Implementation of WGCNA and identification of key module genes
To identify modules significantly associated with NAFLD, we performed WGCNA on the GSE89632 dataset. Initially, we conducted hierarchical clustering analysis on the samples and detected two outlier samples, GSM2385767 and GSM2385782. These outliers were removed by setting the cutHeight to 130 (Fig. 2A). Subsequently, we chose an optimal soft threshold power of 5 (R2 = 0.9) to guarantee a scale-free topological network (Fig. 2B). Then, we defined modules by setting the “minModuleSize” to 50, “mergeCutHeight” to 0.6, and “deepSplit” to 1.0, resulting in the identification of nine distinct modules. Our analysis revealed a strong correlation between the MEbrown and NAFLD (correlation coefficient = -0.87, p = 3e-20) (Fig. 2C). Within the brown module, a highly significant correlation was observed between gene significance (GS) and module membership (MM) (correlation coefficient = 0.92, p < 1e − 200) (Fig. 2D). Ultimately, we identified 2,969 genes significantly associated with NAFLD within the brown module.
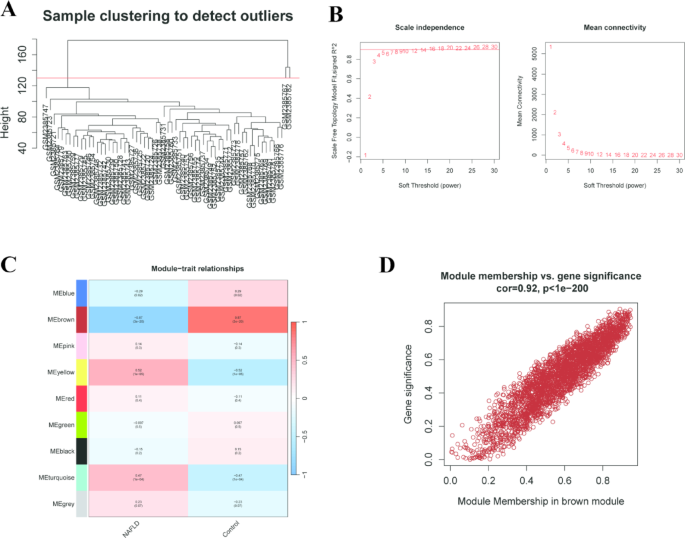
Implementation of WGCNA and identification of key module genes. (A) Hierarchical Clustering Dendrogram for Outlier Detection in Sample Dataset GSE89632. (B) A soft-thresholding power was set at 5, ensuring a scale-free R2 = 0.9. (C) Correlation Heatmap of Module-Trait Relationships in NAFLD and Control Samples. (D) Relationship Between Module Membership and Gene Significance in the Brown Module.
Identification and functional enrichment analysis of palmitoylation-related DEGs in NAFLD
To further elucidate the relationship between palmitoylation-related genes and the development of NAFLD, we subsequently conducted a comprehensive analysis of DEGs between normal and NAFLD samples in the GSE89632 dataset. Ultimately, a total of 872 DEGs were identified in NAFLD samples compared to normal samples (Fig. 3A, B). By intersecting these DEGs with genes from the brown module obtained through WGCNA and palmitoylation-related genes, we successfully identified 60 PR-DEGs (Fig. 3C). GO analysis of these 60 key genes revealed significant enrichment in biological processes (BP), including the regulation of inflammatory response, positive regulation of cytokine production, and response to lipopolysaccharide. In the cellular component (CC) classification, collagen-containing extracellular matrix and endocytic vesicle exhibited the highest enrichment scores. The molecular functions (MF) were primarily enriched in enzyme inhibitor activity, DNA-binding transcription activator activity, and cytokine activity (Fig. 3D, E) (Supplementary Table S2). KEGG pathway enrichment analysis indicated that the majority of these genes were highly enriched in pathways such as the TNF signaling pathway, lipid metabolism and atherosclerosis, IL-17 signaling pathway, and inflammatory bowel disease (Fig. 3F, G) (Supplementary Table S3).
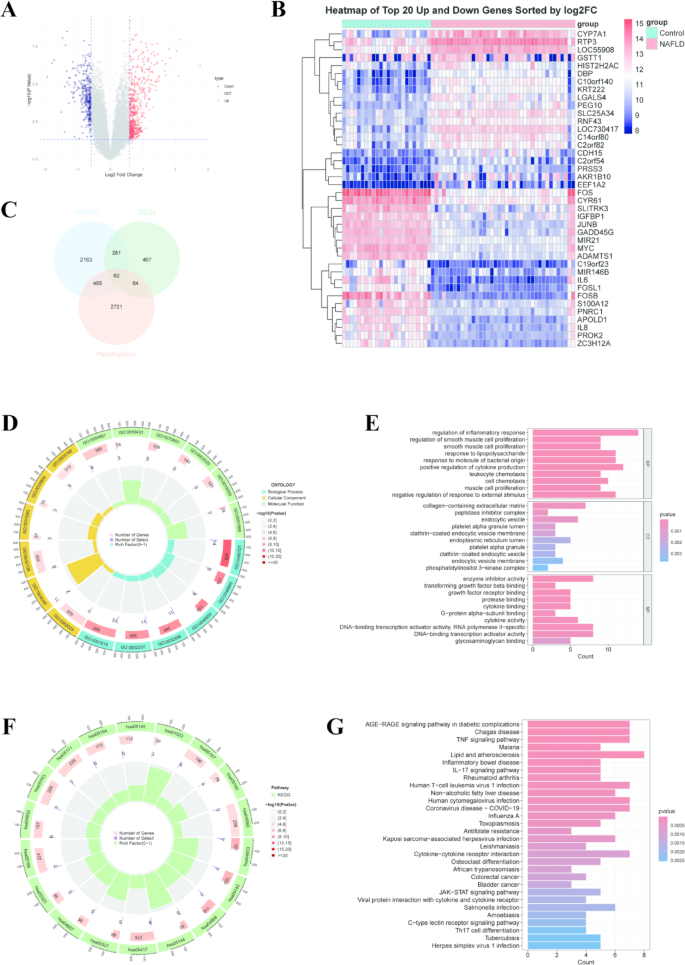
Association Analysis of Palmitoylation-Related Genes with the Development of NAFLD. (A) A volcano plot was constructed to identify DEGs between normal and NAFLD samples. (B) A heatmap was generated to display the top 20 upregulated and downregulated genes, sorted by log2 fold change. (C) A Venn diagram was utilized to illustrate the intersection of genes identified through WGCNA, DEGs, and palmitoylated genes. (D) GO analysis was performed, and the results were presented in a circular plot. (E) GO analysis was further visualized in a bar chart, with annotations for biological processes (BP), cellular components (CC), and molecular functions (MF). (F) KEGG pathway enrichment analysis was conducted, and the findings were depicted in a circular plot. (G) KEGG pathway enrichment analysis was also represented in a bar chart to provide a detailed overview of the enriched pathways.
Identification of PRHGs for NAFLD via machine learning
Based on the expression profiles of 60 PR-DEGs, we constructed seven machine learning models to further identify genes with high diagnostic value. The results indicated that the NNET and DT machine learning models exhibited relatively low residuals (Fig. 4A, B). We assessed the diagnostic performance of the seven machine learning algorithms in the test cohort using the AUC. The AUC values for the RF model, SVM model, GLM model, KNN model, NNET model, LASSO model, and DT model were 0.987, 0.974, 0.675, 0.948, 0.987, 0.974, and 0.955, respectively (Fig. 4C). We further ranked the top 10 significant feature variables in each model using the root mean square error (RMSE) (Fig. 4D). Collectively, the NNET and DT machine learning models demonstrated the highest efficiency in distinguishing between normal individuals and patients with NAFLD among the seven models. We selected the top 10 predictive genes from both the NNET and DT models and identified three PRHGs at their intersection for further investigation: TYMS, WNT5A, and ZFP36 (Fig. 4E). The AUC values for TYMS, WNT5A, and ZFP36 were 0.958, 0.963, and 0.922, respectively, indicating their excellent diagnostic potential (Fig. 4F).
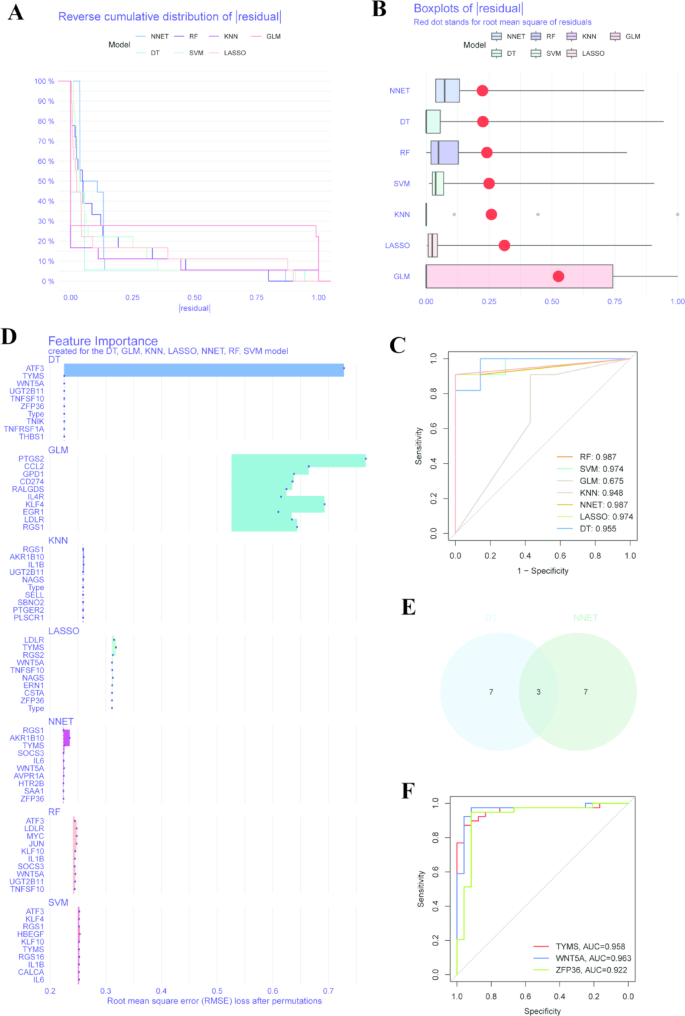
Construction and evaluation of seven machine models. (A) The reverse cumulative distribution of residuals for seven machine learning models (NNET, RF, KNN, GLM, DT, SVM, LASSO). (B) The boxplot of residuals. The boxplot illustrates the distribution of residuals for the seven machine learning models (NNET, RF, KNN, GLM, DT, SVM, LASSO). Red dots represent the root mean square error (RMSE) of the residuals. (C) The ROC curves and corresponding AUC values for seven machine learning models (RF, SVM, GLM, KNN, NNET, LASSO, DT). (D) The top 10 significant feature variables in seven machine learning models (DT, GLM, KNN, LASSO, NNET, RF, SVM), ranked according to root mean square error (RMSE). (E) The intersection of the top 10 predicted genes in NNET and DT models. (F) The ROC curves and corresponding AUC values for TYMS, WNT5A, and ZFP36.
Gene expression analysis and construction and validation of the nomogram
We initially compared the expression levels of TYMS, WNT5A, and ZFP36 genes between the control group and the NAFLD patient group. The results revealed that the expression of TYMS and WNT5A was significantly upregulated, whereas the expression of ZFP36 was significantly downregulated in the NAFLD group (Fig. 5A). Based on these differences, we constructed a nomogram model that converts the relative expression levels of each gene into scores, allowing for the estimation of NAFLD risk based on the cumulative score (Fig. 5B). To evaluate the effectiveness of this model, we further analyzed its calibration and discrimination capabilities. The calibration curve (Fig. 5C) demonstrated a high degree of concordance between predicted probabilities and actual observed probabilities, indicating excellent calibration. The C-index was 0.976 (95% CI: 0.935–1.018) (Fig. 5C), and the area under the ROC (AUC) was 0.976 (Fig. 5D), confirming the model’s superior discrimination ability. Moreover, we validated the accuracy of these candidate biomarkers in an external dataset (GSE164760). The results were largely consistent with our initial findings, demonstrating that TYMS and WNT5A were upregulated, whereas ZFP36 was downregulated in the NAFLD group (Fig. 5E). Further analysis using the nomogram and ROC curve provided additional support for the diagnostic value of these genes (Fig. 5F, H). In summary, our findings suggest that TYMS, WNT5A, and ZFP36 hold promise as diagnostic biomarkers for NAFLD.
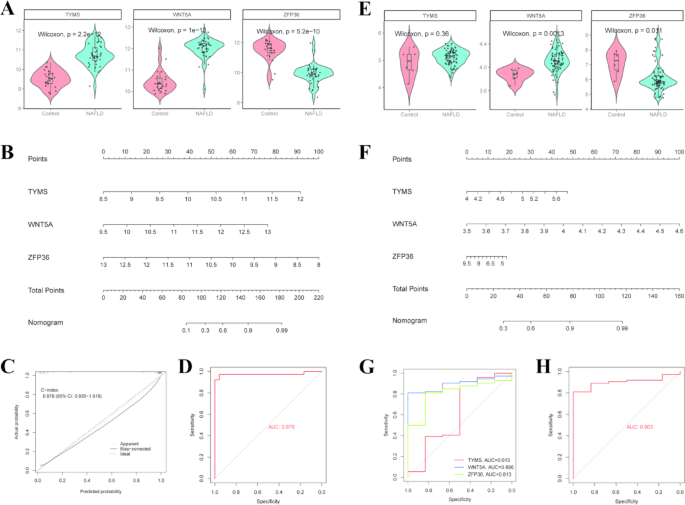
Evaluation of the diagnostic potential of candidate biomarkers and construction of a nomogram in the training (GSE89632) and validation cohorts (GSE164760). (A) The violin plots elucidate the differential expression profiles of TYMS, WNT5A, and ZFP36 between the control and the NAFLD group in the training cohort. (B) A predictive nomogram for NAFLD occurrence constructed from PRHGs. “Points” indicate the scores of hub genes, and “Total Points” represent the sum of all these gene scores. (C) The calibration curve illustrates the relationship between actual and predicted NAFLD rates. A perfectly accurate model would be represented by the dotted diagonal line. The performance of the nomogram is depicted by the solid line, with its closeness to the diagonal indicating greater precision. (D) ROC curve of the nomogram in training cohort for NAFLD. (E) The violin plots elucidate the differential expression profiles of TYMS, WNT5A, and ZFP36 between the control and the NAFLD group in the validation cohort. (F) A predictive nomogram for NAFLD occurrence constructed from PRHGs in the validation cohort. (G) The ROC curves and corresponding AUC values for TYMS, WNT5A, and ZFP36 in the validation cohort. (H) ROC curve of the nomogram in validation cohort for NAFLD.
GSEA and GSVA pathway analyses reveal molecular mechanisms of PRHGs in NAFLD
To further investigate the molecular mechanisms of the three PRHGs in the diagnosis of NAFLD, we performed GSEA-KEGG pathway enrichment analysis for each gene biomarker. The graphical representations highlighted the top five most enriched pathways (Fig. 6A-C and Supplementary Table S4-6). By intersecting the enrichment results of these three hub genes, we identified several key pathways in which they were commonly enriched. These pathways include mitochondrial complex UCP1 in thermogenesis, the impact of inactivated PINK1 on electron transfer in Complex I, aberrant TDP43 on electron transfer in Complex I, electron transfer in Complex I, the effects of aberrant SNCA and ABETA on electron transfer in Complex I, DNA replication origin unwinding and elongation, and the interaction between CENPE and the NDC80 complex (Fig. 6D, E). Subsequently, NAFLD samples were stratified into high- and low-expression groups based on the median expression levels of the hub genes. GSVA enrichment analysis was then performed to investigate the differential pathways between these groups. Comprehensive analysis indicated that high expression of TYMS may activate serotonin metabolism, hormone-like cytokine to JAK-STAT signaling pathway, GH JAK-STAT signaling pathway, and PAX8-PPARG fusion to PPARG-mediated transcription. Conversely, low expression of TYMS was associated with the activation of pathways such as DNA replication origin unwinding and elongation, pre-initiation complex (pre-IC) formation, aberrant ABETA to crosstalk between extrinsic and intrinsic apoptotic pathways, and keratan sulfate degradation (Fig. 6F). High expression of WNT5A was linked to the activation of the KEAP1-NRF2 signaling pathway, tyrosine degradation, and gluconeogenesis, while low expression of WNT5A was associated with the activation of the activin signaling pathway, nodal signaling pathway, and HSV GD to HVEM NFKB signaling pathway (Fig. 6G). Similarly, high expression of ZFP36 was related to the TLR7/8/9-IRF5 signaling pathway and TLR7/9-IRF7 signaling pathway, whereas low expression of ZFP36 was associated with the PERK-ATF4 signaling pathway (Fig. 6H).
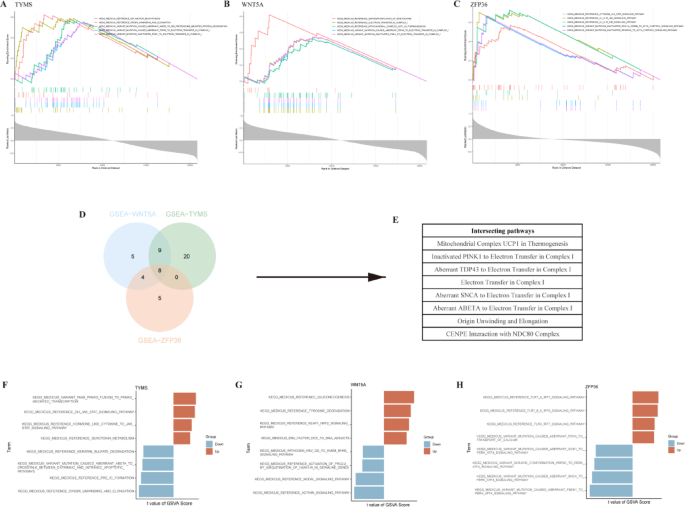
GSEA and GSVA of 3 PRHGs. GSEA of TYMS (A), WNT5A (B) and ZFP36 (C) using KEGG gene sets. (D, E) Intersections of GSEA results for 3 PRHGs. GSVA of TYMS (F), WNT5A (G) and ZFP36 (H) using KEGG gene sets.
The correlation of the PRHGs with singlecell characteristics
In order to evaluate the functions of PRHGs within the hepatic microenvironment at the single-cell transcriptomic level, we comprehensively analyzed the expression profiles of TYMS, WNT5A, and ZFP36 across distinct cell types. Our results revealed that TYMS is predominantly expressed in NK cells, WNT5A is primarily expressed in hepatocytes and monocytes, while ZFP36 exhibits widespread expression across various cell types. Notably, hepatocytes were identified as the main cell type in which all three PRHGs are co-expressed (Fig. 7A-E). Utilizing the “AddModuleScore” function, we assigned the signature-specific score to each cell based on PRHGs expression. Subsequently, cells were categorized into high-score and low-score groups according to their PRHGs scores (Fig. 7F), and differential expression analysis was performed to identify DEGs. GSEA demonstrated that these DEGs were significantly enriched in pathways and functions such as Cytokine Signaling in the Immune System, IL-18 Signaling Pathway, Cytokine Mediated Signaling Pathway, Response to Cytokine, and Inflammatory Response (Fig. 7G, H). Intriguingly, liver cells within the microenvironment exhibited diverse communication patterns depending on their PRHGs scores (Fig. 7I). Within this intricate hepatic microenvironment, various cell types can act as Sender, Receiver, Mediator, and Influencer during cellular communication, ultimately leading to distinct intercellular signaling cues. Our study identified significant alterations and key influencers in the cell communication signals of the high-score group, particularly in MIF, APP, and COLLAGEN signaling (Fig. 7J–L). These findings suggest that such signals may play regulatory roles in inflammation, fibrosis, immune modulation, and metabolic processes within the hepatic microenvironment.
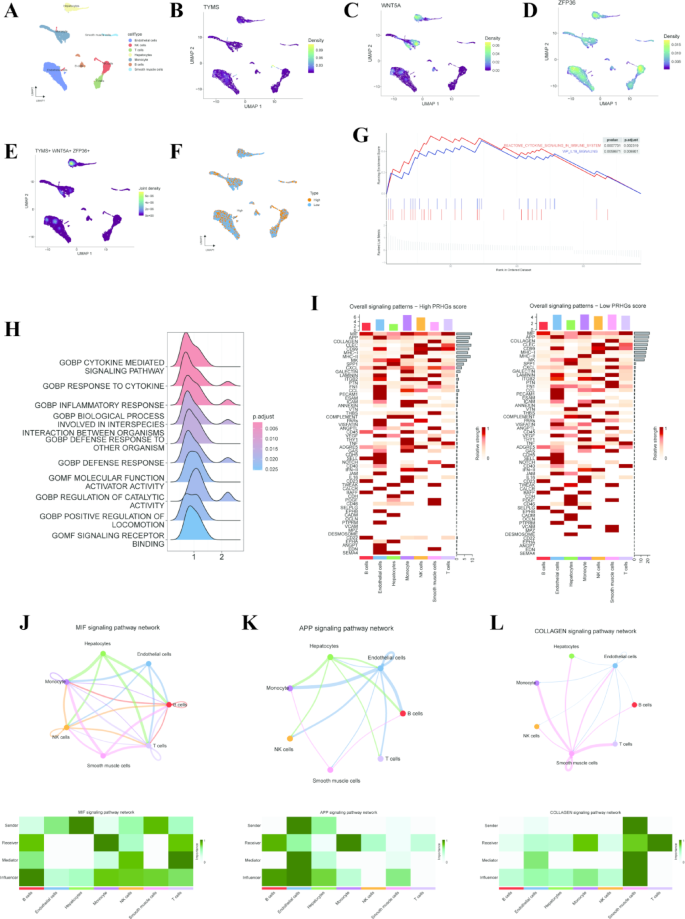
The single-cell characteristics of PRHGs. (A) UMAP plot of cell type annotation using the “SingleR” package. Density of TYMS: (B), WNT5A: (C), ZFP36: (D) Gene Expression. (E) Density of Co-expressed TYMS, WNT5A, and ZFP36 Genes. (F) The UMAP plot categorizes cells into two groups based on their signature-specific scores derived from the expression of PRHGs. (G) The results of GSEA performed using the c2.cp.v2024.1.Hs.symbols.gmt gene set collection. (H) The results of GSEA performed using the c5.go.v2024.1.Hs.symbols.gmt gene set collection. (I) Identifying distinct signal pathways in varying PRHGs score groups. (J–L) The signaling pathway networks for MIF (J), APP (K), and COLLAGEN (L) pathways, with heatmaps showing cell type involvement.
Immune cell infiltration analysis
To investigate the immune response mechanisms in NAFLD, we employed the CIBERSORT algorithm to evaluate the changes in immune cell abundance between NAFLD and control groups (Fig. 8A, B). Our results revealed that, compared to control samples, NAFLD samples exhibited significantly higher levels of plasma cells, activated memory CD4 T cells, monocytes, activated dendritic cells, activated mast cells, and neutrophils. Conversely, there were lower levels of resting memory CD4 T cells, γδ T cells, activated NK cells, M1 macrophages, M2 macrophages, resting dendritic cells, and resting mast cells in NAFLD samples. Subsequent correlation analysis through a heatmap of immune cell interactions demonstrated that in NAFLD samples, M0 macrophages were positively correlated with CD8 T cells (r = 0.55), and γδ T cells were positively correlated with M2 macrophages (r = 0.63). In contrast, monocytes exhibited negative correlations with γδ T cells (r = – 0.75) and M2 macrophages (r = – 0.71) (Fig. 8C). These findings suggest that the immune profile and the interactions among various immune cell types in NAFLD patients are distinct from those in controls. Additionally, a correlation bubble chart was utilized to illustrate the associations between the PRHGs and different immune cells (Fig. 8D).
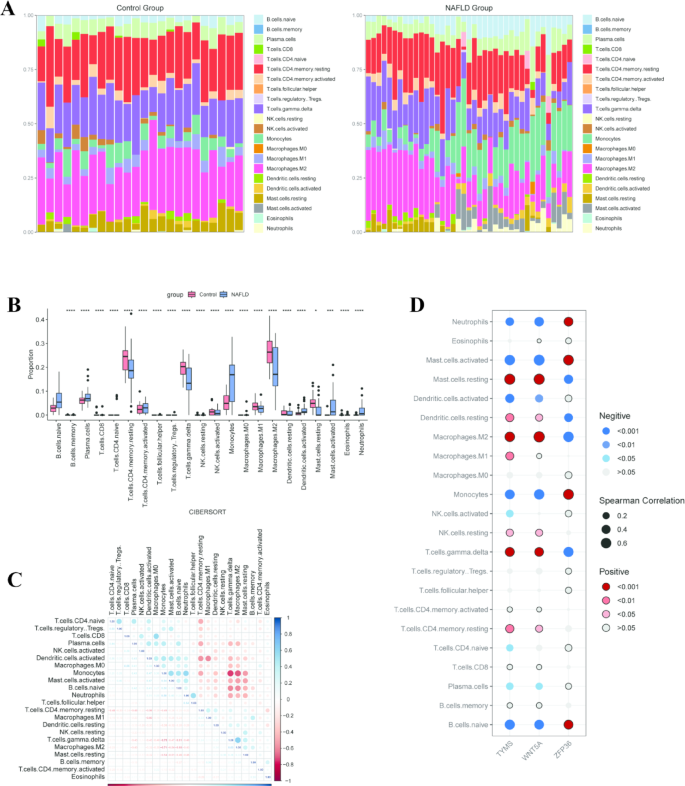
Analysis of immune cell infiltration. (A) The bar plot visualizing the proportion of infiltrating immune cells in control and NAFLD groups. (B) The boxplot comparing the proportion of immune cells between NAFLD and control groups. *p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001. (C) The correlated heatmap representing the correlation between different immune cells in NAFLD. (D) Correlation analysis of infiltrating immune cells with three PRHGs.
Construction of the TFs-genes and genes-miRNAs regulatory network
Given the importance of TFs and miRNAs in elucidating disease progression, we constructed TFs-genes and genes-miRNAs network to identify the transcriptional and post-transcriptional regulators of PRHGs, thereby enhancing our understanding of the disease. The TFs-genes network comprises 202 nodes and 228 edges (Fig. 9A), while the genes-miRNAs network consists of 327 nodes and 501 edges. By selecting the “Minimum Network” option, the genes-miRNAs network was reduced to a subnetwork with only 4 nodes and 3 edges (Fig. 9B). Notably, hsa-let-7b-5p interacts with three hub genes, suggesting that it may serve as a common regulator of these genes. However, these findings require further validation to confirm their roles.
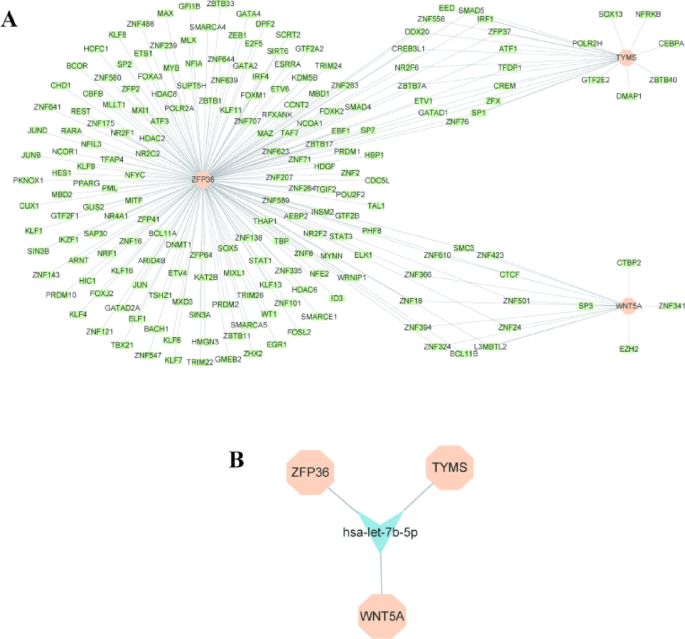
Network for TFs-genes and genes-miRNAs with three hub genes. (A) TFs-genes regulatory network of three hub genes. (B) Genes-miRNAs regulatory network of three hub genes.
Identification of drug candidates
We utilized the DSigDB database on the Enrichr platform to identify potential drugs targeting hub genes. A total of 265 gene-drug associations were identified (Supplementary Table S7). The top 10 compounds were selected based on their p-values and adjusted p-values. These potential therapeutic molecules include troglitazone (CTD 00002415), folic acid (CTD 00005997), TPEN (CTD 00001994), mitomycin C (CTD 00007136), progesterone (CTD 00006624), calcitriol (CTD 00005558), medroxyprogesterone acetate (CTD 00006623), KT 5720 (CTD 00002405), trifluridine (TTD 00011552), and 2-mercaptoethanol (TTD 00000601) (Table 1).




