such as C. vulgaris (0.25%) and O. sativa (0.33%) (Fig. 2).

Average test accuracy across eight plastomes with and without the data augmentation across diverse genomes. The bar chart displays the average test accuracy (%) of the CNN-LSTM hybrid model independently applied to eight different genomes: Chlamydomonas reinhardtii, Chlorella vulgaris, Arabidopsis thaliana, Triticum aestivum, Zea mays, Glycine max, Nicotiana tabacum, and Oryza sativa. The average test accuracies were calculated across five trials. Two conditions are compared: models without data augmentation (red bars) and models with data augmentation (blue bars). Without augmentation, the average test accuracy for C. vulgaris was 0.476, while the average test accuracy for the other models was 0. Therefore, as a result, the red bars indicating the performance of models without data augmentation are not visible (except for C. vulgaris). The Student’s t-test analysis in mean average test accuracy for the model performance with and without the applied data augmentation was separately analyzed for each genome. The significant differences at t < 0.001 are marked by three asterisks (***).
Furthermore, the high accuracy achieved in both training and validation, coupled with minimal discrepancies between these metrics, indicated that the model generalized well to unseen data and did not suffer from overfitting. For instance, the CNN-LSTM model achieved a training accuracy of 97.13% with a corresponding training loss of 0.0641, while the validation accuracy reached 96.37% with a validation loss of 0.0671 on the C. reinhardtii dataset. The final test accuracy was 96.27%, with a test loss as low as 0.0661, further demonstrating the robustness of the model (Fig. 2). To further confirm the latter, the lack of improvement in validation accuracy, coupled with a steady rise in validation loss, demonstrated that the model was unable to generalize and learn effectively without data augmentation (Fig. 3A and S1). However, with data augmentation implementation, the model exhibited remarkable signs of learning and generalization. The training loss steadily decreased throughout the training process, converging to a minimal level close to zero by the final epochs (Fig. 3B and S1). Similarly, the validation loss showed a rapid initial decline, followed by a gradual decrease, ultimately reaching a low and stable value, which indicated the effectiveness of the learning model without substantial overfitting (Fig. 3B and S1). The model’s training accuracy increased continuously with each epoch, reaching over 96%. The validation accuracy followed a similar upward trajectory, achieving a comparable high accuracy, thus indicating successful generalization to unseen data. The close alignment of training and validation accuracy by the final epochs further confirmed that the model did not suffer from overfitting, affirming the robustness of the data augmentation strategy.

Training and validation performance of the CNN-LSTM model on the Chlamydomonas reinhardtii dataset with and without data augmentation. The figure presents the training and validation performance of the CNN-LSTM model across multiple epochs, tracking both the loss and accuracy metrics for the dataset (A) without the data augmentation and (B) with the data augmentation. The x-axis denotes the number of epochs, capturing the progress of model training, while the primary y-axis (left) illustrates the mean loss values for both training and validation sets. Training loss is represented by a blue line with circular markers, while validation loss is indicated by an orange line with cross markers. The secondary y-axis (right) displays accuracy in percentage, with green (squares) and red (diamonds) lines representing training and validation accuracy, respectively. The same training and validation performance of the other seven genome datasets are presented in S1.
Further analyses to investigate the efficiency of the augmentation approach
The model’s predictive efficiency was further quantified using the analysis of i) precision-recall curves and AUC scores, ii) correlation analysis of predicted vs. experimental data, and iii) feature importance analysis.
-
i)
Evaluation of precision-recall performance
The precision-recall curves of the CNN-LSTM hybrid model were evaluated across multiple genomes, both with and without data augmentation, using the average area under the curve (AUC) as a metric. For the non-augmented datasets, the average AUC values ranged between 0.021 and 0.065 (Fig. 4). Specifically, the highest average AUC was observed for T. aestivum (0.0656), followed closely by C. reinhardtii (0.0524) and Z. mays (0.039) when no augmentation was applied (Fig. 4). In contrast, the results indicated significantly higher average AUC scores when the CNN-LSTM hybrid model was trained on the same datasets with data augmentation. For example, the average AUC for augmented data reached its peak for A. thaliana (0.991), followed closely by T. aestivum (0.9867) and C. reinhardtii (0.984). Statistical analysis revealed that the average AUC scores for each genome dataset were significantly higher (t < 0.001) across Augmented datasets compared to those without augmentation (Fig. 4).

Comparison of average area under the curve (AUC) mean scores for the CNN-LSTM hybrid model with and without augmentation across the multiple genome datasets. The bar chart presents the average AUC scores of the model independently evaluated on eight distinct plastid genomes: Chlamydomonas reinhardtii, Chlorella vulgaris, Arabidopsis thaliana, Triticum aestivum, Zea mays, Glycine max, Nicotiana tabacum, and Oryza sativa. Two experimental conditions are shown: model trained without augmentation (red bars) and model trained with data augmentation (blue bars). The Student’s t-test comparing the mean AUC scores for model performance with and without the data augmentation for each genome was separately analyzed, and the mean differences (t < 0.001) were indicated by three asterisks (***).
Complementing the precision-recall analysis, the precision-recall curves for 10 randomly selected classes of distinct datasets were illustrated. The results demonstrated improvements in both the average precision (AP) and area under the curve (AUC) metrics when data augmentation was applied (Fig. 5 and S2). For instance, with data augmentation on C. reinhardtii dataset, the model achieved the highest AP and AUC values (AP = 1.00 and AUC = −1.00) across the 10 classes (Fig. 5 B), underscoring the robustness gained through introducing variability during training. In contrast, the highest AP and AUC values were 0.33 and −0.50, respectively, on the same dataset without augmentation (Fig. 5 A). More specifically, the AUC values without augmentation ranged from −0.06 to −0.50, whereas with augmentation, they remarkably increased to a range between −0.99 and −1.00 (Fig. 5).
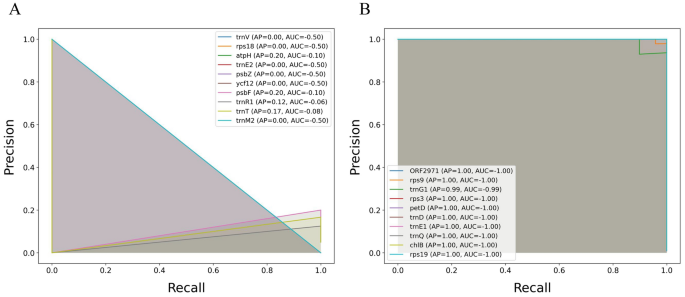
Precision-recall curve analysis for the CNN-LSTM model with and without augmentation with class-specific average precision (AP) and area under the curve (AUC) metrics across the Chlamydomonas reinhardtii dataset. This plot presents the precision-recall curves for a subset of 10 randomly selected classes from the model’s classification of the dataset (A) without the data augmentation and (B) with the data augmentation. The precision-recall curve for each class indicates the trade-off between precision and recall values at varying thresholds. The curves are accompanied by the average precision (AP) score and the area under the curve (AUC) for each selected class. The precision-recall curve analysis of the DNA datasets of the other seven plastomes is shown in S2.
-
ii)
Correlation analysis of predicted versus experimental data
To further assess the impact of data augmentation on model performance, the Pearson correlation coefficient between model predictions and experimental values was analyzed across eight genome datasets, both with and without data augmentation. The data augmentation consistently demonstrated higher significant (t < 0.001) correlation values for all genome datasets (Fig. 6), reflecting an enhanced alignment between the model’s predictions and the actual experimental outcomes. For instance, across the genome dataset from C. reinhardtii, the hybrid model with data augmentation achieved a Pearson correlation coefficient of 0.98 that significantly surpassed those without augmentation with a Pearson correlation coefficient of 0.00 (Fig. 6). This indicated that data augmentation notably enabled the model’s ability to generalize, leading to predictions that match the empirical data more closely.

Model performance via Pearson correlation coefficients between predicted and actual values across multiple genomes with and without the augmentation. The bar chart illustrates the Pearson correlation coefficients between model predictions and experimental values for eight plastomes—Chlamydomonas reinhardtii, Chlorella vulgaris, Arabidopsis thaliana, Triticum aestivum, Zea mays, Glycine max, Nicotiana tabacum, and Oryza sativa. The hybrid model for each genome dataset was evaluated under two conditions: without augmentation (red bars) and with data augmentation (blue bars). The significant mean difference (t < 0.001) for Pearson correlation coefficients was separately analyzed with the Student’s t-test between the model performance for each genome with and without the data augmentation, showing by three asterisks (***).
-
iii)
The feature importance analysis
The feature importance analysis utilizing SHAP (SHapley Additive exPlanations) was used to identify key features contributing to the prediction accuracy of the CNN-LSTM on the datasets applied to the data augmentation (Figs. 7 and S3). The analysis provided valuable insights into elucidating the interactions between selections of influential features (nucleotides in specific positions) in the prediction model for some sequences of the datasets. For instance, by applying data augmentation on the C. reinhardtii dataset, the hybrid model demonstrated a marked ability to interpret the significance of specific nucleotides during classification tasks (Fig. 7 and S3).

Feature importance analysis of nucleotide sequences from the augmented Chlamydomonas reinhardtii plastome dataset was performed using SHAP within the CNN-LSTM model framework. Each feature corresponds to a specific nucleotide type (A, C, G, or T) at one of the 40 positions within each input sequence, as encoded by a one-hot representation. Therefore, each nucleotide position contributes four potential features, resulting in a total of 160 (40*4) distinct feature indices. In the SHAP interaction plot, each feature pair interaction is visualized in two symmetrical positions: where each feature is considered once as the primary and again as the secondary feature. Each feature pair position includes multiple points (five in this plot as a subset of samples), representing how the interaction varies across different samples in the dataset. Each dot’s color indicates the feature value, with blue dots representing lower values and red dots representing higher values. The direction (left or right of zero) and spread of points demonstrate the interaction’s influence, with points to the right indicating positive interaction contributions to the model and points to the left showing negative interactions. The same feature importance analysis of the other seven genome datasets is presented in S3.
The matrix-like plot of the dataset revealed that certain feature pairs, such as those contained features 15 and 90, exhibited a broad distribution across both positive and negative SHAP interaction values. The latter indicated that these features could have a significant impact on model predictions in both enhancing and suppressive capacities (Fig. 7). Conversely, interactions between features 22 and 4 displayed clustering near the baseline, suggesting a relatively minimal impact on the model’s output. Furthermore, high values of feature 88 interacting with feature 67 resulted in a more pronounced positive interaction effect, as indicated by the cluster of red points to the right of the baseline (Fig. 7).
Augmenting of protein sequence datasets
To evaluate the performance of the CNN-LSTM hybrid model, protein datasets that underwent data augmentation were also employed. These datasets presented two remarkable differences from the DNA datasets, which posed additional challenges for applying deep learning models to identify meaningful patterns. First, a substantial portion of the genes of the plastome corresponds to non-coding RNAs, including tRNAs and rRNAs, resulting in a lower number of corresponding protein sequences in each dataset. For example, the DNA dataset for A. thaliana included 113 genes, while its protein dataset contained 56 sequences. Second, the variability in coding sequence (CDS) lengths leads to a highly imbalanced dataset, complicating the training of models. For instance, the varied lengths of the protein sequences in the C. reinhardtii dataset, ranging from 112 to 1995 aa, led to the generation of 73 subsequences from the shorter sequence (rpl20) while 1956 subsequences were generated from the longer sequence (ORF1995) (S4). Consequently, the effectiveness and efficiency of the data augmentation technique were assessed to address limitations associated with low sequence counts and imbalance.
Efficiency of the data augmentation on the model performance
The performance of the hybrid model, with and without the data augmentation, was evaluated across eight genomes using four metrics: macro precision, macro recall, macro F1 score, and their corresponding weighted scores (Fig. 8). The results showed a significant distinction between the performance of the model applied to each dataset with and without data augmentation. Model performance on each dataset with the data augmentation significantly outperformed its corresponding counterparts without it. Specifically, for the macro scores, the model applied to the datasets with data augmentation demonstrated precision values ranging from 0.971 to 0.991, recall values ranging from 0.973 to 0.992, and F1 scores from 0.969 to 0.992 across the genomes, indicating generally high performance (Fig. 8). The weighted scores also revealed similar trends, with precision values ranging from 0.924 to 0.990, recall values from 0.925 to 0.989, and F1 scores from 0.924 to 0.989 (Fig. 8).
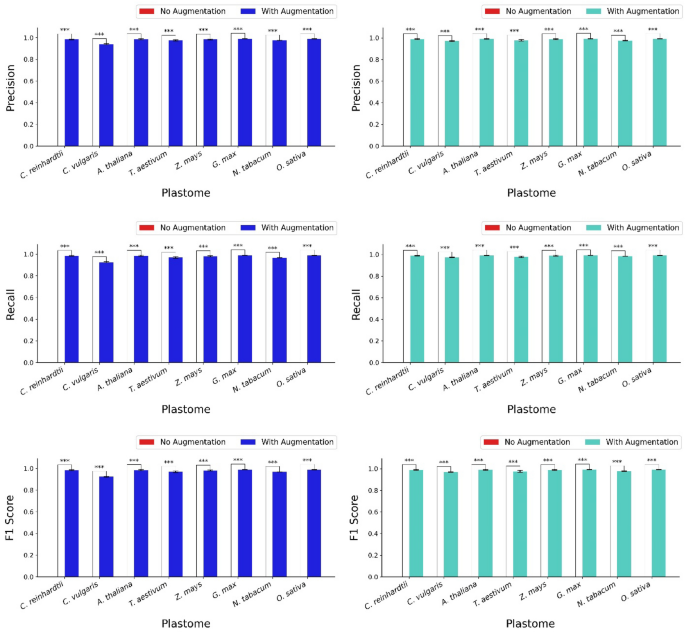
Average precision, recall, and F1 score (both macro and weighted) across eight genomes with and without the data augmentation. The bar chart shows the average macro precision, recall, and F1 score, as well as weighted precision, recall, and F1 score of the CNN-LSTM hybrid model independently applied to eight different plastomes: Chlamydomonas reinhardtii, Chlorella vulgaris, Arabidopsis thaliana, Triticum aestivum, Zea mays, Glycine max, Nicotiana tabacum, and Oryza sativa. Average test accuracies were calculated across three trials and five folds. Two conditions are compared: models with data augmentation (blue bars for macro scores and turquoise bars for weighted scores) and models without the data augmentation (red bars). Since the metric values for all datasets without augmentation were 0, the red bars indicating the performance of models without data augmentation are not visible. Statistical analysis using the Student’s t-test was conducted to compare mean test accuracies for each genome under both conditions, with significant differences at t < 0.001 marked by three asterisks (***).
The improvements in the average precision, recall, and F1 score were statistically significant (t < 0.001) compared to the corresponding metrics of the model’s performance on the datasets without data augmentation, which resulted in zero values for all metrics (Fig. 8). In other words, significant differences comparing the mean performance metrics between the two conditions were observed for all genomes, confirming that data augmentation significantly enhanced the model’s performance across all genomes. Ultimately, the incorporation of data augmentation substantially improved the precision, recall, and F1 score of the CNN-LSTM hybrid model, highlighting its importance in optimizing model performance across various genomes (Fig. 8).
Performance evaluation of plastome protein sequences classifications
The confusion matrix provided a detailed visualization of the CNN-LSTM model’s classification performance across protein sequence categories within the datasets, which were highly imbalanced (Fig. 9 and S5). The results, e.g., for C. reinhardtii dataset, showed that the model achieved high accuracy, as indicated by the dominant presence of higher values along the diagonal of the matrix representing correctly predicted classifications. The diagonal cells reflected strong performance, where the true and predicted labels align across most protein sequence categories. Misclassifications were minimal, as most non-diagonal cells contain zero values, indicating no false predictions (Fig. 9). However, a few non-diagonal cells displayed low values, suggesting occasional misclassifications in certain protein sequence categories. These results highlighted the model’s ability to make accurate predictions overall, with only a small number of misclassifications occurring for specific classes. These low values highlighted specific classes where the model occasionally misclassifies sequences, which may be indicative of class confusion or potential imbalances in the dataset (Fig. 9).
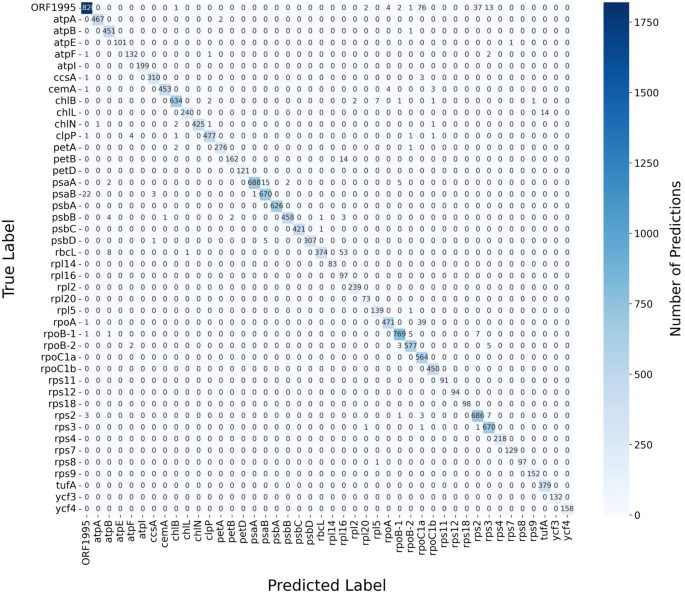
Confusion matrix for classifying Chlamydomonas reinhardtii chloroplast genome protein sequences with data augmentation using CNN-LSTM model. The confusion matrix presented here visualizes the model’s classification performance across different protein sequence categories in the dataset. Each row of the matrix corresponds to the actual class label, while each column represents the predicted label. Cells along the diagonal indicate correctly predicted classifications, where the true label matches the predicted label. Non-diagonal cells represent misclassifications, where the predicted label does not match the true label. The intensity of color in each cell corresponds to the number of instances predicted for that class, with darker shades representing higher counts. The confusion matrix for classifying the protein sequences of other seven plastomes are presented in S5.
Model performance using precision-recall and Receiver Operating Characteristic (ROC) curve analysis on plastome protein datasets
The performance of the CNN-LSTM model on the protein datasets with data augmentation was further evaluated using precision-recall and ROC curves (Fig. 10 and S6). In both curves, e.g., for the C. reinhardtii plastome protein dataset, the model demonstrated exceptionally high classification accuracy (Fig. 10). The precision-recall curve highlighted a near-perfect balance between precision and recall, with the curve consistently close to the top-right corner. This indicated that the model maintained a high precision across a wide range of recall values, minimizing false positives even as recall increased. The AUC for the precision-recall curve averaged 1.00 with a standard error of 0.00, underscoring the model’s strong ability to correctly classify positive instances with minimal error (Fig. 10).
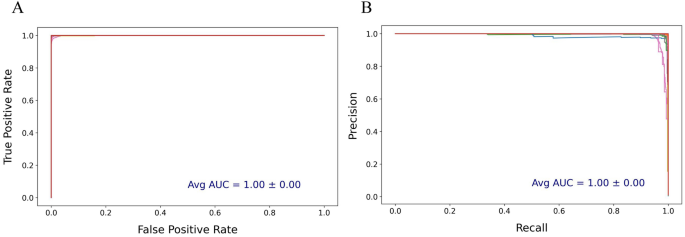
Precision-recall and Receiver Operating Characteristic (ROC) curves for Chlamydomonas reinhardtii plastome protein sequences with the data augmentation. This presents (A) ROC curves and (B) precision-recall generated from a CNN-LSTM model applied to the dataset protein sequence classification data over 50 epochs and 5 cross-validation folds. In the precision-recall curve, the x-axis indicates recall (sensitivity), and the y-axis shows precision. In the ROC curve, the x-axis represents the false positive rate, while the y-axis shows the true positive rate. The average AUC values from all classes was annotated for each plot. The precision-recall and ROC curves for other plastome protein datasets are shown in S6.
Similarly, the ROC curve showed an ideal shape, closely approaching the top-left corner of the plot. The false positive rate remained low as the true positive rate increased, reflecting the model’s high specificity and sensitivity in distinguishing true positives from false positives (Fig. 10). The average AUC for the ROC curve was also 1.00 with a standard error of 0.00, indicating consistent performance across all cross-validation folds and minimal variability. In other words, the ROC curve’s perfect diagonal placement reflected optimal performance, indicating a strong ability to discriminate between the positive and negative classes with minimal false positives and false negatives (Fig. 10).
Model performance across epochs using mean loss, F1 score, and recall on plastome protein datasets
The model demonstrated substantial learning and generalization (Fig. 11 and S7). For C. reinhardtii with the augmentation, the mean training loss exhibited a rapid decline during the first 10 epochs, after which it decreased steadily, approaching a minimal value by the final epochs (Fig. 11). This decrease in loss indicated effective model training with diminishing errors. Likewise, the mean validation loss followed a similar pattern, showing an initial sharp decline, followed by a gradual decrease, ultimately stabilizing at a low level, which showed that the model was learning effectively without significant overfitting (Fig. 11).
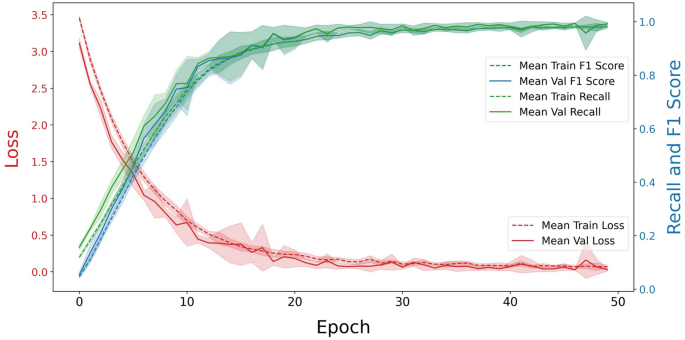
Model performance across epochs via mean loss, F1 score, and recall on plastome protein dataset of Chlamydomonas reinhardtii with augmentation using CNN-LSTM Model. This figure illustrated the model’s training and validation performance across 50 epochs, averaged over 5 cross-validation folds. The x-axis represents the number of epochs, while the y-axes display mean loss values (left y-axis) and performance metrics: recall and F1 score (right y-axis). The red, green, and blue curves depict the mean loss, recall, and F1 score for both the training (dashed line) and validation (solid line) sets. The shaded areas around these lines represent the standard deviation across folds, reflecting the variability in loss for each epoch. The same model performance for other genomes are presented in S7.
Both the training and validation recall and F1 scores showed consistent improvement throughout the epochs. The training recall and F1 scores steadily increased, surpassing 95% by the later epochs. The validation scores mirrored this upward trend, reaching similarly high values, which indicated that the model was able to generalize well to unseen data. The close alignment between the training and validation performance metrics in the final epochs further confirmed that overfitting was not present, emphasizing the effectiveness and robustness of the data augmentation approach (Fig. 11).
Therefore, the developed innovative augmentation strategy effectively addressed the challenges of applying deep learning to omics and other related biological datasets with limited data availability, particularly in cases where each gene or protein is represented by a single sequence. As illustrated in Fig. 12, a graphical flowchart summarizes the existing challenges posed by limited sequence availability for enabling deep learning applications in omics and related datasets, alongside the strategy to overcome these challenges for ease of understanding. This method expanded the datasets without introducing noise or causing overfitting, while preserving the biological integrity of the sequences and ensuring comprehensive coverage of the original data.
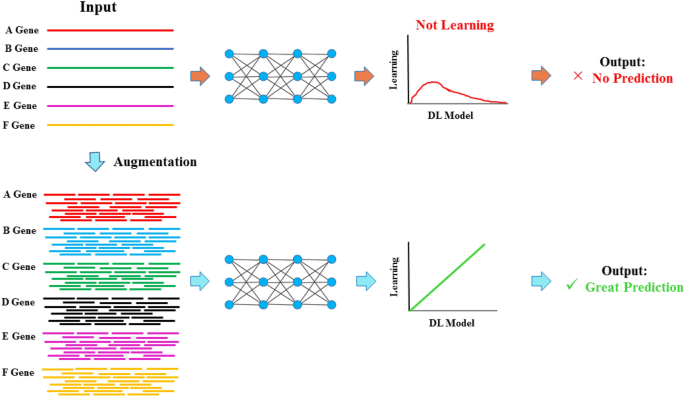
Graphical flowchart illustrating data augmentation strategy to address the challenges of limited sequence availability in omics and related biological datasets, thereby enabling effective deep learning applications.
Unveiling the power of the CNN-LSTM model for short omics sequences
The performance of the CNN-LSTM hybrid model was evaluated against other machine learning models, including Support Vector Machines (SVM), Multi-Layer Perceptron (MLP), Recurrent Neural Networks (RNN), and Convolutional Neural Networks (CNN), on datasets containing sequences of 40 nucleotides or amino acids. These models struggled to effectively learn patterns from the short sequences, achieving less than 18% accuracy on the test datasets (unpublished data). In contrast, the optimized CNN-LSTM model, with its architecture shown in Fig. 13, demonstrated significantly improved performance on both the training and test datasets, highlighting its ability to capture the complex patterns within the input sequences. This superior performance reflects the hybrid model’s capability to address the challenges posed by the short length and inherent complexity of the datasets, which could not be adequately handled by traditional models.
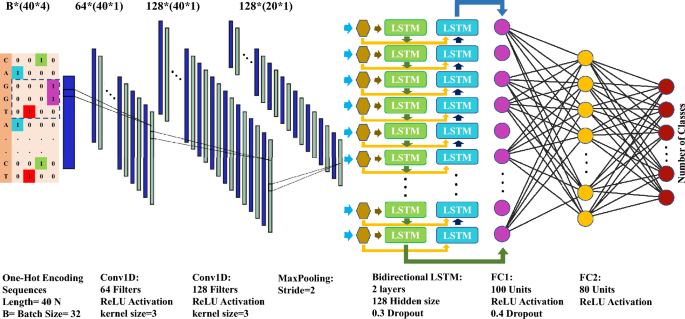
A graphical illustration of the optimized CNN-LSTM model architecture designed for datasets with limited gene representations. This architecture was independently evaluated on multiple nucleotide and protein datasets when applied with and without data augmentation. One-hot encoding sequences with four channels here are for nucleotide sequences that is different from that of amino acid sequences. The architecture comprises two sequential convolutional layers followed by a max-pooling layer. The output is reshaped and passed through a bidirectional LSTM with two layers, followed by two fully connected layers. The final output layer maps the processed features to the number of classes in the dataset. Further details are provided in the material and method section.
Data augmentation strategy for enabling unsupervised analysis
In unlabeled data analysis, an approach was developed to maximize feature generation from a dataset of limited genomic sequences, converting each 300-nucleotide sequence into a high-dimensional set of k-mer features (from 5 to 12 nucleotides). Using the k-mer approach, common motifs between each pair of sequences was quantified, identifying common in k-mer content even among a small dataset. This method produced thousands of pairwise metrics, capturing a high degree of sequence similarity and difference, which underscores the effectiveness of k-mer-based feature generation for producing robust datasets from limited gene samples. To determine all possible pairwise interactions among the dataset, the number of unique gene sequence pairs was calculated using the following formula:
$$\text{C}(\text{n},\text{k})=\frac{\text{k}!}{(\text{n}-\text{k})!\text{n}! }$$
In this context, n represents the total number of gene sequences under consideration, while k is set to 2, indicating that the analysis focuses on pairs of gene sequences.
For example, in the dataset of C. reinhardtii with 100 plastome sequence genes, 4,950 unique combinations of gene pairs were obtained. The resulting dataset comprised a vast number of k-mer counts across sequences, providing meaningful variations and increasing the dataset’s depth for unsupervised learning.
In this unsupervised approach, the k-mer extraction was applied to the same 300-nucleotide sequences that previously used for supervised augmentation. The goal was to capture recurring k-mers across sequence pairs, thus identifying core nucleotide patterns that may indicate biologically relevant relationships among sequences without requiring labeled data. By extracting k-mers across lengths of 5 to 12 nucleotides, the method generated a set of shared k-mers between sequence pairs, providing a framework for comparative analysis. The results revealed diverse levels of k-mer sharing between pairs, quantifying genetic similarity and establishing a foundational distance matrix for subsequent clustering and dimensionality reduction.
To assess the k-mer patterns, a distance matrix representing the inverse of common k-mers between pairs was created, which was then clustered using k-means and visualized through Principal Component Analysis (PCA). This clustering revealed distinct groups of sequences, each representing unique k-mer signatures. The process produced clusters that allowed for the visualization of sequence relationships based on k-mer composition, ultimately emphasizing how even a low count of genes can yield a feature-rich dataset appropriate for unsupervised learning. This analysis demonstrated the flexibility and utility of k-mer-based feature generation in producing sufficient, biologically relevant metrics from limited sequence data, meeting the needs of projects constrained by small gene counts (Fig. 14 and S8).
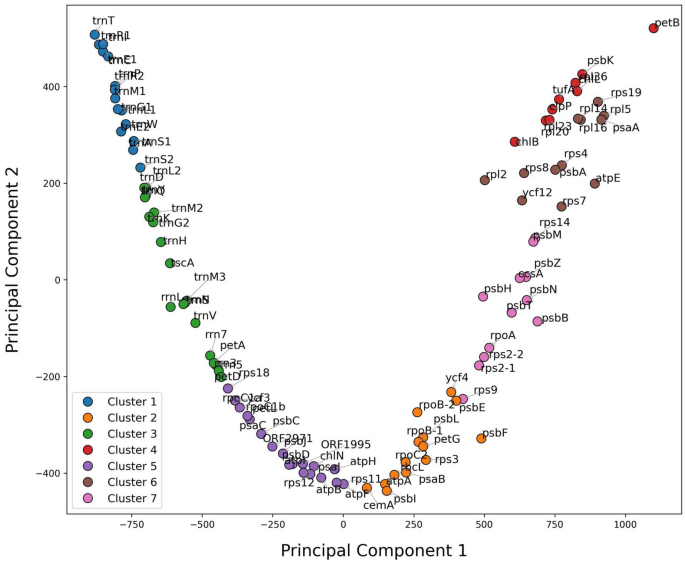
Clustering of the C. reinhardtii plastome dataset of 300-nucleotide sequences based on the total number of common k-mers. Each data point on the scatter plot was annotated with its corresponding sequence, allowing for easy identification and interpretation of individual sequences within each cluster. Every group of sequence clustering in a group was distinguished with the same color. The clustering plots of the DNA datasets of other seven plastomes are presented in S8.
Further analysis of the identified clusters revealed biologically meaningful groupings, where each cluster represented sequences sharing characteristic k-mer motifs. For example, Cluster 1 included sequences that appeared closely related based on recurrent k-mers, potentially indicating shared functions or evolutionary related traits. In contrast, Cluster 4 housed sequences with a different k-mer profile, pointing to distinct lineage or functional properties (Fig. 14). This clustering of unlabeled data demonstrated the potential of k-mer similarity measures to identify sequence relationships, providing insight into the inherent structure of genomic data in a label-free setting.




