This section details the image dataset, analysis, DDRNet training, validation, and testing. A comparison of the proposed methodology with previous research studies is also provided for a better understanding.
Experimental setup of the analysis of DDRNet
In this study, the proposed DDRNet is experimented using a 64-bit Windows 10 operating system. Table 2 lists the computer configuration used for the DDRNet model training and testing. The proposed DDRNet architecture design and implementation were carried out using the deep network designer application of MATLAB 2022 academic version deep learning toolbox 14.4.
Dataset
The experimental analysis was performed through the publicly available database14 from the Kaggle website. The dataset includes 362 images of five types: Basophil, Eosinophil, Lymphocyte, Monocyte, and Neutrophil. Each variety contains images of the count, three images of class Basophil, 80 images of class Eosinophil, 35 images of class Lymphocyte, 25 images of class Monocyte, and 220 images of class Neutrophil, as shown in Fig. 5.

Data augmentation
For easier processing, dataset images were converted to jpeg format and resized to 224 × 224×3. The images were divided into training, validation, and test sets before augmentation was performed. Augmentation techniques were used during the data preprocessing stage to handle class imbalances, especially for under-presented cell types. The augmentation process involves horizontal flipping, rotation to 10 degrees, zooming with a scale of 2, and random contrast enhancement. Since cropping might result in losing some significant information, cropping has not been performed in this work. This augmentation strategy enables the generation of a more balanced and diverse dataset, facilitating effective training and evaluation of DDRNet across all cell types.
Horizontal flipped image \(I_{HF}\) is generated from original Image \(I_{o}\) by flipping the image horizontally referring to image width ‘\(w^{\prime}\) and the transformation is given as Eq. (13).
$$I_{HF} \left( {m,n} \right) = I_{o} (w – (m,n))$$
(13)
When the model sees both original and flipped images during training, it becomes more adept at recognizing leukemic abnormalities regardless of their orientation in real-world diagnostic scenarios. This helps the model learn robust features that are invariant to horizontal orientation.
Rotated image \(I_{RO}\) are generated by rotating the original image \(I_{o}\) to 10 degree clockwise and the transformation is shown in Eq. (14), where is \(\theta\) the rotation angle in radians (10 degrees converted to radians), and m,n are the coordinates of the pixel in the original image \(I_{o}\).
$$I_{RO} \left( {m,n} \right) = I_{o} \left( {\cos \left( \theta \right)m + \sin \left( \theta \right)n, – \sin \left( \theta \right)x + \cos \left( \theta \right)y} \right)$$
(14)
When the model encounters rotated images during training, it becomes better at handling real-world scenarios where cell structures may appear at different angles. For example, a rotated leukemic cell image should still exhibit recognizable patterns. Rotated images help the model learn features that are invariant to rotation.
Zoomed image \(I_{ZO}\) is generated by zooming the image to a scale of 2and the transformation is given in Eq. (15) where m and n are the coordinated of the pixels in the original image \(I_{o}\).
$$I_{ZO} \left( {m,n } \right) = I_{o} \left( {\frac{m}{2},\frac{n}{2}} \right)$$
(15)
Zoomed images allow the model to learn features that remain consistent across different magnifications. Real-world blood cell images contain cells of varying sizes. Augmenting the dataset with zoomed images ensures the model adapts to different cell scales.
Random contrast enhancement increases the contrast of the image in a random manner. This process entails modifying the pixel values to amplify the disparity between the brightest and darkest components in the image, thus enhancing its overall contrast. Clinical blood cell images exhibit diverse contrast levels due to variations in sample preparation and imaging techniques. Varying contrast levels simulate different imaging conditions (e.g., variations in staining, lighting, or equipment settings). The model learns to recognize blood cells even when the lighting varies significantly. By augmenting the dataset with contrast-enhanced images, the model becomes more resilient to such variations.
Sample augmented images of four classes are shown in Fig. 5. A total of 16,249 images were generated across four classes using the augmentation technique, and the distribution is as shown in Fig. 6. Among these, 12,515 images were utilized for training and validation, consisting of 3,133 eosinophil images, 3,109 lymphocyte images, 3,102 monocyte images, and 3,171 neutrophil images. The remaining 3,734 images were reserved for testing, which comprised of 936 eosinophil images, 931 lymphocyte images, 927 monocyte images, and 940 neutrophil images.
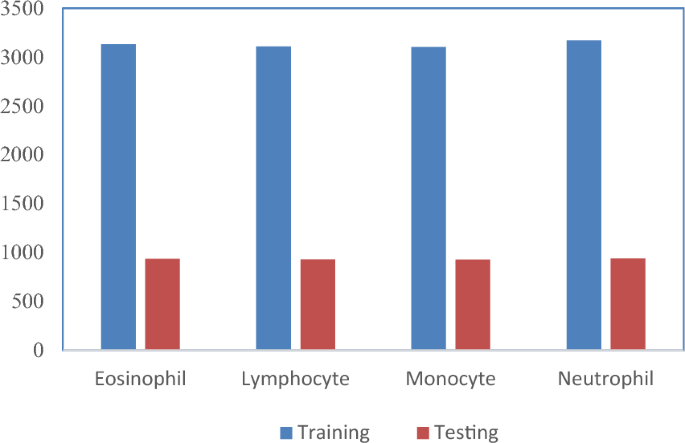
Training and testing image count distribution after the augmentation process.
Hyperparameter tuning
The DDRNet models were fine-tuned to enhance their performance by adjusting various hyperparameters, such as the dropout rate, batch size, epochs, learning rate, and optimization units implemented in the gradient. This study used a gradient optimizer named Adam and a dropout factor of 20%. Adam was the best gradient optimizer, and a dropout of 20% of features was found to reduce overfitting to a greater extent, thus improving the impact of generalization in most processes of the DDRNet models. The customized DDRNet model was trained over 30 epochs using the Adam optimizer and a learning rate of 0.01. The learning rate schedule was also thought to be constant. The DDRNet model was trained for 2040 iterations.
DDRNet model feature analysis
Figure 7 describes the features acquired with the help of the inner layers of DDRNet during the training phase. Gradient-weighted Class Activation Mapping (GradCAM) provides insights into the regions of the input images that contributed most to the model’s classification decision. Visualization of features through GradCAM helps clinicians and researchers interpret the decision-making process of DDRNet, offering transparency and aiding in understanding the model’s focus on relevant features. The intermediate layers’ visual feature inspection demonstrates the resilience of retrieved features that retrain the model. This depiction significantly enhances the understanding of how the DDRNet model internally interprets the multiclass blood image. The outcomes of applying 32, 3 × 3 layer1 convolutional filters are visualized in the first image of Fig. 6. The adjacent image shows the results of applying layer six convolution filters. DRDB, GLFEB, and CSAB blocks capture the edges in this instance are more robust than they were in the prior feature image. Further convolution blocks help to extract even more class-specific and inter-class discriminating feature maps from the image. All 64 image variations reflect a distinct feature and contribute to the fully connected convolution layers utilized for the classifications. Figure 7 confirms the exceptional ability of DDRNet to extract features that discriminate images of eosinophil, lymphocyte, monocyte, and neutrophil. The fact that distinct regions are displayed in different colours demonstrates the feature block’s capacity to discriminate between images belonging to the four classes of Leukemia.
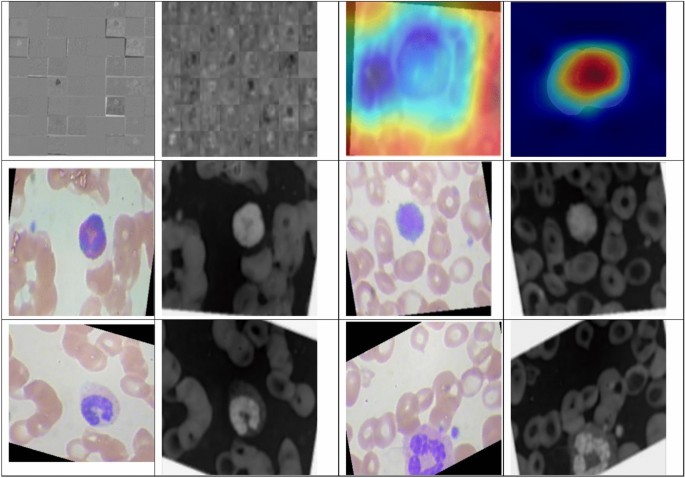
Visualization of the convolution layer features.
Evaluation metrics
Evaluation measures are essential for evaluating the effectiveness of a trained model. The performance metrics used to evaluate the effectiveness of the DDRNet model included accuracy, precision, recall, F1-score, and confusion matrix. The testing accuracy was ascertained by predicting the result of the DDRNet model on the test data. A confusion matrix was utilized to evaluate the performance of each class in the proposed DDRNet model. To assess the accuracy of testing, the results obtained during the training phase were compared with the test set obtained from the partitioned dataset. Equations (16–19) provide the details of the evaluation metrics:
$$Accuracy = \frac{D}{D + E + F + G}$$
(16)
$$Precision = \frac{D}{D + E}$$
(17)
$$Recall = \frac{D}{D + W}$$
(18)
$$F1 Score = \frac{2*Precision*Recall}{{Precision + Recall}}$$
(19)
where “D” represents the true positive result, “E” is the true negative result, “F” is the false positive result, and “G” is the false negative result. When the model accurately predicts the positive class, the results are actual positives (D). When the network successfully estimates the negative class, the results are referred to be True Negative (E). A false positive (FP) or false negative (FN) occurs once the model incorrectly estimates the positive class as the negative class or the negative class as the positive class. Table 3 exhibits the evaluation metrics of the four class classifications using the DDRNet Model, including precision, recall, F1 score, and Mathews Correlation Coefficient (MCC).
The confusion matrix systematically captures and quantifies the model’s predictions across different cell types. Precision, recall, and F1 score metrics were calculated for each cell type to better understand the model’s performance in terms of correct predictions (D and E) and false predictions (F and G). The architecture of DDRNet is built to extract intricate features from input images. This feature enables the model to identify subtle patterns and characteristics that can suggest particular cell types, even when the differences are less pronounced. Augmenting the data through adjusting lighting, rotation, and scaling techniques enhances the model’s capacity to generalize across varied structures. The augmentation of training data diversity proves effective in addressing borderline cases. The utilization of explainability tools aids in identifying the features, which is especially beneficial in situations characterized by ambiguity.
Model training and validation
The classification accuracy of the DDRNet model was evaluated by training and testing it using the augmented data. The model architecture comprises three phases: a deep residual dilated block, followed by blocks that enhance channel and spatial features and global and local feature creation. The ADAM optimizer was employed to train the model over 30 epochs with 2040 iterations. The accuracy and loss curves for each fold of the training and validation sets are illustrated in Fig. 8. The training loss curve decreases as the number of training epochs increases, and similar behavior in the opposing trend is seen for the training accuracy. After 200 iterations, the positive slope’s change rate gradually slows and flattens out. Group normalization was employed during the DDRNet training process, and a learning rate of 0.01 facilitated better and quicker convergence. The model reached saturation more quickly, and the training accuracy improved over time. The DDRNet model incorporates BN and DO techniques to reinforce the stability and efficacy of ALL classification. These methods contribute to sustaining a uniform input distribution across diverse layers and mitigating the risk of overfitting.
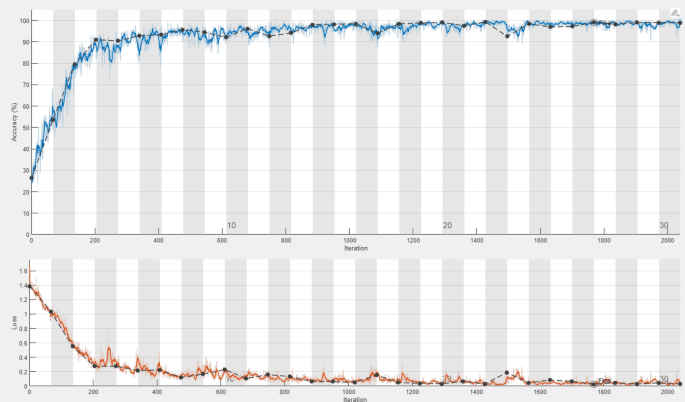
The DDRNet ‘s learning process concerning models’ loss, number of epochs and accuracy .
Although the validation accuracy increased over time, it changed throughout training. DDRNet has the most straightforward architecture of all the listed custom models. DDRNet is capable of learning a variety of characteristics that are pertinent to microscopic cells. The 2000 iteration of 30 epoch execution time is 161 min 20 s. The architecture of DDRNet is computationally more straightforward, with fewer trainable parameters, and more compact taking less time to train.
Figure 9 illustrates the prediction score of DDRNet for eosinophil, lymphocyte, monocyte, and neutrophil images. The DDRNet model achieved the highest prediction scores for eosinophil (1.00), monocyte (1.00), neutrophil (1.00) and slightly lower for lymphocyte (0.99). The high prediction score of DDRNet reveals a greater level of confidence in its predicted class. The higher prediction score for eosinophil, monocyte, neutrophil and lymphocyte imply a higher likelihood of ALL disease, and it can aid the clinical decision to proceed with further treatment based on the prediction.
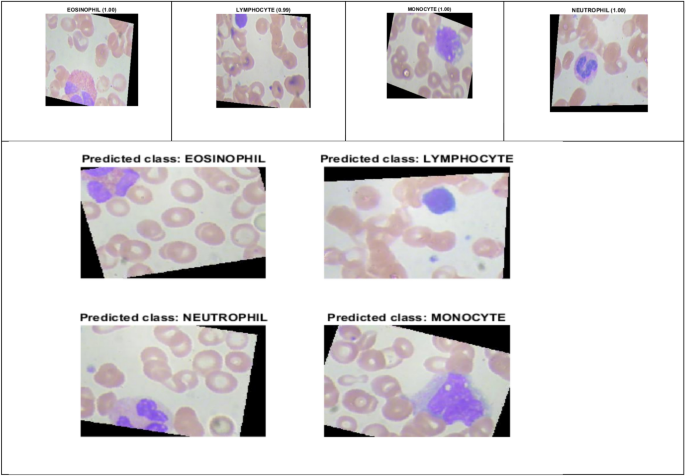
DDRNet Model Prediction Score.
Analysis of DDRNet performance
Deep residual dilated block (DRDB) utilized residual connections to improve gradient flow and decreased the depth of the network by dilated convolution network to reduce the model complexity and trainable parameters. Building an accurate and efficient model for multi-class classification is a complex task that demands careful consideration of appropriate features that lower the ambiguity among the eosinophil, lymphocyte, monocyte, and neutrophil images. It is comparatively more challenging than ALL binary classification. In the context of multi-class ALL classification, data augmentation has demonstrated its effectiveness in enhancing the accuracy of DDRNet model. Through the utilization of data augmentation techniques, the DDRNet model developed the ability to adapt to variations in the input images, including variations in orientation, scale, and intensity. As a result, the DDRNet model achieved better generalization, and ultimately, better performance on test data. Furthermore, data augmentation also tackled the issue of imbalanced data in ALL classification. By increasing the samples in the underrepresented class through augmentation, the DDRNet model learned to detect the unique characteristics that differentiate the four classes, resulting in enhanced classification accuracy. The Global and local feature enhancement block (GLFEB) selects the highest activation in each feature map using global average pooling, where the average activation in each feature map is computed. Group normalization (GN) enhanced the performance of DDRNet by reducing the internal covariate shift that arises during the distribution of ALL input images to layer changes during training. Data augmentation and GN increased the robustness to input variations and reduced the sensitivity to hyperparameters such as learning rate and weight decay. GN has been shown to be more effective with small batch sizes to capture the fine-grained details within the eosinophil, lymphocyte, monocyte, and neutrophil images. GLFEB combined global and local features and captured the overall context of ALL images and their specific details, leading to more discriminative and accurate feature representations. DDRNet model captured more complex inter and intra-class patterns and relationships between the features using the CSAB block, leading to better performance.
Tables 4 and 5 illustrate the efficiency of the recommended DDRNet model with various residual networks such as ResNet 18, ResNet 50, and ResNet 101 based on metrics including MCC, recall, precision, F1-score, accuracy, and execution time. The results demonstrate that the recommended model surpasses the other residual networks with regard to accuracy. Moreover, the proposed model takes only 161m19sec for computation, whereas ResNet 18, ResNet 50, and ResNet 101 require 399m17s, 1203m21s, and 1679m39s, respectively, indicating superior computational efficiency. Notably, the proposed model’s computational time is almost 90% less than that of ResNet 101. Moreover, the proposed model achieves an MCC, precision, recall, and F1-score of 1, whereas ResNet 18 and ResNet 101 show variations in performance values.
The confusion matrix generated for the proposed DDRNet is compared with the residual network to show the impacts of the classification of leukemia. The confusion matrices obtained for all the models are shown in Table 5.
Convolutional layers acquire hierarchical features that include local patterns and structures that are improved by translational invariance, enabling the recognition of diverse cell morphologies. The model extracts and learns features at different levels, enabling it to capture both subtle and prominent morphological characteristics associated with each cell type. Augmentation techniques, such as rotation, scaling, and contrast enhancement, are applied during training to introduce variations in cell morphology. This ensures that the model is exposed to a diverse range of morphological features, enhancing its ability to generalize across different cell types. Group normalization stabilizes and normalizes these properties, enabling the model to effectively handle variances in cell morphology across distinct cell types like eosinophils, lymphocytes, monocytes, and neutrophils. The utilization of the deep residual dilated block enables the network to effectively adjust to various spatial scales and accurately capture intricate class-specific features included in the cell images. Dilated convolutions aid in expanding the receptive field, which is essential for detecting patterns at different levels. By allocating varying weights to distinct channels according to their significance, channel attention enables the model to concentrate on the most illuminating characteristics for cell type differentiation. The model highlights crucial regions and discards unnecessary ones with the use of spatial attention. Simultaneously capturing both spatial and channel-wise information, makes it capable of capturing intricate correlations among the data. The attention mechanism has the potential to be useful in identifying tiny details and patterns that differentiate various cell types.
Ablation study
The proposed DDRNet model’s various blocks were examined using an ablation study to determine their effects on accurately classifying ALL. It is clear that adding the various blocks, as stated in Table 6, causes the model’s overall performance to improve steadily. Accuracy, Precision, Recall, F1-score, and Confusion matrix of the ALL images are the metrics used to reveal the efficiency of the suggested DDRNet classification architecture. Figure 10 depicts the impact of the ablation study on the suggested blocks for classifying ALL.
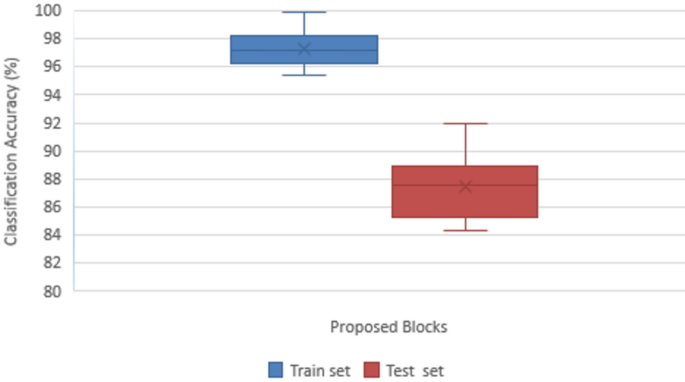
Impact of ablation study on proposed blocks in ALL classification.
Convolution block is considered the baseline of the proposed model, which attained a classification accuracy of 84.30%, comprising only a convolutional layer followed by a max pooling layer and a group normalization, which normalizes to global features for enhanced classification. Group normalization simultaneously normalizes the channel and batch dimensions by separating and normalizing the channels into groups. Compared to batch normalization, group normalization is less reliant and stable on huge mini-batch sizes, making it advantageous in situations involving smaller batch sizes. Group normalization eliminates the dependence on the mini-batch and batch size sensitivity during the training phase.
The DRDB block is implemented along with the baseline convolution model that attained a classification accuracy of 85.23%. The conventional residual block between the layers retained more original information that extracted the general feature maps for improved accuracy of test data. The generated features are enhanced using CSAB embedded in the Convolution and DRDB block where global averaging and max-pooling are employed, resulting in excellent feature representation and improving accuracy to 86.14%.
Similarly, another study with the GLFEB module is performed with the baseline convolution layer, resulting in a better accuracy of 87.56%. The resultant features are subjected to the channel and attention model for enhanced feature representation, achieving an accuracy of 87.98%. The baseline model is connected with DRDB, GLFEB, and CSAB blocks for testing and training the ALL dataset since both blocks showed improvement in classification accuracy. The internal covariate shift developed during the distribution is minimized through global normalization as the GLFEB finds the highest activation in each feature map. GLFEB generates more accurate and discriminative feature representations by combining global and local features, capturing the overall context of ALL images and the individual details therein. Using the CSAB block, the DDRNet model was able to capture associations and inter- and intra-class patterns more intricately, improving accuracy to 91.98%.
The proposed DDRNet has been thoroughly trained and assessed using the Acute Lymphoblastic Leukemia (ALL) database49 in addition to the multi-class Kaggle dataset14. The ALL training data consisted of 73 subjects, including 47 with ALL (7272 cancer cell images) and 26 normal individuals (3389 images), totaling 10,661 cell images. The test set comprised of 2586 cell images.The experimental findings demonstrate that on the ALL dataset, the DDRNet obtained a remarkable accuracy of 99.86% and the training and testing accuracy plot is shown in Fig. 11. This additional validation demonstrates the model’s robustness in handling a variety of data distributions and shows that it can generalize effectively across diverse datasets.
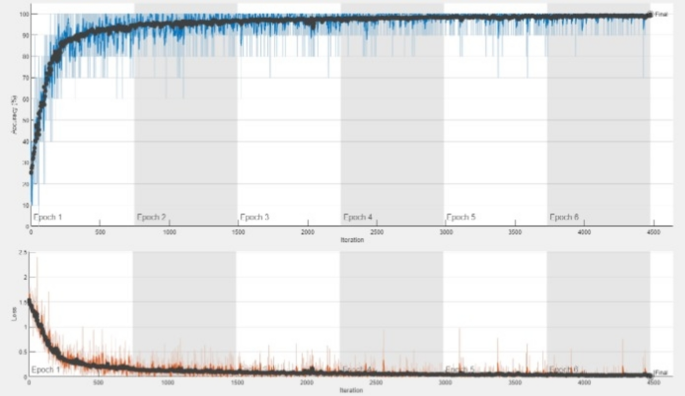
Performance of the proposed model in ALL49 dataset.
Performance comparison with existing leukemia classification research models
Table 7 presents a comparative analysis of the accuracy and efficacy of the DDRNet architecture with cutting-edge models for ALL classification. Karthikeyan et al.7 analyzed microscopic images by extracting GLCM features and classifying them using random forest, achieving an accuracy of 90%. Neoh et al.20 utilized an intelligent decision support system with machine learning models and achieved an accuracy of 96.72%. Rehman et al.10 achieved an accuracy of approximately 97.78% using convolutional neural networks for a dataset from Pakistan. Additionally, YOLOv4 was used to analyze ALLIDB1 and C NMC 2019 datasets and attained an accuracy of 96.06%32. The proposed DDRNet model achieved a training accuracy of 99.86% for blood cell dataset and 99.86% accuracy for Leukemia Dataset49 with fewer parameters and less computation time compared to the ML and DL approaches.




