GMAI models promise to solve more diverse and challenging tasks than current medical AI models, even while requiring little to no labels for specific tasks. Of the three defining capabilities of GMAI, two enable flexible interactions between the GMAI model and the user: first, the ability to carry out tasks that are dynamically specified; and second, the ability to support flexible combinations of data modalities. The third capability requires that GMAI models formally represent medical domain knowledge and leverage it to carry out advanced medical reasoning. Recent foundation models already exhibit individual aspects of GMAI, by flexibly combining several modalities2 or making it possible to dynamically specify a new task at test time5, but substantial advances are still required to build a GMAI model with all three capabilities. For example, existing models that show medical reasoning abilities (such as GPT-3 or PaLM) are not multimodal and do not yet generate reliably factual statements.
Flexible interactions
GMAI offers users the ability to interact with models through custom queries, making AI insights easier for different audiences to understand and offering unprecedented flexibility across tasks and settings. In current practice, AI models typically handle a narrow set of tasks and produce a rigid, predetermined set of outputs. For example, a current model might detect a specific disease, taking in one kind of image and always outputting the likelihood of that disease. By contrast, a custom query allows users to come up with questions on the fly: “Explain the mass appearing on this head MRI scan. Is it more likely a tumour or an abscess?”. Furthermore, queries can allow users to customize the format of their outputs: “This is a follow-up MRI scan of a patient with glioblastoma. Outline any tumours in red”.
Custom queries will enable two key capabilities—dynamic task specification and multimodal inputs and outputs—as follows.
Dynamic task specification
Custom queries can teach AI models to solve new problems on the fly, dynamically specifying new tasks without requiring models to be retrained. For example, GMAI can answer highly specific, previously unseen questions: “Given this ultrasound, how thick is the gallbladder wall in millimetres?”. Unsurprisingly, a GMAI model may struggle to complete new tasks that involve unknown concepts or pathologies. In-context learning then allows users to teach the GMAI about a new concept with few examples: “Here are the medical histories of ten previous patients with an emerging disease, an infection with the Langya henipavirus. How likely is it that our current patient is also infected with Langya henipavirus?”17.
Multimodal inputs and outputs
Custom queries can allow users to include complex medical information in their questions, freely mixing modalities. For example, a clinician might include multiple images and laboratory results in their query when asking for a diagnosis. GMAI models can also flexibly incorporate different modalities into responses, such as when a user asks for both a text answer and an accompanying visualization. Following previous models such as Gato, GMAI models can combine modalities by turning each modality’s data into ‘tokens’, each representing a small unit (for example, a word in a sentence or a patch in an image) that can be combined across modalities. This blended stream of tokens can then be fed into a transformer architecture18, allowing GMAI models to integrate a given patient’s entire history, including reports, waveform signals, laboratory results, genomic profiles and imaging studies.
Medical domain knowledge
In stark contrast to a clinician, conventional medical AI models typically lack prior knowledge of the medical domain before they are trained for their particular tasks. Instead, they have to rely solely on statistical associations between features of the input data and the prediction target, without having contextual information (for example, about pathophysiological processes). This lack of background makes it harder to train models for specific medical tasks, particularly when data for the tasks are scarce.
GMAI models can address these shortcomings by formally representing medical knowledge. For example, structures such as knowledge graphs can allow models to reason about medical concepts and relationships between them. Furthermore, building on recent retrieval-based approaches, GMAI can retrieve relevant context from existing databases, in the form of articles, images or entire previous cases19,20.
The resulting models can raise self-explanatory warnings: “This patient is likely to develop acute respiratory distress syndrome, because the patient was recently admitted with a severe thoracic trauma and because the patient’s partial pressure of oxygen in the arterial blood has steadily decreased, despite an increased inspired fraction of oxygen”.
As a GMAI model may even be asked to provide treatment recommendations, despite mostly being trained on observational data, the model’s ability to infer and leverage causal relationships between medical concepts and clinical findings will play a key role for clinical applicability21.
Finally, by accessing rich molecular and clinical knowledge, a GMAI model can solve tasks with limited data by drawing on knowledge of related problems, as exemplified by initial works on AI-based drug repurposing22.
Use cases of GMAI
We present six potential use cases for GMAI that target different user bases and disciplines, although our list is hardly exhaustive. Although there have already been AI efforts in these areas, we expect GMAI will enable comprehensive solutions for each problem.
Grounded radiology reports
GMAI enables a new generation of versatile digital radiology assistants, supporting radiologists throughout their workflow and markedly reducing workloads. GMAI models can automatically draft radiology reports that describe both abnormalities and relevant normal findings, while also taking into account the patient’s history. These models can provide further assistance to clinicians by pairing text reports with interactive visualizations, such as by highlighting the region described by each phrase. Radiologists can also improve their understanding of cases by chatting with GMAI models: “Can you highlight any new multiple sclerosis lesions that were not present in the previous image?”.
A solution needs to accurately interpret various radiology modalities, noticing even subtle abnormalities. Furthermore, it must integrate information from a patient’s history, including sources such as indications, laboratory results and previous images, when describing an image. It also needs to communicate with clinicians using multiple modalities, providing both text answers and dynamically annotated images. To do so, it must be capable of visual grounding, accurately pointing out exactly which part of an image supports any statement. Although this may be achieved through supervised learning on expert-labelled images, explainability methods such as Grad-CAM could enable self-supervised approaches, requiring no labelled data23.
Augmented procedures
We anticipate a surgical GMAI model that can assist surgical teams with procedures: “We cannot find the intestinal rupture. Check whether we missed a view of any intestinal section in the visual feed of the last 15 minutes”. GMAI models may carry out visualization tasks, potentially annotating video streams of a procedure in real time. They may also provide information in spoken form, such as by raising alerts when steps of a procedure are skipped or by reading out relevant literature when surgeons encounter rare anatomical phenomena.

a, A GMAI model is trained on multiple medical data modalities, through techniques such as self-supervised learning. To enable flexible interactions, data modalities such as images or data from EHRs can be paired with language, either in the form of text or speech data. Next, the GMAI model needs to access various sources of medical knowledge to carry out medical reasoning tasks, unlocking a wealth of capabilities that can be used in downstream applications. The resulting GMAI model then carries out tasks that the user can specify in real time. For this, the GMAI model can retrieve contextual information from sources such as knowledge graphs or databases, leveraging formal medical knowledge to reason about previously unseen tasks. b, The GMAI model builds the foundation for numerous applications across clinical disciplines, each requiring careful validation and regulatory assessment.
This model can also assist with procedures outside the operating room, such as with endoscopic procedures. A model that captures topographic context and reasons with anatomical knowledge can draw conclusions about previously unseen phenomena. For instance, it could deduce that a large vascular structure appearing in a duodenoscopy may indicate an aortoduodenal fistula (that is, an abnormal connection between aorta and the small intestine), despite never having encountered one before (Fig. 2, right panel). GMAI can solve this task by first detecting the vessel, second identifying the anatomical location, and finally considering the neighbouring structures.
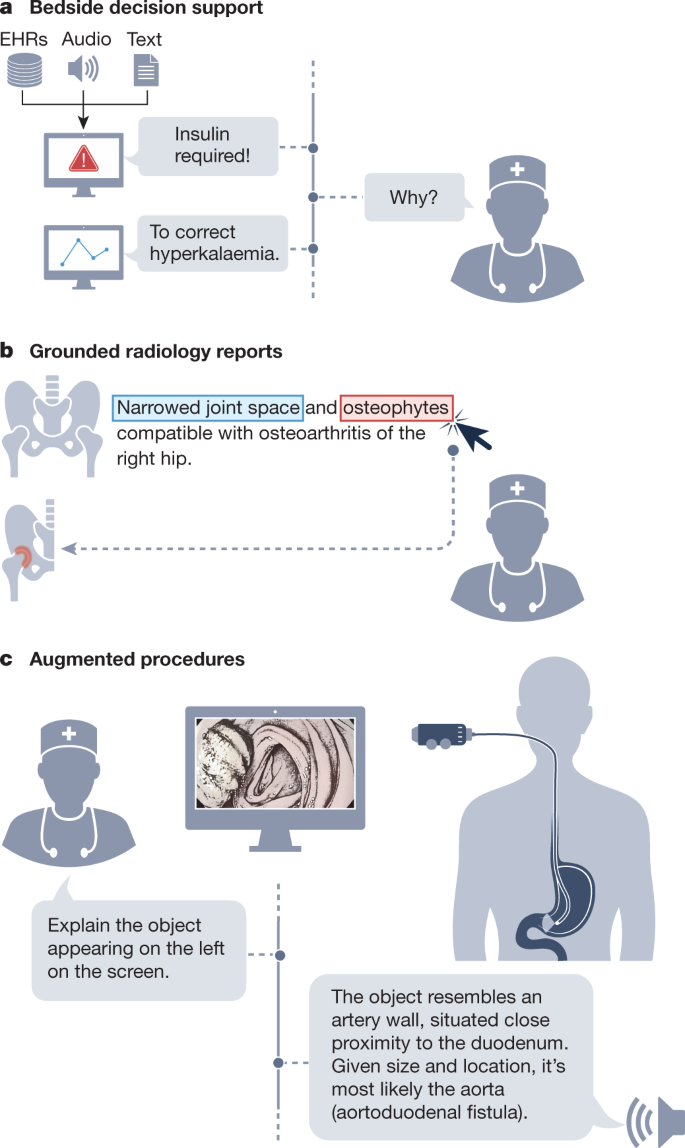
a, GMAI could enable versatile and self-explanatory bedside decision support. b, Grounded radiology reports are equipped with clickable links for visualizing each finding. c, GMAI has the potential to classify phenomena that were never encountered before during model development. In augmented procedures, a rare outlier finding is explained with step-by-step reasoning by leveraging medical domain knowledge and topographic context. The presented example is inspired by a case report58. Image of the fistula in panel c adapted from ref. 58, CC BY 3.0.
A solution needs to integrate vision, language and audio modalities, using a vision–audio–language model to accept spoken queries and carry out tasks using the visual feed. Vision–language models have already gained traction, and the development of models that incorporate further modalities is merely a question of time24. Approaches may build on previous work that combines language models and knowledge graphs25,26 to reason step-by-step about surgical tasks. Additionally, GMAI deployed in surgical settings will probably face unusual clinical phenomena that cannot be included during model development, owing to their rarity, a challenge known as the long tail of unseen conditions27. Medical reasoning abilities will be crucial for both detecting previously unseen outliers and explaining them, as exemplified in Fig. 2.
Bedside decision support
GMAI enables a new class of bedside clinical decision support tools that expand on existing AI-based early warning systems, providing more detailed explanations as well as recommendations for future care. For example, GMAI models for bedside decision support can leverage clinical knowledge and provide free-text explanations and data summaries: “Warning: This patient is about to go into shock. Her circulation has destabilized in the last 15 minutes <link to data summary>. Recommended next steps: <link to checklist>”.
A solution needs to parse electronic health record (EHR) sources (for example, vital and laboratory parameters, and clinical notes) that involve multiple modalities, including text and numeric time series data. It needs to be able to summarize a patient’s current state from raw data, project potential future states of the patient and recommend treatment decisions. A solution may project how a patient’s condition will change over time, by using language modelling techniques to predict their future textual and numeric records from their previous data. Training datasets may specifically pair EHR time series data with eventual patient outcomes, which can be collected from discharge reports and ICD (International Classification of Diseases) codes. In addition, the model must be able to compare potential treatments and estimate their effects, all while adhering to therapeutic guidelines and other relevant policies. The model can acquire the necessary knowledge through clinical knowledge graphs and text sources such as academic publications, educational textbooks, international guidelines and local policies. Approaches may be inspired by REALM, a language model that answers queries by first retrieving a single relevant document and then extracting the answer from it, making it possible for users to identify the exact source of each answer20.
Interactive note-taking
Documentation represents an integral but labour-intensive part of clinical workflows. By monitoring electronic patient information as well as clinician–patient conversations, GMAI models will preemptively draft documents such as electronic notes and discharge reports for clinicians to merely review, edit and approve. Thus, GMAI can substantially reduce administrative overhead, allowing clinicians to spend more time with patients.
A GMAI solution can draw from recent advances in speech-to-text models28, specializing techniques for medical applications. It must accurately interpret speech signals, understanding medical jargon and abbreviations. Additionally, it must contextualize speech data with information from the EHRs (for example, diagnosis list, vital parameters and previous discharge reports) and then generate free-text notes or reports. It will be essential to obtain consent before recording any interaction with a patient. Even before such recordings are collected in large numbers, early note-taking models may already be developed by leveraging clinician–patient interaction data collected from chat applications.
Chatbots for patients
GMAI has the potential to power new apps for patient support, providing high-quality care even outside clinical settings. For example, GMAI can build a holistic view of a patient’s condition using multiple modalities, ranging from unstructured descriptions of symptoms to continuous glucose monitor readings to patient-provided medication logs. After interpreting these heterogeneous types of data, GMAI models can interact with the patient, providing detailed advice and explanations. Importantly, GMAI enables accessible communication, providing clear, readable or audible information on the patient’s schedule. Whereas similar apps rely on clinicians to offer personalized support at present29, GMAI promises to reduce or even remove the need for human expert intervention, making apps available on a larger scale. As with existing live chat applications, users could still engage with a human counsellor on request.
Building patient-facing chatbots with GMAI raises two special challenges. First, patient-facing models must be able to communicate clearly with non-technical audiences, using simple, clear language without sacrificing the accuracy of the content. Including patient-focused medical texts in training datasets may enable this capability. Second, these models need to work with diverse data collected by patients. Patient-provided data may represent unusual modalities; for example, patients with strict dietary requirements may submit before-and-after photos of their meals so that GMAI models can automatically monitor their food intake. Patient-collected data are also likely to be noisier compared to data from a clinical setting, as patients may be more prone to error or use less reliable devices when collecting data. Again, incorporating relevant data into training can help overcome this challenge. However, GMAI models also need to monitor their own uncertainty and take appropriate action when they do not have enough reliable data.
Text-to-protein generation
GMAI could generate protein amino acid sequences and their three-dimensional structures from textual prompts. Inspired by existing generative models of protein sequences30, such a model could condition its generation on desired functional properties. By contrast, a biomedically knowledgeable GMAI model promises protein design interfaces that are as flexible and easy to use as concurrent text-to-image generative models such as Stable Diffusion or DALL-E31,32. Moreover, by unlocking in-context learning capabilities, a GMAI-based text-to-protein model may be prompted with a handful of example instructions paired with sequences to dynamically define a new generation task, such as the generation of a protein that binds with high affinity to a specified target while fulfilling additional constraints.
There have already been early efforts to develop foundation models for biological sequences33,34, including RFdiffusion, which generates proteins on the basis of simple specifications (for example, a binding target)35. Building on this work, GMAI-based solution can incorporate both language and protein sequence data during training to offer a versatile text interface. A solution could also draw on recent advances in multimodal AI such as CLIP, in which models are jointly trained on paired data of different modalities16. When creating such a training dataset, individual protein sequences must be paired with relevant text passages (for example, from the body of biological literature) that describe the properties of the proteins. Large-scale initiatives, such as UniProt, that map out protein functions for millions of proteins, will be indispensable for this effort36.



