Application of the FALCON-on-the-fly calculator to a MD simulation
The concept of the OTF calculator is based on the ability of a ML model to predict not only the energy and forces of a structure but also the uncertainty of these values. In this work, the GPR model implemented in AGOX is used for this purpose (see Methods section for details). If the predicted uncertainty of the energy ΔE exceeds a customizable accuracy threshold ϵ, an exact calculation (e.g. using DFT) is performed, and the model is updated and trained with energy and forces of the structure. Otherwise, the structure is accepted. In MD simulations, this means that the next MD step is computed directly using the forces predicted by the ML model (Fig. 1).
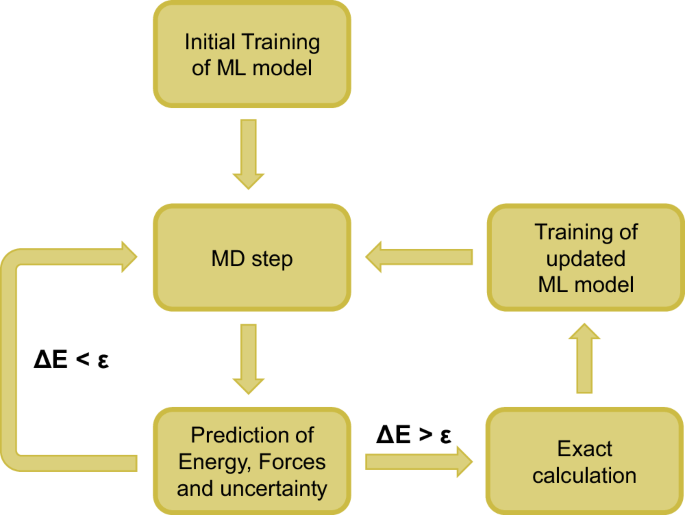
After the initial training of the ML model energy, forces and the energy uncertainty ΔE is predicted by the model. An exact calculation and training of the ML model is only required if ΔE exceeds the accuracy threshold ϵ.
A simple input example is given below. Note however, that much more parameters can be used to control the FALCON OTF calculator and the ML model.
-
from ase.build import bulk
from ase.calculators.emt import EMT
from ase import units
from ase.io.trajectory import Trajectory
from ase.optimize import QuasiNewton
from ase.io import read
from falcon_md.otf_calculator import FALCON
from falcon_md.models.agox_models import GPR
###############################################
from ase.md.langevin import Langevin
# General Setup
T = 600 # Temperature in K
accuracy_e = 0.10 # Accuracy Threshold (ε) in eV
# Create a Pt bulk, (3x3x3) fcc structure
atoms = bulk(‘Pt’, ‘fcc’, a=3.92, cubic=True).repeat((3, 3, 3)) # Setup ASE atoms object
atoms.rattle(0.1) #rattle the positions slightly
exact_calc = EMT() # Calculator for exact calculations during OTF training.
###############################################
# Geometry optimization with EMT potential to generate intial training structures
atoms.calc = exact_calc
qn = QuasiNewton(atoms, trajectory=’opt.traj’)
qn.run(0.001, 10)
training_data = read(‘opt.traj@0:’)
###############################################
# Setup of the minimal FALCON-OTF-Calculator
atoms.calc = FALCON(model = GPR(atoms), # The default AGOX GPR model is used
calc = exact_calc
training_data = training_data
accuracy_e = accuracy_e)
# Setup of the MD Simulation
dyn = Langevin(atoms, 1 * units.fs, temperature_K=T, friction=0.002, trajectory=’Md.traj’)
###############################################
# Now run the OTF-MD Simulation!
dyn.run(1000) # 1000 MD steps
As a demonstration, a MD simulation of an icosahedral Pt55 cluster using an effective-medium theory (EMT) potential43 is employed as implemented in ASE. In particular, Pt-based nanoalloys have attracted great interest due to their applications in heterogeneous catalysis44. Especially, they are useful as empirical potentials exist which are well benchmarked against DFT43,45, which is why they have been used in other studies introducing ML potentials46. For the following example, the usage of an EMT potential instead of DFT makes it easy for every reader to reproduce the data shown in this chapter on their personal computer. All scripts for reproducing these data can be found in the Supporting Information.
The structures for the initial training of the ML model are generated from a geometry optimization of the cluster. The subsequent MD simulation is conducted at a temperature of 600 K with a time step of 1 fs and an accuracy threshold of 0.125 eV is used to determine whether an EMT calculation is required. Initially, the ML model requires frequent retraining, with 53 training steps occurring within the first 1000 time steps (1 ps). As the simulation progresses, the number of required EMT calculations decreases significantly (Fig. 2). After 120,000 MD steps (120 ps), the energy of the system stabilizes, oscillating around a value of 0.51 eV/atom. For the remaining 880,000 MD steps, only 31 additional training steps with exact calculations are necessary. This shows that after 153 training steps in the first 120,000 MD steps, the model serves as an accurate surrogate for the potential and only requires minimal further training. This is further supported by plausible results as the initially well-ordered icosahedral cluster undergoes structural transformation into a more disordered, spherical shape after 30 ps at 600 K, which is kept until reaching 1 ns simulation time.
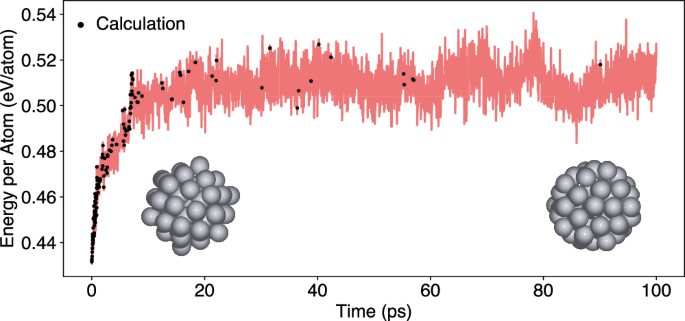
If the predicted energy uncertainty ΔE of the ML model exceeds the accuracy threshold ϵ of 0.125 eV, an EMT calculation is performed and the ML model is trained (black dots).
MD simulations with varying accuracy thresholds ϵ
To demonstrate the importance of a sensible selection of the accuracy threshold ϵ, MD simulations of the Pt55 cluster at 600 K are performed with various thresholds ranging from 0.050 to 0.200 eV. For accuracy thresholds of 0.100 eV and above, the ML model initially requires frequent retraining. However, after a certain number of training steps, a stable state is reached in which only occasional training steps are necessary (Fig. 3a).
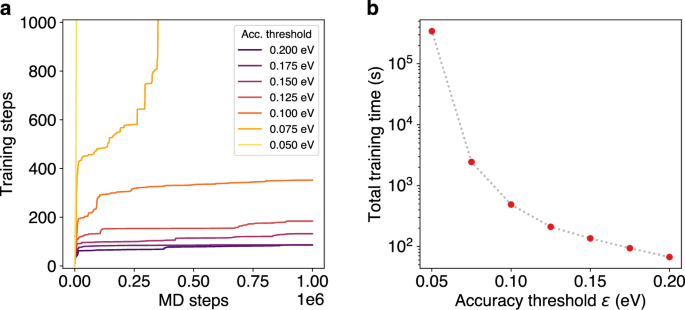
a Number of required training steps over the course of 1 ns MD simulations of a Pt55 cluster at 600 K with different accuracy thresholds ϵ. b Total training times (logarithmic scale) required for 20,000 MD steps (20 ps) during the MD simulation with different accuracy thresholds ϵ.
The exact number of training steps required for 1,000,000 MD steps (1 ns) depends on the specific threshold, e.g. at a threshold of 0.100 eV, more than 300 training steps are needed, while this number drops to 86 for a threshold of 0.200 eV. In this case, accuracy thresholds under 0.010 eV require a very high number of training steps and never reach a stable state of the model. This significantly slows down the MD simulation, since the duration of a single training step depends on the size of the ML model, which grows over the course of the simulation. Therefore, the total training time does not increase linearly with the number of training steps. As a result, very small accuracy thresholds lead to extremely high total training times. For example, at a threshold of 0.075 eV, the total training time required for 20,000 MD steps is an order of magnitude higher than at 0.100 eV (Fig. 3b). In such cases, the benefits of OTF training compared to conventional AIMD are reduced, as the computational cost of frequent training steps outweighs the time saved by avoiding exact calculations.
Naturally, the reduced number of training steps at high accuracy thresholds decreases the prediction accuracy of the ML model. To analyze this effect, the energies of the OTF-MD trajectories were recomputed using EMT, and the root mean squared error (RMSE) of the energy was calculated. As expected, the RMSE increases with larger accuracy thresholds (Fig. 4). However, deviations from this trend are visible. Most notably, the RMSE at an accuracy threshold of 0.125 eV is significantly lower than at a threshold of 0.100 eV. This deviation can be attributed to the nature of MD simulations, where the initial velocities of the atoms are randomly sampled from a velocity distribution. Therefore, each of the conducted MD simulations is unique, and the deviations in prediction quality are also influenced by the initial velocities. For a more precise statement about the correlation between the accuracy threshold and RMSE, it would be required to either compute the same trajectory with various accuracy thresholds or to perform multiple MD simulations to obtain statistically significant results. However, for all accuracy thresholds, the error lies in the region of RMSE(E) ≤ 2ϵ.
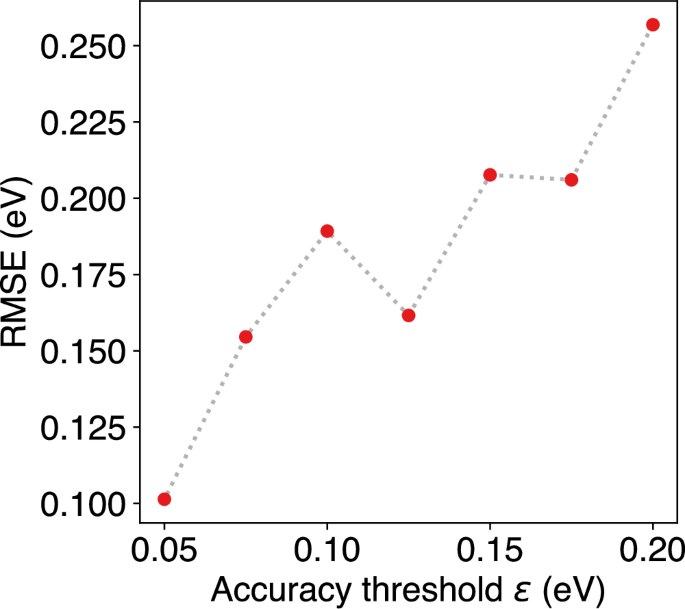
Root mean squared errors (RMSE) of the energy during the MD simulations of a Pt55 cluster at 600 K with different accuracy thresholds ϵ.
As a general guideline, an accuracy threshold of 0.10 eV offers a reasonable balance between prediction accuracy and computational cost for most MD simulations. However, this value may be modified according to the specific needs of the user, such as higher accuracy or reduced simulation times. The lower the threshold is chosen, the more often an electronic structure calculation needs to be performed, which should be the computational bottleneck in most cases. However, an alternative approach to directly choosing a low accuracy threshold is to run a first simulation with a higher value and then add additional training data based on their uncertainty as described in section “Diffusion of Water in a Carbon Nanotube”.
MD simulations of different-sized systems
In addition to the choice of the accuracy threshold, the size of the simulated system also impacts training times. To investigate this effect, MD simulations at 600 K are conducted for three Pt clusters of different sizes (Pt55, Pt147, Pt561) with an accuracy threshold ϵ of 0.100 eV. The results show that the increase in training times does not scale linearly with the increasing system size (Fig. 5). Linear fits for each MD simulation with different cluster sizes reveal that, while the Pt561 cluster is around ten times larger than the Pt55 cluster, the increase in training time per training step is approximately 80 times higher. Note, for this comparison, we completely neglect the calculation time for the energy and forces of the calculation in the real potential. In case of EMT, as used for these calculations, the time difference is neglecteable but in cases of using DFT, e.g. using the PBE exchange-correlation functional these methods formally scale like \({\mathcal{O}}({N}^{3})\) with respect to system size N.
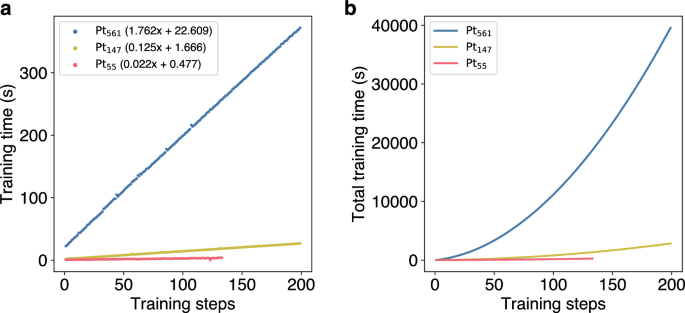
a Evolution of training times of the ML model over the number of training steps for MD simulations of different-sized Pt clusters at 600 K. Linear fits added for each size indicate that increase in training times does not scale linearly with the increase in system size. b Total training times over the number of training steps during the MD simulations of different Pt clusters.
Naturally, the increase in the training time for every single training step does impact the total training times. Training of the first 100 training steps takes 156 s and 803 s for the Pt55 and Pt147 clusters, respectively. For the Pt561 it takes 11,666 s. With an increasing number of training steps, this difference grows even further, demonstrating the problems of this simple OTF calculator, when dealing with larger systems (For completeness: All calculations have been performed on an Intel Xeon 8362 CPU using 16 cores. The GPR model as implemented in AGOX is parallelized via RAY.)
MD simulations at different temperatures
A factor that can also impact the training times of OTF-MD simulations is the temperature. At elevated temperatures, the kinetic energy and the atomic movement in the simulated system increase. This results in structures that differ more strongly from each other, increasing the number of necessary training steps to correctly describe the potential. This effect is evident when comparing MD simulations of the Pt55 cluster at different temperatures. At 300 K and 600 K a stable state is reached after approximately 10,000 MD steps and only few additional training steps are necessary (Fig. 6). However, this is not the case at 900 K, where 3500 training steps are necessary for the first 35,000 MD steps. This translates to a significant difference in total training times. While the simulation at 600 K reaches 200,000 MD steps in 2735 s, at 900 K only 13,000 MD steps are computed in the same amount of time.
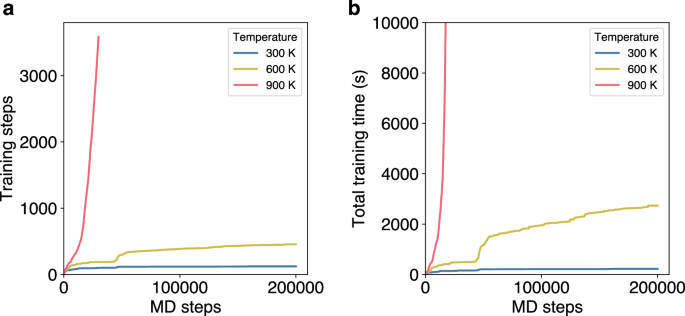
a Number of required training steps over the course of 20 ps MD simulations of a Pt55 cluster at different temperatures. b Total training times over the course of the MD simulations at different temperatures.
This demonstrates that simulations involving significant structural changes, such as those induced by high temperatures or in general materials showing high structural disorder, result in a large training dataset. For many ML models, such as the here used GPR this poses to be a challenge, since the computational cost of GPR scales cubically with the size of the dataset47. Therefore, a large set of training data significantly increases the computational cost, when using GPR. Solutions to this issue are discussed in the following sections.
Clustering of training data—the subset of data approach
A simple way of decreasing the training time is to restrict the number of times the model is trained and only train in batches e.g. every nth time after an exact calculation has been performed. Although this approach might work well in cases, the model is already pre-trained well, we have realized that this approach might lead to a higher number of exact calculations. This may especially be found in cases where non-well-explored regions of the potential energy surface are explored and instant retraining of the model does not improve the uncertainty in this area directly (see Supporting Information).
In addition, the restricted optimization of hyperparameters can reduce the computational cost of the OTF calculator. However, it only partially solves the problem of very long training times for large systems high temperatures or strongly disordered systems. (c.f. SI).
Another commonly used approach is the use of a SparseGPR, a technique used to make GPR computationally feasible for large datasets by approximating the model with a smaller set of so-called inducing points. These act as representatives for clusters of training data, while other similar data points are neglected. This sparsification is helpful as will be discussed shortly in section “Melting of aluminum using density functional theory”. However, especially if highly expensive computational methods, like DFT calculations or high-level post-HF ab initio correlation methods like CCSD(T) have been used to create the training data, one wants to make use of all of it.
A possible solution is clustering of the training data into subsets and operating multiple smaller ML models instead of a single very large model. A similar approach has been used by Kisi et al. in the area of rainfall runoff modeling48 and might be useful in other applications of active learning in general. To achieve smaller subsets of the training data in FALCON, first an average model size is defined. If the total number of training structures reaches this value or its integer multiple, a new ML model is added. All structures are then distributed to the models using k-means clustering (see sec. “k-Means Clustering”), where k is the number of active ML models. After this clustering all ML models are trained (Fig. 7).
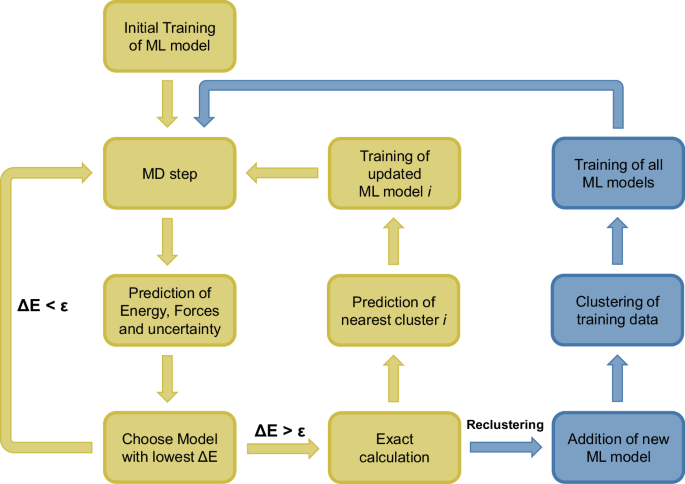
The original algorithm is extended by the prediction of the nearest cluster for every structure that is added to the training data. When the number of training structures reaches the average model size or its integer multiple a new ML model is added and all structures are reclustered (blue part).
During the MD simulation, each structure’s energy, energy uncertainty and forces are predicted by all active ML models. The model with the lowest uncertainty is chosen and compared to the accuracy threshold ϵ, to determine if an exact calculation is necessary. If this is the case, the energy and forces of the structure are computed e.g. by means of DFT and added to the ML model, whose cluster mean it is closest to. Only this single ML model has to be retrained and not the whole set of models. The limitation of cluster size avoids the formation of very large ML models that slow down the simulation. Therefore, clustering can significantly reduce training times (Fig. 8).
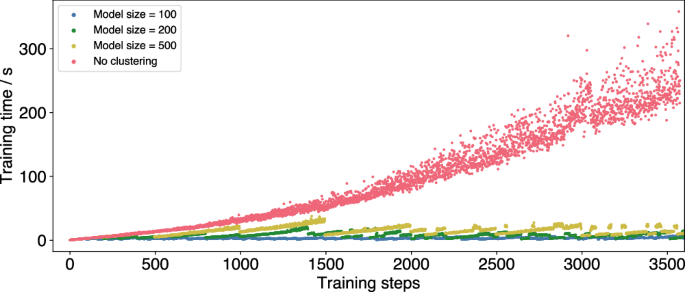
MD simulations of a Pt55 cluster at 600 K with no clustering of training data (red) and clustering with different average model sizes (yellow, green, blue).
The average model size parameter is highly relevant for the performance of the OTF calculator. While a smaller model size reduces the training times, it negatively impacts the prediction accuracy. This leads to an increased number of required training steps for small model sizes (Fig. 9). However, MD simulations of the icosahedral Pt55 cluster indicate that clustering generally provides a significant time advantage over the standard OTF calculator. Nevertheless, the total time required for predictions and training strongly depends on the average model size. For the simulation of the Pt cluster, the lowest total time is achieved with an average model size of 200, whereas both smaller (100) and larger (500) sizes lead to an increase in total time (Fig. 9). While the larger models take more individual training time, the smaller models need frequent retraining that the advantage of shorter training times is neglected. This shows that optimization of the average model size is essential for the use clustering in the OTF calculator.
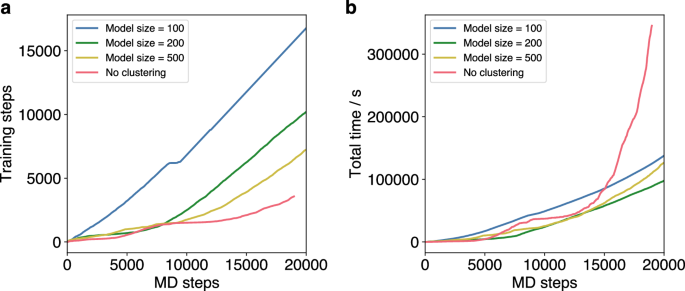
a Number of required training steps over the course of 20 ps MD simulations of a Pt55 cluster at 600 K with different average model sizes (blue, green, yellow) and without clustering (red). b Total time of trainings, predictions and clusterings over the course of the OTF-MD simulations with different average model sizes (blue, green, yellow) and without clustering (red).
For further analysis of the impact of different model sizes, a training dataset of 2500 structures generated during a FALCON-OTF run of Pt55 cluster at 600 K for 10 ps was clustered into subsets using various model sizes. This results in various FALCON models with different numbers of clusters, ranging from one single model to 100 small models. With each number of clusters, the energies of an MD trajectory were predicted using and the corresponding errors were computed. The results indicate that the overall RMSE remains consistent when using between one and 20 clusters (Fig. 10). While the RMSE of some individual models becomes large when clustering into more than five models, the combination of multiple models is sufficient to achieve an overall RMSE comparable to that obtained with a single large model. However, when the individual clusters become too small, even the combination of them cannot predict the energies with sufficient accuracy. This is evident for runs with 50 and 100 clusters, which result in much higher RMSEs than runs with fewer clusters. In these cases, clusters contain fewer than 100 training structures, demonstrating that model sizes below 100 are not advisable without compromising prediction accuracy.
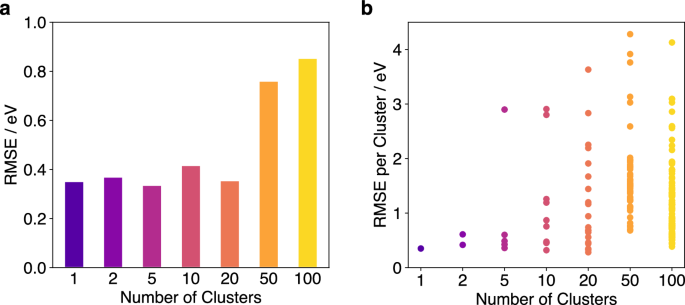
a Total RMSE of the energy over a 10 ps MD trajectory of a Pt55 cluster for various numbers of clusters for the FALCON predictions. b RMSEs of the energy for each individual cluster over the course of the MD trajectory.
For better understanding of the physical differences among structures in the individual clusters, the training dataset of 2500 structures was divided into five clusters A to E. For each structure their similarity to the initial icosahedral Pt55 cluster and to a spherical structure from the late stages of the MD simulation was computed. The Smooth Overlap of Atomic Positions (SOAP)49 descriptor was used to describe the local environments in each structure. Further, the SOAP Average Kernel,50 as implemented in the DScribe package,51,52 was used to compare the averaged local environments between pairs of structures on a scale from 0 to 1. This metric provides a clear visualization of the different compositions of the clusters. (Fig. 11).
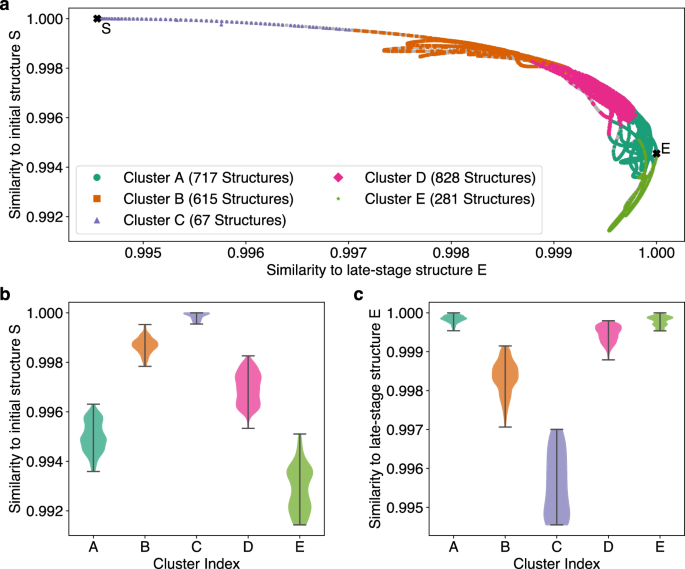
a Similarity of structures in a FALCON-OTF training dataset, divided into five clusters, to both the initial icosahedral Pt55 cluster (S) and a spherical structure from the end of the simulation (E). Light gray points in the background represent all structures from the MD trajectory that were not included in the training dataset. b Distribution of structures in each cluster with respect to their similarity to the initial structure S. c Distribution of structures in each cluster with respect to their similarity to the late-stage structure E.
It is evident that cluster D consists of more ordered structures similar to the initial structure S, while clusters A and E contain disordered structures from later stages of the simulation. This demonstrates that each cluster effectively describes a subset of structures from a specific part of the potential energy surface, resulting in the combination of all clusters providing a full representation of the entire training dataset. While the five clusters differ strongly from each other, increasing the number of clusters results in some clusters becoming very similar and describing the same structural region (see Supporting Information). Therefore, although from a mathematical point of view a higher number of clusters might give a reasonable fit, whereas from a physical/chemical point of view a lower number of clusters might be reasonable to get an intuitive fit of specific regions of the PES.
However, the representation of the data in Fig. 11 clearly shows the general benefit of the active learning approach, where only the regions of the PES are sampled in the real potential, which are actually needed for the MD simulation.
Overall, clustering the training data into subsets and distributing it across multiple smaller ML models, leads to a significant reduction in computational cost during OTF calculation. Since the required training time increases more than linearly with the number of training structures, the distribution on multiple models avoids the formation of a single very large and extremely slow ML model. Therefore, clustering provides the foundation for more complex OTF-MD simulations of large systems or at high temperatures, which could not be performed in a reasonable amount of time with the classical OTF calculator.
Melting of aluminum using density functional theory
To demonstrate the ability of the OTF-MD calculator to adapt to changing conditions during the simulation and to provide an example using DFT, a simulation of the melting of a fcc-phase bulk aluminum 2 × 2 × 2 supercell was conducted using a 0.5 fs time step. The 32-atom system was initialized at 300 K for 5 ps, followed by a temperature ramp to 3000 K for 1.25 ps. Subsequently, the temperature was held at 3000 K for the remainder of the simulation. A long simulation of 250 ps was conducted using GPR with a low accuracy threshold of 0.05 eV. During the heating phase and subsequent equilibration of the system, the number of required training steps is relatively high, totaling at 500 in the first 25 ps of the simulation (Fig. 12). After the system has adapted to the temperature change only 69 additional DFT calculations are required in the remaining 225 ps of the simulation.
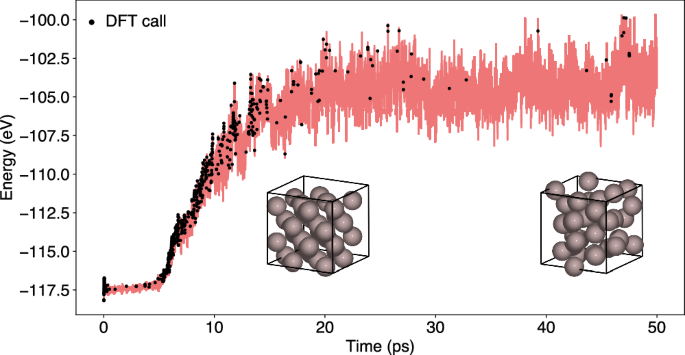
The simulation was initialized at 300 K for 5 ps, followed by a linear temperature ramp to 3000 K over 1.25 ps. The system was then maintained at 3000 K for the remaining simulation. If the predicted energy uncertainty ΔE of the GPR model exceeds the accuracy threshold ϵ of 0.05 eV, a DFT calculation is performed and the ML model is trained (black dots).
The radial distribution function (RDF) of the bulk aluminum system was sampled over two time intervals during the simulation: from 2 ps to 5 ps, corresponding to the solid phase at 300 K, and from 19 ps to 25 ps, after the system had been heated to 3000 K and the melting was complete. The RDFs sampled from the OTF-MD simulation show very good agreement with those sampled from an identical AIMD simulation, showing the ability of the ML model to accurately simulate the melting process of a bulk metal system with periodic boundary conditions using DFT (Fig. 13).
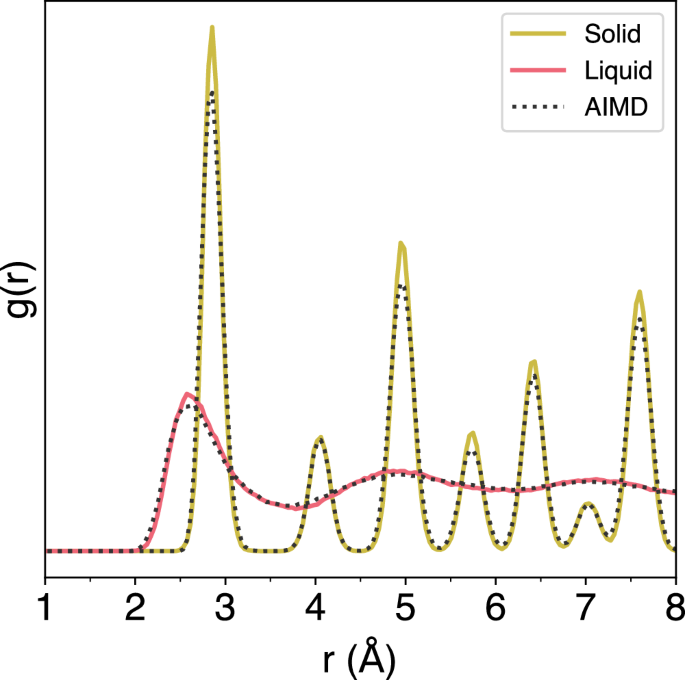
The colored RDFs are sampled from an OTF-MD simulation using classical GPR with an accuracy threshold of 0.05 eV. They show agreement with the RDFs obtained an identical AIMD simulation.
While the GPR model used in all previous simulations is only trained on the energy of the structures, it is possible to additionally train on the atomic forces. Naturally, this leads to increased training times, which is why it is sensible to use the SparseGPR model implemented in AGOX, as it applies sparsification to speed up the training process41. To compare the effects of force training and sparsification, the melting of bulk aluminum was simulated using three different ML models: GPR trained only on energies (GPR), SparseGPR trained on both energy and atomic forces (SparseGPR) and standard GPR without sparsification trained on energies and forces (GPR + Forces). For each model two simulations were performed with accuracy thresholds of 0.05 eV and 0.10 eV and the corresponding errors of the energy and the maximum atomic force were computed (Fig. 14). As expected, the models with force training show lower errors of the force. However, force training results in a higher error of the energy. This can be attributed to the fact that a model trained exclusively on energies is more optimized for energy prediction, whereas the additional training on atomic forces increases the complexity of the model and decreases accuracy of energy predictions.
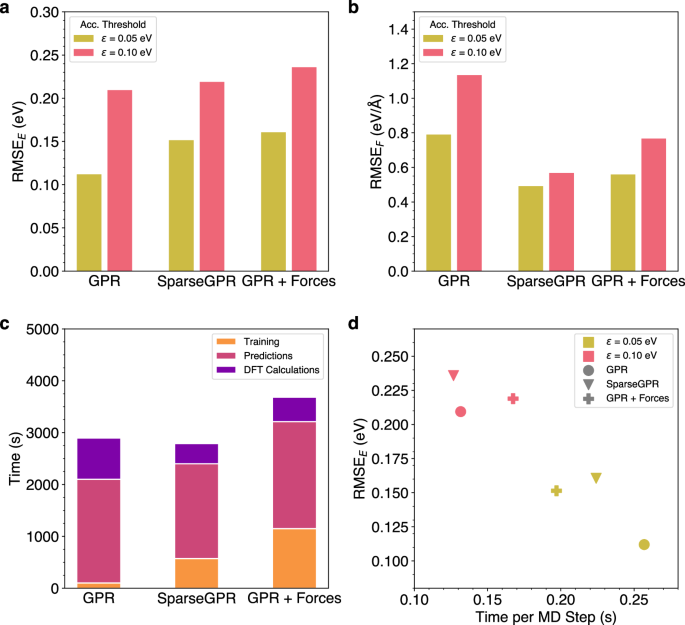
a RMSE of the Energy for OTF-MD-Simulations of a melting Al crystal for three different ML models at accuracy thresholds of 0.05 eV and 0.10 eV. b RMSE of the maximum Force for OTF-MD-Simulations of a melting Al crystal for three different ML models at accuracy thresholds of 0.05 eV and 0.10 eV. c Breakdown of the time required for Training, Predictions and DFT calculations during the OTF-MD simulations with three different ML models. d RMSE of the Energy plotted against the Time per MD step for three different ML models during OTF MD simulations of an Al crystal with accuracy thresholds of 0.10 eV and 0.05 eV, respectively.
Additionally, diffusion coefficients of the aluminum in the melt were computed for different time frames during the simulations, comparing AIMD to GPR and SparseGPR with force training (Table 1). While the order of magnitude of the diffusion coefficients obtained from the OTF-MD simulations is comparable to that derived from the AIMD simulations over all analyzed time frames, both the deviation from the AIMD values and the uncertainties of the coefficients are higher for the time frames that start earlier in the trajectory. This can be attributed to the general effect that, due to a lower number of exact calculations, the OTF simulations require a longer time to equilibrate than the AIMD simulations.
Diffusion of water in a carbon nanotube
Beyond the ability to simulate single-element systems, the OTF-MD calculator can be applied to large multi-element systems. This is demonstrated by a simulation of the diffusion of a water molecule through a carbon nanotube (CNT). The study of such water diffusion inside CNTs has attracted great interest because they can be used as a model system for a variety of nanoporous systems having potential in many applications53. In our calculations, the system contains a single water molecule confined in a (5,5)-CNT of length 17.2 Å and diameter 5.424 Å within a 20 Å × 20 Å × 17.2 Å simulation cell under periodic boundary conditions. This results in a 115-atom system, which is initialized at 10 K. Subsequently, the system is heated to 400 K over 1000 time steps (0.5 fs per time step) and simulated at this temperature for a total of 50,000 time steps (25 ps). An energy threshold of 0.10 eV was chosen for the simulation. The simulation shows correct diffusion of the water molecule through the nanotube (Fig. 15). Only 151 exact DFT calculations are needed for the 50,000 time steps. This is especially valuable since one single-point calculation takes on average 15.2 min (again on 16 cores of an Intel Xeon 8362 CPU). As a result, DFT calculations for all time steps would require over 500 days, while the OTF-MD calculator completes the simulation in 38.4 h. This makes it more than 300 times faster than the conventional AIMD method, showcasing its value in MD simulations of large systems.
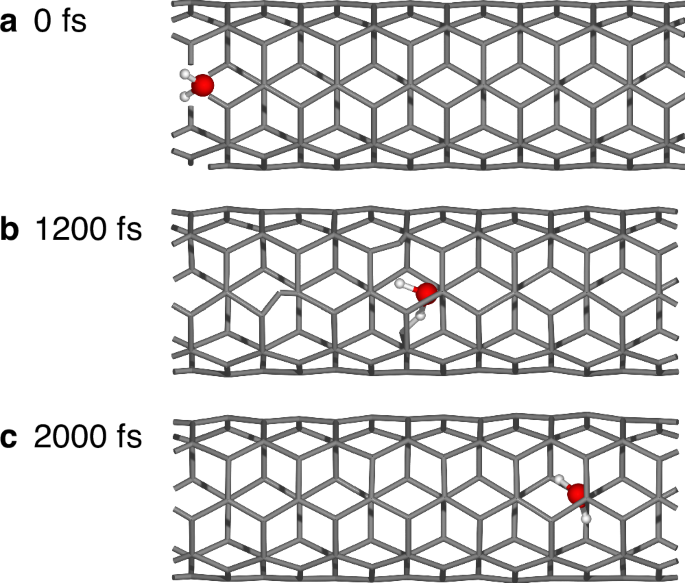
Snapshots taken at a 0 fs, b 1200 fs and c 2000 fs.
The ML model generated during a FALCON-OTF can be easily employed in subsequent non-OTF simulations using a static model. The training dataset produced during OTF training is saved in an ASE trajectory file and can be reused as training data for any static ML model of choice. This can either be performed in FALCON or other packages. Furthermore, FALCON provides functions to save the trained AGOX model to a file and reload it for different simulations. To ensure a complete representation of all relevant structures in the training data, FALCON offers a postprocessing script that identifies the most uncertain points in a MD trajectory after an OTF run. These structures are selected, saved in a trajectory file, and can be added to the training data before generating a static model. To avoid duplicate entries, all structures in the MD trajectory are clustered, and only the most uncertain structure from each cluster is selected.
For example, in the simulation of water in a CNT the script was employed to return relevant structures with an uncertainty higher than 0.09 eV (i.e., very close to the chosen accuracy threshold of 0.10 eV) as uncertain structures. This approach resulted in 33 individual, uncertain structures, which could be included in the training data to further improve a static model. Visualization shows that most of these uncertain structures originate from the end of the simulation (Fig. 16). Here, the uncertainty frequently was close the accuracy threshold, but only very infrequently exceeded it and triggered a retraining. Therefore, adding the uncertain structures to the training data can improve the representation of data, also in the late stages of the simulation.
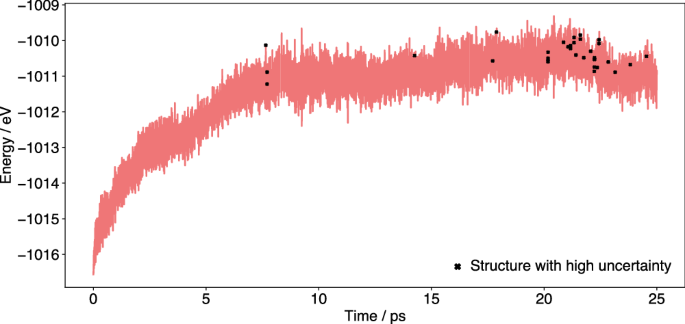
The black markers indicate 33 relevant structures with the high uncertainty, identified by postprocessing, which could be added to the training dataset before generating a static ML model for a non-OTF simulation.
Transfer learning
The benefit of OTF training is the possibility of training a very accurate model designed for a specific system. However, the constant retraining might become a bottleneck, especially, if the system size increases. Therefore, FALCON has the ability to train on smaller systems and use the data as training data for repeated systems. Note, currently only systems with a number of atoms that is a multiple of the small simulation cell (e.g. a supercell of a periodic system) can be used. To demonstrate the ability, we use LiFePO4 as a toy system. We start with three independent runs for 5 ps and an accuracy threshold of 0.25 eV at 400 K, 600 K and 800 K, respectively. The FALCON simulations are performed on a single unit cell consisting of 28 atoms. The training dataset of this cell is used to create the corresponding 1 × 2 × 2 data, which serves as the initial training data for the FALCON simulation of the supercell. Here, an increased accuracy threshold (less frequent retraining) of 0.5 eV for a timeframe of 5 ps is employed with a timestep of 0.5 fs. We advise performing at least a short active learning phase for the bigger cell before starting a non-OTF run, because the training data only consists of symmetrically displaced atoms, as a result of repeating the single cell when creating the supercell. Adding a few motifs of less symmetric displacements to the bigger cell, will help to create a better ML model (or multiple models in case of clustering). In a final step, we run the simulation without further retraining in a non-OTF FALCON run. This can be done in the same script. No restart is needed. However, alternatively one could use an increased threshold of e.g. 1 eV instead which additionally increases stepwise (e.g. +0.05 eV) with each MD step. The resulting energy profile can be found in Fig. 17. Just for the first 5 ps a traditional AIMD run would take 85 days (using 64 cores of an Intel Xeon 8362 CPU). We were able to perform this active learning run in less than 12 h, including the creation of the training data in the smaller cells. The trained ML model can be saved and loaded for additional runs. In addition, structures, energies and forces of all exact calculations during the OTF part of the calculation will always be saved in a separate trajectory and can be used for training of new models. Here we only show the next 45 ps of the simulation, although now the model can reach the ns scale within reasonable time.
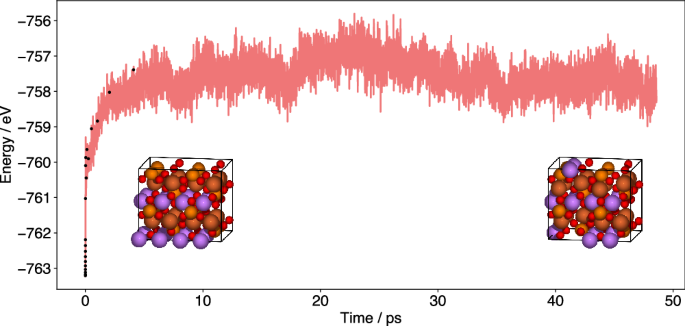
After generating the initial training data from a smaller cell, only the first 10 ps were performed as an OTF run with active learning, while afterwards the generated static model was used in a non-OTF run.




