This retrospective study was approved by the Institutional Review Board (IRB) of Yongnam University Hospital. The requirement for informed consent was exempt by the IRB due to a retrospective design of the study. This study was conducted in accordance with the Declaration of Helsinki.
patient
All VFSS data from the hospital were included to diagnose or monitor the progress of patients with dysphagia between March 2019 and July 2022. Exclusion criteria include (1) patients under 20 years of age, (2) tracheotomy, (3) having facial or cranial abnormalities, and (4) with metal plates in the VFSS field.
Video fluid scopy study
The VFSS was performed with a patient standing upright in a wheelchair or chair, with head position upright and staring forward. VFSS images were stored as digital media at 30 frames per second using a scan converter. A Bonorex 300 injection (Iohexol 647 mg/ml, Central Medical Service, Seoul, Korea) was used as the contrast medium. VFSS contains multiple swallows of different consistency (liquid, semi-solid, and solid) food contrast mixtures, but only the first 3 mL liquid swallows were analyzed to avoid residual contrast affecting image analysis.
Deep learning model development
A deep learning model for automatic swallowing dysphagia diagnosis was developed using Python 3.10.15, Pytorch 2.14.1, Scikit-Learn1.5.2, and Deepspeed 0.16.3.
VFSS video frame-by-frame analysis involves converting the original 1280\(\:\ times \:\)1024 Pixel MPG files are filed into individual frames using custom Python scripts that leverage the OPENCV library. Frames were extracted at a rate of 10 frames per second (i.e., one frame for each of the three original frames). Images containing the larynx (Epiglottis, Supraglottis, Glottis, and Subgrotis) and upper esophageal sphincter were then automatically extracted (Fig. 1) and sized to 224 × 224 pixels. ROI was extracted using the REDET object detection model. Redet Model offers great advantages for VFSS image analysis with its inherent rotation equality and accurate depiction of orientation ROI. This allows Redet to accurately detect and separate anatomy or bolus using a rotated bounding box. Chall often interferes with traditional object detectors even when rotational variations occur due to patient movement or positioning. Redet's ability to accommodate rotational variations allows for the robust and accurate extraction of critical spatio-time information from VFSS sequences, thereby improving the reliability and accuracy of downstream evaluations despite image-oriented discrepancies. Table 1 summarizes the hyperparameters used for this ROI detection, including the “random flip” enhancement, the input image scale of (1024, 1024), the stable gradient descent (SGD) optimizer, the learning rate of 1E-4, and the 50 training epoch, achieving a recurrence of 0.944 and an average accuracy (map) of 0.904.
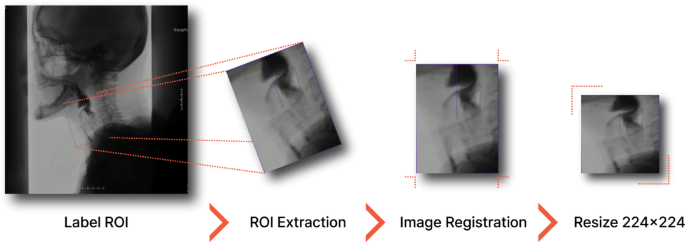
A workflow for extracting region of interest (ROI) images from a VFSS frame. ROI Areas of interest, VFSS Video fluid scopy study.
To reduce the burden of training and select select frames associated with deep learning model development, we extracted frames in which the filamentous bones with posterior to forward movement reached the highest position (high peak) and frames depicting frames that reached the lowest position (low peak) following the methodology used in previous studies.twenty one. The frame corresponding to the highest position (high peak) of the lobe was selected, reflecting the opening moment of the upper esophageal sphincter during engulfment, and the lowest position frame (low peak) indicates the phase after folding as before and after swallowing22,23. However, in previous studies, five frames were extracted at the highest and lowest positions of the Hyoid bones, whereas in this study, all frames corresponding to both peaks were extracted. Additionally, selection of these peak frames has been automated. Peak frame detection was performed using a dedicated classification model based on the EfficientNetB0 CNN architecture trained with Adam Optimizer, Relu Activation, and a batch size of 64.\(\:\ times \:\)224 pixels. It was included in the model architecture for normalization to prevent overfitting, dropouts, and batch normalization layers. This model was trained on 2,880 images (80%) and verified on 720 images (20%). The dataset is fully distributed in three classes: high peak frames (1,200; 33.3%), low peak frames (1,200; 33.3%), and non-peak frames (1,200; 33.3%). This model achieved a training accuracy of 96.6% and a validation accuracy of 94.3% to classify high, low and non-peak frames (Table 2). Using this model, 18,145 frames were extracted from VFSS data obtained from 1,467 patients.
Each image was labelled as “normal” (no penetration or aspiration), “penetration (entering the airway but remaining above the true vocal cords)” or “aspiration (contrast passing under the true vocal cords). Based on a single Bachelor of Science degree with more than 15 years of clinical experience in the diagnosis and management of dysphagia. The dataset included 14,535 training and 3,610 validation images. The dataset showed class imbalances: normal (63.9%), penetration (21.4%), and aspiration (14.7%), with the regular class being the most common. Synthetic minority oversampling technology (SMOTE) was applied to alleviate class imbalancestwenty fourGenerates a composite sample of a minority class by interpolating an existing instance with its k-nearest Neighbors. Class imbalances in medical image analysis can bias classification results, particularly in small datasets, leading to poor model performance. Small reduces this bias by increasing the sensitivity and accuracy of the rare but clinically significant dysphagia classifier.
A deep learning model for detecting penetration and aspiration in VFSS-based dysphagia severity classification adopted the Convnext-Tiny CNN architecture25,26. This model was optimized using an ADAMW optimizer and incorporated a sigmoid linear unit (SILU) activation function. Dropout and batch normalization were applied to alleviate overfitting and improve generalization. The DeepSpeed library integration facilitates larger batch sizes, accelerates training and improves model performance.
To ensure rigorous model training and evaluation and prevent data leakage, the data set was split based on a set of specific patient training (80%, 1,173 patients, 14,535 images) and validation (20%, 294 patients, 3,610 images). This patient-level outflow ensured that images of the same patient did not appear in both the training and validation sets, preventing overfitting, promoting model generalization, and ensuring fairness and process validity. Specifically, avoiding data leakage prevented artificially inflated performance metrics due to inherent similarities to patient data within the training and validation set.
Systematic hyperparameter optimization was performed to fine-tune the model architecture and training parameters. This process was used to assess performance metrics, including accuracy, accuracy, recall, F1 scores, and areas under the receiver operating characteristic curve (AUC ROC), providing a comprehensive assessment of the model's ability to detect penetration and aspiration. Rather than relying on a single iterative approach, multiple optimization tests were performed to evaluate different hyperparameter combinations and to identify configurations that achieved optimal performance. Figure 2 illustrates the model development workflow.
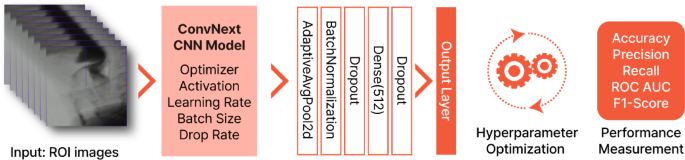
An overview of the deep learning model development process for detecting penetration and aspiration events in VFSS images. ROI Areas of interest, CNN Convolutional neural networks, rock Receiver operating characteristics, auc Area under the curve.
The diagnostic criteria were defined as follows: (1) Aspiration – classified when aspiration was observed in two or more images from a set of high- and low-peak images. (2) Penetration – We diagnosed the presence of 1 or fewer aspiration images and ≥2 penetration images. (3) Normal – Assigned when both the aspiration and penetration images were less than or equal to 1.
Statistical analysis
Statistical analysis was performed using Python and Scikit-Learn libraries. The diagnostic effects of deep learning classification models of dysphagia were assessed using accuracy, accuracy, recall, F1 scores, and AUC ROC. These performance indicators were calculated and reported at both the image level (each extracted image was analyzed individually) and at the patient level (all images extracted from the same patient were analyzed collectively) to ensure a comprehensive assessment of model performance.
Microaveraging and macroaveraging provide contrasting views on multiclass classification in deep learning. Microaveraging calculates global metrics by aggregating the contributions per sample and highlighting the performance of frequently used classes. Macro averaging calculates the average of metrics by class, weighs all classes evenly, and emphasizes performance on less frequent classes. Microaveraging typically evaluates overall performance, whereas macroaveraging evaluates class-balanced performance, particularly using unbalanced data.




